Metagenome on:
[Wikipedia]
[Google]
[Amazon]
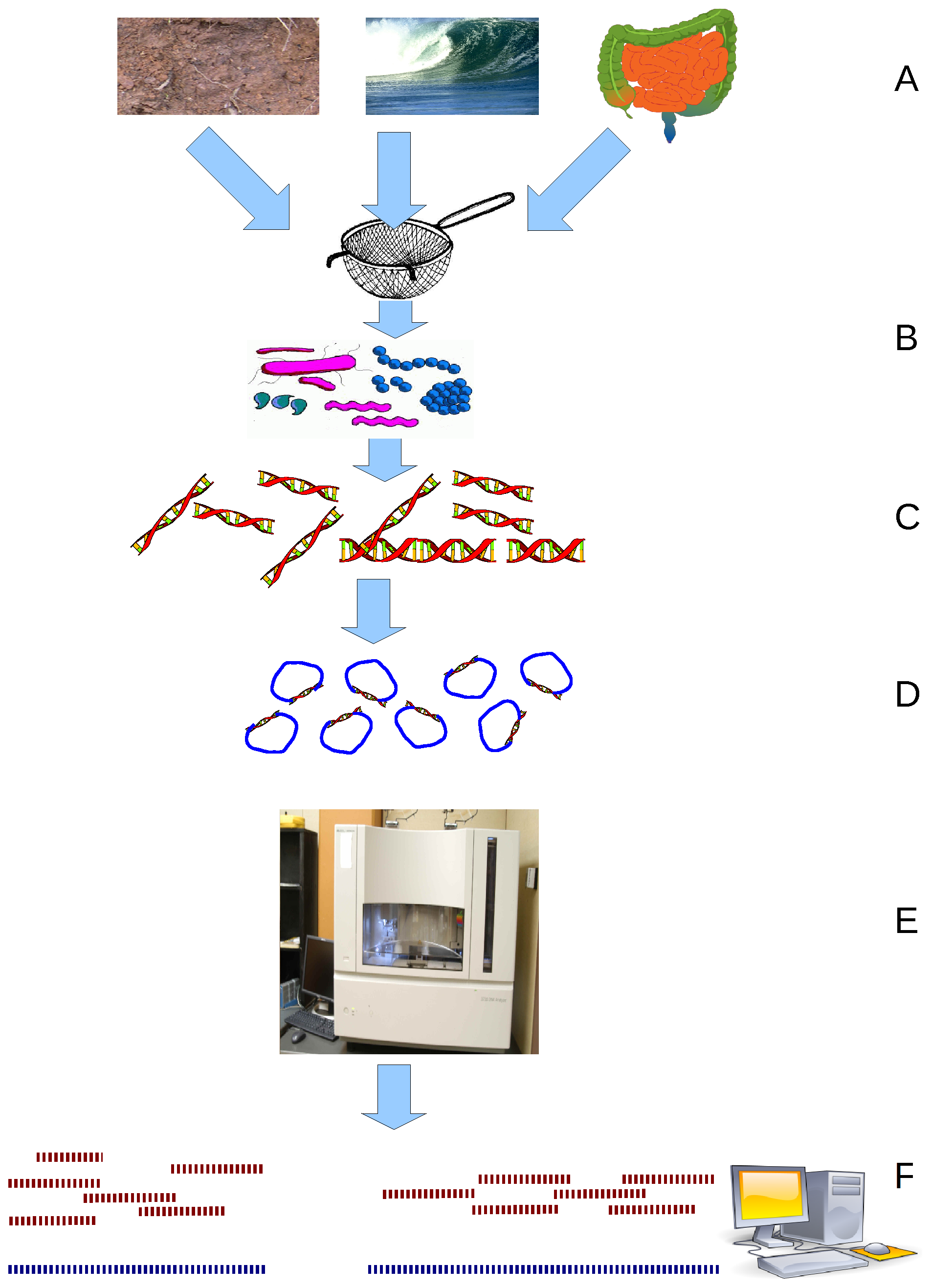 Metagenomics is the study of genetic material recovered directly from
Metagenomics is the study of genetic material recovered directly from
 The data generated by metagenomics experiments are both enormous and inherently noisy, containing fragmented data representing as many as 10,000 species. The sequencing of the cow
The data generated by metagenomics experiments are both enormous and inherently noisy, containing fragmented data representing as many as 10,000 species. The sequencing of the cow
and
mOTUs
ref name="sunagawa2013"/> and MetaPhyler, use universal marker genes to profile prokaryotic species. With th
mOTUs profiler
is possible to profile species without a reference genome, improving the estimation of microbial community diversity. Recent methods, such a
SLIMM
use read coverage landscape of individual reference genomes to minimize false-positive hits and get reliable relative abundances. In composition based binning, methods use intrinsic features of the sequence, such as oligonucleotide frequencies or
Integrated Microbial Genomes
(IMG) system and th
project. One of the first standalone tools for analysing high-throughput metagenome shotgun data was
SEED
or
 Metagenomics can provide valuable insights into the functional ecology of environmental communities. Metagenomic analysis of the bacterial consortia found in the defecations of Australian sea lions suggests that nutrient-rich sea lion faeces may be an important nutrient source for coastal ecosystems. This is because the bacteria that are expelled simultaneously with the defecations are adept at breaking down the nutrients in the faeces into a bioavailable form that can be taken up into the food chain.
DNA sequencing can also be used more broadly to identify species present in a body of water, debris filtered from the air, sample of dirt, or animal's faeces, and even detect diet items from blood meals. This can establish the range of
Metagenomics can provide valuable insights into the functional ecology of environmental communities. Metagenomic analysis of the bacterial consortia found in the defecations of Australian sea lions suggests that nutrient-rich sea lion faeces may be an important nutrient source for coastal ecosystems. This is because the bacteria that are expelled simultaneously with the defecations are adept at breaking down the nutrients in the faeces into a bioavailable form that can be taken up into the food chain.
DNA sequencing can also be used more broadly to identify species present in a body of water, debris filtered from the air, sample of dirt, or animal's faeces, and even detect diet items from blood meals. This can establish the range of
Focus on Metagenomics
at ''
The “Critical Assessment of Metagenome Interpretation” (CAMI) initiative
to evaluate methods in metagenomics {{Portal bar, Biology, Medicine Bioinformatics Genomics Environmental microbiology Microbiology techniques
 Metagenomics is the study of genetic material recovered directly from
Metagenomics is the study of genetic material recovered directly from environmental
A biophysical environment is a biotic and abiotic surrounding of an organism or population, and consequently includes the factors that have an influence in their survival, development, and evolution. A biophysical environment can vary in scal ...
or clinical samples by a method called sequencing
In genetics and biochemistry, sequencing means to determine the primary structure (sometimes incorrectly called the primary sequence) of an unbranched biopolymer. Sequencing results in a symbolic linear depiction known as a sequence which suc ...
. The broad field may also be referred to as environmental genomics, ecogenomics, community genomics or microbiomics.
While traditional microbiology
Microbiology () is the scientific study of microorganisms, those being unicellular (single cell), multicellular (cell colony), or acellular (lacking cells). Microbiology encompasses numerous sub-disciplines including virology, bacteriology, ...
and microbial genome sequencing
Whole genome sequencing (WGS), also known as full genome sequencing, complete genome sequencing, or entire genome sequencing, is the process of determining the entirety, or nearly the entirety, of the DNA sequence of an organism's genome at a ...
and genomics
Genomics is an interdisciplinary field of biology focusing on the structure, function, evolution, mapping, and editing of genomes. A genome is an organism's complete set of DNA, including all of its genes as well as its hierarchical, three-dim ...
rely upon cultivated clonal cultures
Culture () is an umbrella term which encompasses the social behavior, institutions, and norms found in human societies, as well as the knowledge, beliefs, arts, laws, customs, capabilities, and habits of the individuals in these groups.Tylo ...
, early environmental gene sequencing cloned specific genes (often the 16S rRNA 16S rRNA may refer to:
* 16S ribosomal RNA
16 S ribosomal RNA (or 16 S rRNA) is the RNA component of the 30S subunit of a prokaryotic ribosome ( SSU rRNA). It binds to the Shine-Dalgarno sequence and provides most of the SSU structure.
The g ...
gene) to produce a profile of diversity in a natural sample. Such work revealed that the vast majority of microbial biodiversity had been missed by cultivation-based methods.
Because of its ability to reveal the previously hidden diversity of microscopic life, metagenomics offers a powerful lens for viewing the microbial world that has the potential to revolutionize understanding of the entire living world. As the price of DNA sequencing continues to fall, metagenomics now allows microbial ecology
Microbial ecology (or environmental microbiology) is the ecology of microorganisms: their relationship with one another and with their environment. It concerns the three major domains of life—Eukaryota, Archaea, and Bacteria—as well as viru ...
to be investigated at a much greater scale and detail than before. Recent studies use either "shotgun
A shotgun (also known as a scattergun, or historically as a fowling piece) is a long-barreled firearm designed to shoot a straight-walled cartridge known as a shotshell, which usually discharges numerous small pellet-like spherical sub- pr ...
" or PCR directed sequencing to get largely unbiased samples of all genes from all the members of the sampled communities.
Etymology
The term "metagenomics" was first used byJo Handelsman
Jo Emily Handelsman (born March 19, 1959 in New York City) is the Director of the Wisconsin Institute for Discovery at University of Wisconsin–Madison. She is also a Vilas Research Professor and a Howard Hughes Medical Institute Professor. Dr. H ...
, Robert M. Goodman
Robert "Bob" M. Goodman (born December 30, 1945) is a prominent plant biologist and virologist, and has served as the executive dean of agriculture and natural resources at Rutgers, The State University of New Jersey since June 2005. He is the exe ...
, Michelle R. Rondon, Jon Clardy
Jon Clardy (born May 16, 1943, Washington, D.C., United States) is currently the Hsien Wu and Daisy Yen Wu professor of biological chemistry and molecular pharmacology at Harvard Medical School. His research focuses on the isolation and structur ...
, and Sean F. Brady, and first appeared in publication in 1998. The term metagenome referenced the idea that a collection of genes sequenced from the environment could be analyzed in a way analogous to the study of a single genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ...
. In 2005, Kevin Chen and Lior Pachter
Lior Samuel Pachter is a computational biologist. He works at the California Institute of Technology, where he is the Bren Professor of Computational Biology. He has widely varied research interests including genomics, combinatorics, computational ...
(researchers at the University of California, Berkeley
The University of California, Berkeley (UC Berkeley, Berkeley, Cal, or California) is a public land-grant research university in Berkeley, California. Established in 1868 as the University of California, it is the state's first land-grant un ...
) defined metagenomics as "the application of modern genomics technique without the need for isolation and lab cultivation of individual species".
History
Conventionalsequencing
In genetics and biochemistry, sequencing means to determine the primary structure (sometimes incorrectly called the primary sequence) of an unbranched biopolymer. Sequencing results in a symbolic linear depiction known as a sequence which suc ...
begins with a culture of identical cells as a source of DNA. However, early metagenomic studies revealed that there are probably large groups of microorganisms in many environments that cannot be cultured and thus cannot be sequenced. These early studies focused on 16S ribosomal
Ribosomes ( ) are macromolecular machines, found within all cells, that perform biological protein synthesis (mRNA translation). Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to for ...
RNA (rRNA) sequences which are relatively short, often conserved within a species, and generally different between species. Many 16S rRNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribos ...
sequences have been found which do not belong to any known cultured species
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriat ...
, indicating that there are numerous non-isolated organisms. These surveys of ribosomal RNA genes taken directly from the environment revealed that cultivation based methods find less than 1% of the bacterial and archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaeba ...
l species in a sample. Much of the interest in metagenomics comes from these discoveries that showed that the vast majority of microorganisms had previously gone unnoticed.
In the 1980s early molecular work in the field was conducted by Norman R. Pace and colleagues, who used PCR to explore the diversity of ribosomal RNA sequences. The insights gained from these breakthrough studies led Pace to propose the idea of cloning DNA directly from environmental samples as early as 1985. This led to the first report of isolating and cloning
Cloning is the process of producing individual organisms with identical or virtually identical DNA, either by natural or artificial means. In nature, some organisms produce clones through asexual reproduction. In the field of biotechnology, c ...
bulk DNA from an environmental sample, published by Pace and colleagues in 1991 while Pace was in the Department of Biology at Indiana University
Indiana University (IU) is a system of public universities in the U.S. state of Indiana.
Campuses
Indiana University has two core campuses, five regional campuses, and two regional centers under the administration of IUPUI.
*Indiana Universi ...
. Considerable efforts ensured that these were not PCR false positives and supported the existence of a complex community of unexplored species. Although this methodology was limited to exploring highly conserved, non-protein coding genes, it did support early microbial morphology-based observations that diversity was far more complex than was known by culturing methods. Soon after that in 1995, Healy reported the metagenomic isolation of functional genes from "zoolibraries" constructed from a complex culture of environmental organisms grown in the laboratory on dried grass
Poaceae () or Gramineae () is a large and nearly ubiquitous family of monocotyledonous flowering plants commonly known as grasses. It includes the cereal grasses, bamboos and the grasses of natural grassland and species cultivated in lawns a ...
es. After leaving the Pace laboratory, Edward DeLong
Edward Francis DeLong (born 1958), is a marine microbiologist and professor in the Department of Oceanography at the University of Hawaii, Manoa, and is considered a pioneer in the field of metagenomics. He is best known for his discovery of the ...
continued in the field and has published work that has largely laid the groundwork for environmental phylogenies based on signature 16S sequences, beginning with his group's construction of libraries from marine
Marine is an adjective meaning of or pertaining to the sea or ocean.
Marine or marines may refer to:
Ocean
* Maritime (disambiguation)
* Marine art
* Marine biology
* Marine debris
* Marine habitats
* Marine life
* Marine pollution
Military ...
samples.
In 2002, Mya Breitbart, Forest Rohwer, and colleagues used environmental shotgun sequencing
In genetics, shotgun sequencing is a method used for sequencing random DNA strands. It is named by analogy with the rapidly expanding, quasi-random shot grouping of a shotgun.
The chain-termination method of DNA sequencing ("Sanger sequencing ...
(see below) to show that 200 liters of seawater contains over 5000 different viruses. Subsequent studies showed that there are more than a thousand viral species in human stool and possibly a million different viruses per kilogram of marine sediment
Marine sediment, or ocean sediment, or seafloor sediment, are deposits of insoluble particles that have accumulated on the seafloor. These particles have their origins in soil and rocks and have been transported from the land to the sea, mai ...
, including many bacteriophages
A bacteriophage (), also known informally as a ''phage'' (), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν ('), meaning "to devour". Bacterio ...
. Essentially all of the viruses in these studies were new species. In 2004, Gene Tyson, Jill Banfield, and colleagues at the University of California, Berkeley
The University of California, Berkeley (UC Berkeley, Berkeley, Cal, or California) is a public land-grant research university in Berkeley, California. Established in 1868 as the University of California, it is the state's first land-grant un ...
and the Joint Genome Institute
The U.S. Department of Energy (DOE) Joint Genome Institute (JGI), first located in Walnut Creek then Berkeley, California, was created in 1997 to unite the expertise and resources in genome mapping, DNA sequencing, technology development, and i ...
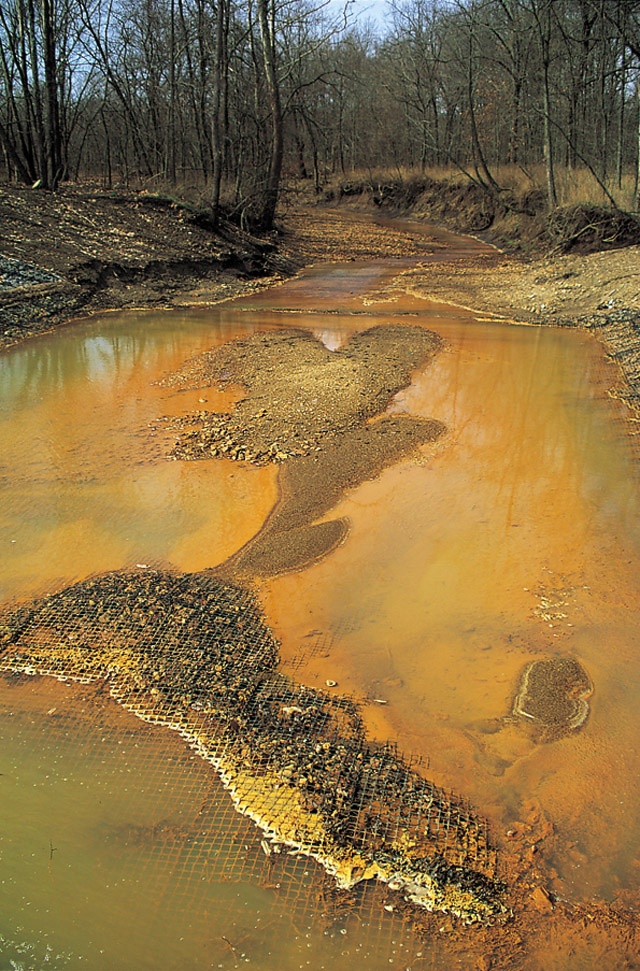
sequenced DNA extracted from an acid mine drainage
Acid mine drainage, acid and metalliferous drainage (AMD), or acid rock drainage (ARD) is the outflow of acidic water from metal mines or coal mines.
Acid rock drainage occurs naturally within some environments as part of the rock weathering ...
system. This effort resulted in the complete, or nearly complete, genomes for a handful of bacteria and archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaeba ...
that had previously resisted attempts to culture them.
Beginning in 2003, Craig Venter
John Craig Venter (born October 14, 1946) is an American biotechnologist and businessman. He is known for leading one of the first draft sequences of the human genome and assembled the first team to transfect a cell with a synthetic chromosome. ...
, leader of the privately funded parallel of the Human Genome Project
The Human Genome Project (HGP) was an international scientific research project with the goal of determining the base pairs that make up human DNA, and of identifying, mapping and sequencing all of the genes of the human genome from both ...
, has led the Global Ocean Sampling Expedition
The Global Ocean Sampling Expedition (GOS) is an ocean exploration genome project whose goal is to assess genetic diversity in marine microbial communities and to understand their role in nature's fundamental processes. It was begun as a Sargass ...
(GOS), circumnavigating the globe and collecting metagenomic samples throughout the journey. All of these samples were sequenced using shotgun sequencing, in hopes that new genomes (and therefore new organisms) would be identified. The pilot project, conducted in the Sargasso Sea
The Sargasso Sea () is a region of the Atlantic Ocean bounded by four currents forming an ocean gyre. Unlike all other regions called seas, it has no land boundaries. It is distinguished from other parts of the Atlantic Ocean by its charac ...
, found DNA from nearly 2000 different species
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriat ...
, including 148 types of bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were am ...
never before seen. Venter thoroughly explored the West Coast of the United States
The West Coast of the United States, also known as the Pacific Coast, Pacific states, and the western seaboard, is the coastline along which the Western United States meets the North Pacific Ocean. The term typically refers to the contiguous U.S ...
, and completed a two-year expedition to explore the Baltic
Baltic may refer to:
Peoples and languages
*Baltic languages, a subfamily of Indo-European languages, including Lithuanian, Latvian and extinct Old Prussian
*Balts (or Baltic peoples), ethnic groups speaking the Baltic languages and/or originatin ...
, Mediterranean
The Mediterranean Sea is a sea connected to the Atlantic Ocean, surrounded by the Mediterranean Basin and almost completely enclosed by land: on the north by Western and Southern Europe and Anatolia, on the south by North Africa, and on ...
and Black
Black is a color which results from the absence or complete absorption of visible light. It is an achromatic color, without hue, like white and grey. It is often used symbolically or figuratively to represent darkness. Black and white ha ...
Seas. Analysis of the metagenomic data collected during this journey revealed two groups of organisms, one composed of taxa adapted to environmental conditions of 'feast or famine', and a second composed of relatively fewer but more abundantly and widely distributed taxa primarily composed of plankton
Plankton are the diverse collection of organisms found in water (or air) that are unable to propel themselves against a current (or wind). The individual organisms constituting plankton are called plankters. In the ocean, they provide a cruc ...
.
In 2005 Stephan C. Schuster at Penn State University
The Pennsylvania State University (Penn State or PSU) is a Public university, public Commonwealth System of Higher Education, state-related Land-grant university, land-grant research university with campuses and facilities throughout Pennsylvan ...
and colleagues published the first sequences of an environmental sample generated with high-throughput sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. Th ...
, in this case massively parallel pyrosequencing
Pyrosequencing is a method of DNA sequencing (determining the order of nucleotides in DNA) based on the "sequencing by synthesis" principle, in which the sequencing is performed by detecting the nucleotide incorporated by a DNA polymerase. Pyrosequ ...
developed by 454 Life Sciences
454 Life Sciences was a biotechnology company based in Branford, Connecticut that specialized in high-throughput DNA sequencing. It was acquired by Roche in 2007 and shut down by Roche in 2013 when its technology became noncompetitive, although ...
. Another early paper in this area appeared in 2006 by Robert Edwards, Forest Rohwer, and colleagues at San Diego State University
San Diego State University (SDSU) is a public research university in San Diego, California. Founded in 1897 as San Diego Normal School, it is the third-oldest university and southernmost in the 23-member California State University (CSU) system ...
.
Sequencing
Recovery of DNA sequences longer than a few thousandbase pair
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both D ...
s from environmental samples was very difficult until recent advances in molecular biological techniques allowed the construction of libraries
A library is a collection of Document, materials, books or media that are accessible for use and not just for display purposes. A library provides physical (hard copies) or electronic media, digital access (soft copies) materials, and may be a ...
in bacterial artificial chromosome A bacterial artificial chromosome (BAC) is a DNA construct, based on a functional fertility plasmid (or F-plasmid), used for transforming and cloning in bacteria, usually '' E. coli''. F-plasmids play a crucial role because they contain partiti ...
s (BACs), which provided better vectors for molecular cloning
Molecular cloning is a set of experimental methods in molecular biology that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word '' cloning'' refers to the fact that the meth ...
.
Shotgun metagenomics
Advances inbioinformatics
Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combi ...
, refinements of DNA amplification, and the proliferation of computational power have greatly aided the analysis of DNA sequences recovered from environmental samples, allowing the adaptation of shotgun sequencing
In genetics, shotgun sequencing is a method used for sequencing random DNA strands. It is named by analogy with the rapidly expanding, quasi-random shot grouping of a shotgun.
The chain-termination method of DNA sequencing ("Sanger sequencing ...
to metagenomic samples (known also as whole metagenome shotgun or WMGS sequencing). The approach, used to sequence many cultured microorganisms and the human genome
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the ...
, randomly shears DNA, sequences many short sequences, and reconstructs them into a consensus sequence
In molecular biology and bioinformatics, the consensus sequence (or canonical sequence) is the calculated order of most frequent residues, either nucleotide or amino acid, found at each position in a sequence alignment. It serves as a simplified r ...
. Shotgun sequencing reveals genes present in environmental samples. Historically, clone libraries were used to facilitate this sequencing. However, with advances in high throughput sequencing technologies, the cloning step is no longer necessary and greater yields of sequencing data can be obtained without this labour-intensive bottleneck step. Shotgun metagenomics provides information both about which organisms are present and what metabolic processes are possible in the community. Because the collection of DNA from an environment is largely uncontrolled, the most abundant organisms in an environmental sample are most highly represented in the resulting sequence data. To achieve the high coverage needed to fully resolve the genomes of under-represented community members, large samples, often prohibitively so, are needed. On the other hand, the random nature of shotgun sequencing ensures that many of these organisms, which would otherwise go unnoticed using traditional culturing techniques, will be represented by at least some small sequence segments.
High-throughput sequencing
An advantage to high throughput sequencing is that this technique does not require cloning the DNA before sequencing, removing one of the main biases and bottlenecks in environmental sampling. The first metagenomic studies conducted usinghigh-throughput sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. Th ...
used massively parallel 454 pyrosequencing. Three other technologies commonly applied to environmental sampling are the Ion Torrent Personal Genome Machine, the Illumina MiSeq or HiSeq and the Applied Biosystems SOLiD system. These techniques for sequencing DNA generate shorter fragments than Sanger sequencing
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Fred ...
; Ion Torrent PGM System and 454 pyrosequencing typically produces ~400 bp reads, Illumina MiSeq produces 400-700bp reads (depending on whether paired end options are used), and SOLiD produce 25–75 bp reads. Historically, these read lengths were significantly shorter than the typical Sanger sequencing read length of ~750 bp, however the Illumina technology is quickly coming close to this benchmark. However, this limitation is compensated for by the much larger number of sequence reads. In 2009, pyrosequenced metagenomes generate 200–500 megabases, and Illumina platforms generate around 20–50 gigabases, but these outputs have increased by orders of magnitude in recent years.
An emerging approach combines shotgun sequencing and chromosome conformation capture
Chromosome conformation capture techniques (often abbreviated to 3C technologies or 3C-based methods) are a set of molecular biology methods used to analyze the spatial organization of chromatin in a cell. These methods quantify the number of int ...
(Hi-C), which measures the proximity of any two DNA sequences within the same cell, to guide microbial genome assembly. Long read sequencing technologies, including PacBio RSII and PacBio Sequel by Pacific Biosciences
Pacific Biosciences of California, Inc. (aka PacBio) is an American biotechnology company founded in 2004 that develops and manufactures systems for gene sequencing and some novel real time biological observation. PacBio describes its platform ...
, and Nanopore MinION, GridION, PromethION by Oxford Nanopore Technologies
Oxford Nanopore Technologies Limited is a UK-based company which is developing and selling nanopore sequencing products (including the portable DNA sequencer, MinION) for the direct, electronic analysis of single molecules.
History
The company ...
, is another choice to get long shotgun sequencing reads that should make ease in assembling process.
Bioinformatics
 The data generated by metagenomics experiments are both enormous and inherently noisy, containing fragmented data representing as many as 10,000 species. The sequencing of the cow
The data generated by metagenomics experiments are both enormous and inherently noisy, containing fragmented data representing as many as 10,000 species. The sequencing of the cow rumen
The rumen, also known as a paunch, is the largest stomach compartment in ruminants and the larger part of the reticulorumen, which is the first chamber in the alimentary canal of ruminant animals. The rumen's microbial favoring environment al ...
metagenome generated 279 gigabases, or 279 billion base pairs of nucleotide sequence data, while the human gut microbiome
A microbiome () is the community of microorganisms that can usually be found living together in any given habitat. It was defined more precisely in 1988 by Whipps ''et al.'' as "a characteristic microbial community occupying a reasonably wel ...
gene catalog identified 3.3 million genes assembled from 567.7 gigabases of sequence data. Collecting, curating, and extracting useful biological information from datasets of this size represent significant computational challenges for researchers.
Sequence pre-filtering
The first step of metagenomic data analysis requires the execution of certain pre-filtering steps, including the removal of redundant, low-quality sequences and sequences of probableeukaryotic
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bacte ...
origin (especially in metagenomes of human origin). The methods available for the removal of contaminating eukaryotic genomic DNA sequences include Eu-Detect and DeConseq.
Assembly
DNA sequence data from genomic and metagenomic projects are essentially the same, but genomic sequence data offers highercoverage
Coverage may refer to:
Filmmaking
* Coverage (lens), the size of the image a lens can produce
* Camera coverage, the amount of footage shot and different camera setups used in filming a scene
* Script coverage, a short summary of a script, writ ...
while metagenomic data is usually highly non-redundant. Furthermore, the increased use of second-generation sequencing technologies with short read lengths means that much of future metagenomic data will be error-prone. Taken in combination, these factors make the assembly of metagenomic sequence reads into genomes difficult and unreliable. Misassemblies are caused by the presence of repetitive DNA sequences that make assembly especially difficult because of the difference in the relative abundance of species present in the sample. Misassemblies can also involve the combination of sequences from more than one species into chimeric contig
A contig (from ''contiguous'') is a set of overlapping DNA segments that together represent a consensus region of DNA.Gregory, S. ''Contig Assembly''. Encyclopedia of Life Sciences, 2005.
In bottom-up sequencing projects, a contig refers to ov ...
s.
There are several assembly programs, most of which can use information from paired-end tag Paired-end tags (PET) (sometimes "Paired-End diTags", or simply "ditags") are the short sequences at the 5’ and 3' ends of a DNA fragment which are unique enough that they (theoretically) exist together only once in a genome, therefore making th ...
s in order to improve the accuracy of assemblies. Some programs, such as Phrap Phrap is a widely used program for DNA sequence assembly. It is part of the Phred-Phrap- Consed package.
History
Phrap was originally developed by Prof. Phil Green for the assembly of cosmids in large-scale cosmid shotgun sequencing within th ...
or Celera
Celera is a subsidiary of Quest Diagnostics which focuses on genetic sequencing and related technologies. It was founded in 1998 as a business unit of Applera, spun off into an independent company in 2008, and finally acquired by Quest Diagnostic ...
Assembler, were designed to be used to assemble single genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ...
s but nevertheless produce good results when assembling metagenomic data sets. Other programs, such as Velvet assembler
Velvet is an algorithm package that has been designed to deal with ''de novo'' genome assembly and short read sequencing alignments. This is achieved through the manipulation of de Bruijn graphs for genomic sequence assembly via the removal of er ...
, have been optimized for the shorter reads produced by second-generation sequencing through the use of de Bruijn graph
In graph theory, an -dimensional De Bruijn graph of symbols is a directed graph representing overlaps between sequences of symbols. It has vertices, consisting of all possible sequences of the given symbols; the same symbol may appear multipl ...
s. The use of reference genomes allows researchers to improve the assembly of the most abundant microbial species, but this approach is limited by the small subset of microbial phyla for which sequenced genomes are available. After an assembly is created, an additional challenge is "metagenomic deconvolution", or determining which sequences come from which species in the sample.
Gene prediction
Metagenomic analysis pipelines use two approaches in the annotation of coding regions in the assembled contigs. The first approach is to identify genes based upon homology with genes that are already publicly available insequence database
In the field of bioinformatics, a sequence database is a type of biological database that is composed of a large collection of computerized (" digital") nucleic acid sequences, protein sequences, or other polymer sequences stored on a computer. T ...
s, usually by BLAST
Blast or The Blast may refer to:
*Explosion, a rapid increase in volume and release of energy in an extreme manner
*Detonation, an exothermic front accelerating through a medium that eventually drives a shock front
Film
* ''Blast'' (1997 film), ...
searches. This type of approach is implemented in the program MEGAN
Megan is a Welsh feminine given name, originally a diminutive form of Margaret. Margaret is from the Greek μαργαρίτης (''margarítēs''), Latin ''margarīta'', "pearl". Megan is one of the most popular Welsh-language names for women in ...
4. The second, ''ab initio
''Ab initio'' ( ) is a Latin term meaning "from the beginning" and is derived from the Latin ''ab'' ("from") + ''initio'', ablative singular of ''initium'' ("beginning").
Etymology
Circa 1600, from Latin, literally "from the beginning", from ab ...
'', uses intrinsic features of the sequence to predict coding regions based upon gene training sets from related organisms. This is the approach taken by programs such as GeneMark
GeneMark is a generic name for a family of ab initio gene prediction programs developed at the Georgia Institute of Technology in Atlanta. Developed in 1993, original GeneMark was used in 1995 as a primary gene prediction tool for annotation of ...
and GLIMMER
In bioinformatics, GLIMMER (Gene Locator and Interpolated Markov ModelER) is used to find genes in prokaryotic DNA. "It is effective at finding genes in bacteria, archea, viruses, typically finding 98-99% of all relatively long protein coding g ...
. The main advantage of ''ab initio'' prediction is that it enables the detection of coding regions that lack homologs in the sequence databases; however, it is most accurate when there are large regions of contiguous genomic DNA available for comparison.
Species diversity
Gene annotations provide the "what", while measurements ofspecies diversity
Species diversity is the number of different species that are represented in a given community (a dataset). The effective number of species refers to the number of equally abundant species needed to obtain the same mean proportional species abundan ...
provide the "who". In order to connect community composition and function in metagenomes, sequences must be binned. Binning is the process of associating a particular sequence with an organism. In similarity-based binning, methods such as BLAST
Blast or The Blast may refer to:
*Explosion, a rapid increase in volume and release of energy in an extreme manner
*Detonation, an exothermic front accelerating through a medium that eventually drives a shock front
Film
* ''Blast'' (1997 film), ...
are used to rapidly search for phylogenetic markers or otherwise similar sequences in existing public databases. This approach is implemented in MEGAN
Megan is a Welsh feminine given name, originally a diminutive form of Margaret. Margaret is from the Greek μαργαρίτης (''margarítēs''), Latin ''margarīta'', "pearl". Megan is one of the most popular Welsh-language names for women in ...
. Another tool, PhymmBL, uses interpolated Markov models to assign reads.MetaPhlAnand
AMPHORA
An amphora (; grc, ἀμφορεύς, ''amphoreús''; English plural: amphorae or amphoras) is a type of container with a pointed bottom and characteristic shape and size which fit tightly (and therefore safely) against each other in storag ...
are methods based on unique clade-specific markers for estimating organismal relative abundances with improved computational performances. Other tools, likmOTUs
ref name="sunagawa2013"/> and MetaPhyler, use universal marker genes to profile prokaryotic species. With th
mOTUs profiler
is possible to profile species without a reference genome, improving the estimation of microbial community diversity. Recent methods, such a
SLIMM
use read coverage landscape of individual reference genomes to minimize false-positive hits and get reliable relative abundances. In composition based binning, methods use intrinsic features of the sequence, such as oligonucleotide frequencies or
codon usage bias
Codon usage bias refers to differences in the frequency of occurrence of synonymous codons in coding DNA. A codon is a series of three nucleotides (a triplet) that encodes a specific amino acid residue in a polypeptide chain or for the termination ...
. Once sequences are binned, it is possible to carry out comparative analysis of diversity and richness.
Data integration
The massive amount of exponentially growing sequence data is a daunting challenge that is complicated by the complexity of themetadata
Metadata is "data that provides information about other data", but not the content of the data, such as the text of a message or the image itself. There are many distinct types of metadata, including:
* Descriptive metadata – the descriptive ...
associated with metagenomic projects. Metadata includes detailed information about the three-dimensional (including depth, or height) geography and environmental features of the sample, physical data about the sample site, and the methodology of the sampling. This information is necessary both to ensure replicability
Reproducibility, also known as replicability and repeatability, is a major principle underpinning the scientific method. For the findings of a study to be reproducible means that results obtained by an experiment or an observational study or in a ...
and to enable downstream analysis. Because of its importance, metadata and collaborative data review and curation require standardized data formats located in specialized databases, such as the Genomes OnLine Database (GOLD).
Several tools have been developed to integrate metadata and sequence data, allowing downstream comparative analyses of different datasets using a number of ecological indices. In 2007, Folker Meyer and Robert Edwards and a team at Argonne National Laboratory
Argonne National Laboratory is a science and engineering research United States Department of Energy National Labs, national laboratory operated by University of Chicago, UChicago Argonne LLC for the United States Department of Energy. The facil ...
and the University of Chicago
The University of Chicago (UChicago, Chicago, U of C, or UChi) is a private research university in Chicago, Illinois. Its main campus is located in Chicago's Hyde Park neighborhood. The University of Chicago is consistently ranked among the b ...
released the Metagenomics Rapid Annotation using Subsystem Technology server (MG-RAST
MG-RAST is an open-source software, open-source web application server that suggests automatic phylogenetic and functional analysis of metagenomes. It is also one of the biggest repositories for metagenomic data. The name is an abbreviation of ''M ...
) a community resource for metagenome data set analysis. As of June 2012 over 14.8 terabases (14x1012 bases) of DNA have been analyzed, with more than 10,000 public data sets freely available for comparison within MG-RAST. Over 8,000 users now have submitted a total of 50,000 metagenomes to MG-RAST. The Integrated Microbial Genomes/Metagenomes (IMG/M) system also provides a collection of tools for functional analysis of microbial communities based on their metagenome sequence, based upon reference isolate genomes included from thIntegrated Microbial Genomes
(IMG) system and th
project. One of the first standalone tools for analysing high-throughput metagenome shotgun data was
MEGAN
Megan is a Welsh feminine given name, originally a diminutive form of Margaret. Margaret is from the Greek μαργαρίτης (''margarítēs''), Latin ''margarīta'', "pearl". Megan is one of the most popular Welsh-language names for women in ...
(MEta Genome ANalyzer). A first version of the program was used in 2005 to analyse the metagenomic context of DNA sequences obtained from a mammoth bone. Based on a BLAST comparison against a reference database, this tool performs both taxonomic and functional binning, by placing the reads onto the nodes of the NCBI taxonomy using a simple lowest common ancestor (LCA) algorithm or onto the nodes of thSEED
or
KEGG
KEGG (Kyoto Encyclopedia of Genes and Genomes) is a collection of databases dealing with genomes, biological pathways, diseases, drugs, and chemical substances. KEGG is utilized for bioinformatics research and education, including data analysis i ...
classifications, respectively.
With the advent of fast and inexpensive sequencing instruments, the growth of databases of DNA sequences is now exponential (e.g., the NCBI GenBank database ). Faster and efficient tools are needed to keep pace with the high-throughput sequencing, because the BLAST-based approaches such as MG-RAST or MEGAN run slowly to annotate large samples (e.g., several hours to process a small/medium size dataset/sample ). Thus, ultra-fast classifiers have recently emerged, thanks to more affordable powerful servers. These tools can perform the taxonomic annotation at extremely high speed, for example CLARK (according to CLARK's authors, it can classify accurately "32 million metagenomic short reads per minute"). At such a speed, a very large dataset/sample of a billion short reads can be processed in about 30 minutes.
With the increasing availability of samples containing ancient DNA and due to the uncertainty associated with the nature of those samples (ancient DNA damage), a fast tool capable of producing conservative similarity estimates has been made available. According to FALCON's authors, it can use relaxed thresholds and edit distances without affecting the memory and speed performance.
Comparative metagenomics
Comparative analyses between metagenomes can provide additional insight into the function of complex microbial communities and their role in host health. Pairwise or multiple comparisons between metagenomes can be made at the level of sequence composition (comparingGC-content
In molecular biology and genetics, GC-content (or guanine-cytosine content) is the percentage of nitrogenous bases in a DNA or RNA molecule that are either guanine (G) or cytosine (C). This measure indicates the proportion of G and C bases out o ...
or genome size), taxonomic diversity, or functional complement. Comparisons of population structure and phylogenetic diversity can be made on the basis of 16S rRNA 16S rRNA may refer to:
* 16S ribosomal RNA
16 S ribosomal RNA (or 16 S rRNA) is the RNA component of the 30S subunit of a prokaryotic ribosome ( SSU rRNA). It binds to the Shine-Dalgarno sequence and provides most of the SSU structure.
The g ...
and other phylogenetic marker genes, or—in the case of low-diversity communities—by genome reconstruction from the metagenomic dataset. Functional comparisons between metagenomes may be made by comparing sequences against reference databases such as COG or KEGG
KEGG (Kyoto Encyclopedia of Genes and Genomes) is a collection of databases dealing with genomes, biological pathways, diseases, drugs, and chemical substances. KEGG is utilized for bioinformatics research and education, including data analysis i ...
, and tabulating the abundance by category and evaluating any differences for statistical significance. This gene-centric approach emphasizes the functional complement of the ''community'' as a whole rather than taxonomic groups, and shows that the functional complements are analogous under similar environmental conditions. Consequently, metadata on the environmental context of the metagenomic sample is especially important in comparative analyses, as it provides researchers with the ability to study the effect of habitat upon community structure and function.
Additionally, several studies have also utilized oligonucleotide usage patterns to identify the differences across diverse microbial communities. Examples of such methodologies include the dinucleotide relative abundance approach by Willner et al. and the HabiSign approach of Ghosh et al. This latter study also indicated that differences in tetranucleotide usage patterns can be used to identify genes (or metagenomic reads) originating from specific habitats. Additionally some methods as TriageTools or Compareads detect similar reads between two read sets. The similarity measure
In statistics and related fields, a similarity measure or similarity function or similarity metric is a real-valued function that quantifies the similarity between two objects. Although no single definition of a similarity exists, usually such meas ...
they apply on reads is based on a number of identical words of length ''k'' shared by pairs of reads.
A key goal in comparative metagenomics is to identify microbial group(s) which are responsible for conferring specific characteristics to a given environment. However, due to issues in the sequencing technologies artifacts need to be accounted for like in metagenomeSeq. Others have characterized inter-microbial interactions between the resident microbial groups. A GUI
The GUI ( "UI" by itself is still usually pronounced . or ), graphical user interface, is a form of user interface that allows users to interact with electronic devices through graphical icons and audio indicator such as primary notation, inste ...
-based comparative metagenomic analysis application called Community-Analyzer has been developed by Kuntal et al.
which implements a correlation-based graph layout algorithm that not only facilitates a quick visualization of the differences in the analyzed microbial communities (in terms of their taxonomic composition), but also provides insights into the inherent inter-microbial interactions occurring therein. Notably, this layout algorithm also enables grouping of the metagenomes based on the probable inter-microbial interaction patterns rather than simply comparing abundance values of various taxonomic groups. In addition, the tool implements several interactive GUI-based functionalities that enable users to perform standard comparative analyses across microbiomes.
Data analysis
Community metabolism
In many bacterial communities, natural or engineered (such asbioreactor
A bioreactor refers to any manufactured device or system that supports a biologically active environment. In one case, a bioreactor is a vessel in which a chemical reaction, chemical process is carried out which involves organisms or biochemistry, ...
s), there is significant division of labor in metabolism (syntrophy In biology, syntrophy, synthrophy, or cross-feeding (from Greek ''syn'' meaning together, ''trophe'' meaning nourishment) is the phenomenon of one species feeding on the metabolic products of another species to cope up with the energy limitations b ...
), during which the waste products of some organisms are metabolites for others. In one such system, the methanogen
Methanogens are microorganisms that produce methane as a metabolic byproduct in hypoxic conditions. They are prokaryotic and belong to the domain Archaea. All known methanogens are members of the archaeal phylum Euryarchaeota. Methanogens are com ...
ic bioreactor, functional stability requires the presence of several syntrophic In biology, syntrophy, synthrophy, or cross-feeding (from Greek ''syn'' meaning together, ''trophe'' meaning nourishment) is the phenomenon of one species feeding on the metabolic products of another species to cope up with the energy limitations by ...
species ( Syntrophobacterales and Synergistia
The Synergistota is a phylum of anaerobic bacteria that show Gram-negative staining and have rod/vibrioid cell shape. Although Synergistota have a diderm cell envelope,Gupta, R. S. (2011) Origin of Diderm (Gram-negative) Bacteria: Antibiotic Sel ...
) working together in order to turn raw resources into fully metabolized waste (methane
Methane ( , ) is a chemical compound with the chemical formula (one carbon atom bonded to four hydrogen atoms). It is a group-14 hydride, the simplest alkane, and the main constituent of natural gas. The relative abundance of methane on Eart ...
). Using comparative gene studies and expression experiments with microarray
A microarray is a multiplex lab-on-a-chip. Its purpose is to simultaneously detect the expression of thousands of genes from a sample (e.g. from a tissue). It is a two-dimensional array on a solid substrate—usually a glass slide or silicon t ...
s or proteomics
Proteomics is the large-scale study of proteins. Proteins are vital parts of living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replication of DNA. In ...
researchers can piece together a metabolic network that goes beyond species boundaries. Such studies require detailed knowledge about which versions of which proteins are coded by which species and even by which strains of which species. Therefore, community genomic information is another fundamental tool (with metabolomics and proteomics) in the quest to determine how metabolites are transferred and transformed by a community.
Metatranscriptomics
Metagenomics allows researchers to access the functional and metabolic diversity of microbial communities, but it cannot show which of these processes are active. The extraction and analysis of metagenomicmRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
(the metatranscriptome) provides information on the regulation
Regulation is the management of complex systems according to a set of rules and trends. In systems theory, these types of rules exist in various fields of biology and society, but the term has slightly different meanings according to context. For ...
and expression
Expression may refer to:
Linguistics
* Expression (linguistics), a word, phrase, or sentence
* Fixed expression, a form of words with a specific meaning
* Idiom, a type of fixed expression
* Metaphorical expression, a particular word, phrase, o ...
profiles of complex communities. Because of the technical difficulties (the short half-life of mRNA, for example) in the collection of environmental RNA there have been relatively few ''in situ
''In situ'' (; often not italicized in English) is a Latin phrase that translates literally to "on site" or "in position." It can mean "locally", "on site", "on the premises", or "in place" to describe where an event takes place and is used in ...
'' metatranscriptomic studies of microbial communities to date. While originally limited to microarray
A microarray is a multiplex lab-on-a-chip. Its purpose is to simultaneously detect the expression of thousands of genes from a sample (e.g. from a tissue). It is a two-dimensional array on a solid substrate—usually a glass slide or silicon t ...
technology, metatranscriptomics studies have made use of transcriptomics technologies
Transcriptomics technologies are the techniques used to study an organism's transcriptome, the sum of all of its RNA transcripts. The information content of an organism is recorded in the DNA of its genome and expressed through transcription. ...
to measure whole-genome expression and quantification of a microbial community, first employed in analysis of ammonia oxidation in soils.
Viruses
Metagenomic sequencing is particularly useful in the study of viral communities. As viruses lack a shared universal phylogenetic marker (as16S RNA
16 S ribosomal RNA (or 16 S rRNA) is the RNA component of the 30S subunit of a prokaryotic ribosome (SSU rRNA). It binds to the Shine-Dalgarno sequence and provides most of the SSU structure.
The genes coding for it are referred to as 16S rR ...
for bacteria and archaea, and 18S RNA for eukarya), the only way to access the genetic diversity of the viral community from an environmental sample is through metagenomics. Viral metagenomes (also called viromes) should thus provide more and more information about viral diversity and evolution. For example, a metagenomic pipeline called Giant Virus Finder
The Giant Virus Finder is a free bioinformatics software for finding giant viruses in metagenomes.
Applications
The Giant Virus Finder tool integrates and applies thGiant Virus Toplist the list of the largest virus genomes. With the tool, gi ...
showed the first evidence of existence of giant viruses
A giant virus, sometimes referred to as a girus, is a very large virus, some of which are larger than typical bacteria. All known giant viruses belong to the phylum ''Nucleocytoviricota''.
Description
While the exact criteria as defined in the s ...
in a saline desert
and in Antarctic dry valleys.
Applications
Metagenomics has the potential to advance knowledge in a wide variety of fields. It can also be applied to solve practical challenges inmedicine
Medicine is the science and practice of caring for a patient, managing the diagnosis, prognosis, prevention, treatment, palliation of their injury or disease, and promoting their health. Medicine encompasses a variety of health care pract ...
, engineering
Engineering is the use of scientific method, scientific principles to design and build machines, structures, and other items, including bridges, tunnels, roads, vehicles, and buildings. The discipline of engineering encompasses a broad rang ...
, agriculture
Agriculture or farming is the practice of cultivating plants and livestock. Agriculture was the key development in the rise of sedentary human civilization, whereby farming of domesticated species created food surpluses that enabled people to ...
, sustainability
Specific definitions of sustainability are difficult to agree on and have varied in the literature and over time. The concept of sustainability can be used to guide decisions at the global, national, and individual levels (e.g. sustainable livi ...
and ecology
Ecology () is the study of the relationships between living organisms, including humans, and their physical environment. Ecology considers organisms at the individual, population, community, ecosystem, and biosphere level. Ecology overlaps wi ...
.
Agriculture
Thesoil
Soil, also commonly referred to as earth or dirt, is a mixture of organic matter, minerals, gases, liquids, and organisms that together support life. Some scientific definitions distinguish ''dirt'' from ''soil'' by restricting the former te ...
s in which plants grow are inhabited by microbial communities, with one gram of soil containing around 109-1010 microbial cells which comprise about one gigabase of sequence information. The microbial communities which inhabit soils are some of the most complex known to science, and remain poorly understood despite their economic importance. Microbial consortia perform a wide variety of ecosystem service
Ecosystem services are the many and varied benefits to humans provided by the natural environment and healthy ecosystems. Such ecosystems include, for example, agroecosystems, forest ecosystem, grassland ecosystems, and aquatic ecosystems. Th ...
s necessary for plant growth, including fixing atmospheric nitrogen, nutrient cycling
A nutrient cycle (or ecological recycling) is the movement and exchange of inorganic and organic matter back into the production of matter. Energy flow is a unidirectional and noncyclic pathway, whereas the movement of mineral nutrients is cycli ...
, disease suppression, and sequester
Sequestration or sequester may refer to:
Law and government
* Sequestration (law), the seizure of property for creditors or the state. See also Committee for Compounding with Delinquents
* Jury sequestration, the isolation of a jury
* Bankruptcy, ...
iron
Iron () is a chemical element with symbol Fe (from la, ferrum) and atomic number 26. It is a metal that belongs to the first transition series and group 8 of the periodic table. It is, by mass, the most common element on Earth, right in f ...
and other metal
A metal (from Greek μέταλλον ''métallon'', "mine, quarry, metal") is a material that, when freshly prepared, polished, or fractured, shows a lustrous appearance, and conducts electricity and heat relatively well. Metals are typicall ...
s. Functional metagenomics strategies are being used to explore the interactions between plants and microbes through cultivation-independent study of these microbial communities. By allowing insights into the role of previously uncultivated or rare community members in nutrient cycling and the promotion of plant growth, metagenomic approaches can contribute to improved disease detection in crop
A crop is a plant that can be grown and harvested extensively for profit or subsistence. When the plants of the same kind are cultivated at one place on a large scale, it is called a crop. Most crops are cultivated in agriculture or hydroponic ...
s and livestock
Livestock are the domesticated animals raised in an agricultural setting to provide labor and produce diversified products for consumption such as meat, eggs, milk, fur, leather, and wool. The term is sometimes used to refer solely to animals ...
and the adaptation of enhanced farming
Agriculture or farming is the practice of cultivating plants and livestock. Agriculture was the key development in the rise of sedentary human civilization, whereby farming of domesticated species created food surpluses that enabled people to ...
practices which improve crop health by harnessing the relationship between microbes and plants.
Biofuel
Biofuel
Biofuel is a fuel that is produced over a short time span from biomass, rather than by the very slow natural processes involved in the formation of fossil fuels, such as oil. According to the United States Energy Information Administration (E ...
s are fuel
A fuel is any material that can be made to react with other substances so that it releases energy as thermal energy or to be used for work. The concept was originally applied solely to those materials capable of releasing chemical energy but ...
s derived from biomass
Biomass is plant-based material used as a fuel for heat or electricity production. It can be in the form of wood, wood residues, energy crops, agricultural residues, and waste from industry, farms, and households. Some people use the terms bi ...
conversion, as in the conversion of cellulose
Cellulose is an organic compound with the formula , a polysaccharide consisting of a linear chain of several hundred to many thousands of β(1→4) linked D-glucose units. Cellulose is an important structural component of the primary cell wall ...
contained in corn
Maize ( ; ''Zea mays'' subsp. ''mays'', from es, maíz after tnq, mahiz), also known as corn (North American and Australian English), is a cereal grain first domesticated by indigenous peoples in southern Mexico about 10,000 years ago. Th ...
stalks, switchgrass
''Panicum virgatum'', commonly known as switchgrass, is a perennial warm season bunchgrass native to North America, where it occurs naturally from 55°N latitude in Canada southwards into the United States and Mexico. Switchgrass is one of the ...
, and other biomass into cellulosic ethanol
Cellulosic ethanol is ethanol (ethyl alcohol) produced from cellulose (the stringy fiber of a plant) rather than from the plant's seeds or fruit. It can be produced from grasses, wood, algae, or other plants. It is generally discussed for use as a ...
. This process is dependent upon microbial consortia(association) that transform the cellulose into sugar
Sugar is the generic name for sweet-tasting, soluble carbohydrates, many of which are used in food. Simple sugars, also called monosaccharides, include glucose, fructose, and galactose. Compound sugars, also called disaccharides or double ...
s, followed by the fermentation
Fermentation is a metabolic process that produces chemical changes in organic substrates through the action of enzymes. In biochemistry, it is narrowly defined as the extraction of energy from carbohydrates in the absence of oxygen. In food ...
of the sugars into ethanol
Ethanol (abbr. EtOH; also called ethyl alcohol, grain alcohol, drinking alcohol, or simply alcohol) is an organic compound. It is an Alcohol (chemistry), alcohol with the chemical formula . Its formula can be also written as or (an ethyl ...
. Microbes also produce a variety of sources of bioenergy
Biomass is plant-based material used as a fuel for heat or electricity production. It can be in the form of wood, wood residues, energy crops, agricultural residues, and waste from industry, farms, and households. Some people use the terms biom ...
including methane
Methane ( , ) is a chemical compound with the chemical formula (one carbon atom bonded to four hydrogen atoms). It is a group-14 hydride, the simplest alkane, and the main constituent of natural gas. The relative abundance of methane on Eart ...
and hydrogen
Hydrogen is the chemical element with the symbol H and atomic number 1. Hydrogen is the lightest element. At standard conditions hydrogen is a gas of diatomic molecules having the formula . It is colorless, odorless, tasteless, non-toxic, an ...
.
The efficient industrial-scale deconstruction of biomass requires novel enzymes
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different molecule ...
with higher productivity and lower cost. Metagenomic approaches to the analysis of complex microbial communities allow the targeted screening
Screening may refer to:
* Screening cultures, a type a medical test that is done to find an infection
* Screening (economics), a strategy of combating adverse selection (includes sorting resumes to select employees)
* Screening (environmental), a ...
of enzymes
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different molecule ...
with industrial applications in biofuel production, such as glycoside hydrolase
Glycoside hydrolases (also called glycosidases or glycosyl hydrolases) catalyze the hydrolysis of glycosidic bonds in complex sugars. They are extremely common enzymes with roles in nature including degradation of biomass such as cellulose (ce ...
s. Furthermore, knowledge of how these microbial communities function is required to control them, and metagenomics is a key tool in their understanding. Metagenomic approaches allow comparative analyses between convergent microbial systems like biogas
Biogas is a mixture of gases, primarily consisting of methane, carbon dioxide and hydrogen sulphide, produced from raw materials such as agricultural waste, manure, municipal waste, plant material, sewage, green waste and food waste. It is a ...
fermenters or insect
Insects (from Latin ') are pancrustacean hexapod invertebrates of the class Insecta. They are the largest group within the arthropod phylum. Insects have a chitinous exoskeleton, a three-part body ( head, thorax and abdomen), three pairs ...
herbivore
A herbivore is an animal anatomically and physiologically adapted to eating plant material, for example foliage or marine algae, for the main component of its diet. As a result of their plant diet, herbivorous animals typically have mouthpart ...
s such as the fungus garden of the leafcutter ant
Leafcutter ants, a non-generic name, are any of 47 species of leaf-chewing ants belonging to the two genera '' Atta'' and ''Acromyrmex''.
These species of tropical, fungus-growing ants are all endemic to South and Central America, Mexico, and par ...
s.
Biotechnology
Microbial communities produce a vast array of biologically active chemicals that are used in competition and communication. Many of the drugs in use today were originally uncovered in microbes; recent progress in mining the rich genetic resource of non-culturable microbes has led to the discovery of new genes, enzymes, and natural products. The application of metagenomics has allowed the development ofcommodity
In economics, a commodity is an economic good, usually a resource, that has full or substantial fungibility: that is, the market treats instances of the good as equivalent or nearly so with no regard to who produced them.
The price of a comm ...
and fine chemicals
In chemistry, fine chemicals are complex, single, pure chemical substances, produced in limited quantities in multipurpose plants by multistep batch chemical or biotechnological processes. They are described by exacting specifications, used fo ...
, agrochemical
An agrochemical or agrichemical, a contraction of ''agricultural chemical'', is a chemical product used in industrial agriculture. Agrichemical refers to biocides ( pesticides including insecticides, herbicides, fungicides and nematicides) an ...
s and pharmaceuticals
A medication (also called medicament, medicine, pharmaceutical drug, medicinal drug or simply drug) is a drug used to diagnose, cure, treat, or prevent disease. Drug therapy (pharmacotherapy) is an important part of the medical field and rel ...
where the benefit of enzyme-catalyzed chiral synthesis
Enantioselective synthesis, also called asymmetric synthesis, is a form of chemical synthesis. It is defined by IUPAC as "a chemical reaction (or reaction sequence) in which one or more new elements of chirality are formed in a substrate molecul ...
is increasingly recognized.
Two types of analysis are used in the bioprospecting
Bioprospecting (also known as biodiversity prospecting) is the exploration of natural sources for small molecules, macromolecules and biochemical and genetic information that could be developed into commercialization, commercially valuable prod ...
of metagenomic data: function-driven screening for an expressed trait, and sequence-driven screening for DNA sequences of interest. Function-driven analysis seeks to identify clones expressing a desired trait or useful activity, followed by biochemical characterization and sequence analysis. This approach is limited by availability of a suitable screen and the requirement that the desired trait be expressed in the host cell. Moreover, the low rate of discovery (less than one per 1,000 clones screened) and its labor-intensive nature further limit this approach. In contrast, sequence-driven analysis uses conserved DNA sequences to design PCR primers to screen clones for the sequence of interest. In comparison to cloning-based approaches, using a sequence-only approach further reduces the amount of bench work required. The application of massively parallel sequencing also greatly increases the amount of sequence data generated, which require high-throughput bioinformatic analysis pipelines. The sequence-driven approach to screening is limited by the breadth and accuracy of gene functions present in public sequence databases. In practice, experiments make use of a combination of both functional and sequence-based approaches based upon the function of interest, the complexity of the sample to be screened, and other factors. An example of success using metagenomics as a biotechnology for drug discovery is illustrated with the malacidin antibiotics.
Ecology
 Metagenomics can provide valuable insights into the functional ecology of environmental communities. Metagenomic analysis of the bacterial consortia found in the defecations of Australian sea lions suggests that nutrient-rich sea lion faeces may be an important nutrient source for coastal ecosystems. This is because the bacteria that are expelled simultaneously with the defecations are adept at breaking down the nutrients in the faeces into a bioavailable form that can be taken up into the food chain.
DNA sequencing can also be used more broadly to identify species present in a body of water, debris filtered from the air, sample of dirt, or animal's faeces, and even detect diet items from blood meals. This can establish the range of
Metagenomics can provide valuable insights into the functional ecology of environmental communities. Metagenomic analysis of the bacterial consortia found in the defecations of Australian sea lions suggests that nutrient-rich sea lion faeces may be an important nutrient source for coastal ecosystems. This is because the bacteria that are expelled simultaneously with the defecations are adept at breaking down the nutrients in the faeces into a bioavailable form that can be taken up into the food chain.
DNA sequencing can also be used more broadly to identify species present in a body of water, debris filtered from the air, sample of dirt, or animal's faeces, and even detect diet items from blood meals. This can establish the range of invasive species
An invasive species otherwise known as an alien is an introduced organism that becomes overpopulated and harms its new environment. Although most introduced species are neutral or beneficial with respect to other species, invasive species ad ...
and endangered species
An endangered species is a species that is very likely to become extinct in the near future, either worldwide or in a particular political jurisdiction. Endangered species may be at risk due to factors such as habitat loss, poaching and inv ...
, and track seasonal populations.
Environmental remediation
Metagenomics can improve strategies for monitoring the impact ofpollutant
A pollutant or novel entity is a substance or energy introduced into the environment that has undesired effects, or adversely affects the usefulness of a resource. These can be both naturally forming (i.e. minerals or extracted compounds like oi ...
s on ecosystem
An ecosystem (or ecological system) consists of all the organisms and the physical environment with which they interact. These biotic and abiotic components are linked together through nutrient cycles and energy flows. Energy enters the syste ...
s and for cleaning up contaminated environments. Increased understanding of how microbial communities cope with pollutants improves assessments of the potential of contaminated sites to recover from pollution and increases the chances of bioaugmentation
Biological augmentation is the addition of archaea or bacterial cultures required to speed up the rate of degradation of a contaminant. Organisms that originate from contaminated areas may already be able to break down waste, but perhaps inefficien ...
or biostimulation
Biostimulation involves the modification of the environment to stimulate existing bacteria capable of bioremediation. This can be done by addition of various forms of rate limiting nutrients and electron acceptors, such as phosphorus, nitrogen, ...
trials to succeed.
Gut microbe characterization
Microbial communities Microbial population biology is the application of the principles of population biology to microorganisms.
Distinguishing from other biological disciplines
Microbial population biology, in practice, is the application of population ecology and popu ...
play a key role in preserving human health
Health, according to the World Health Organization, is "a state of complete physical, mental and social well-being and not merely the absence of disease and infirmity".World Health Organization. (2006)''Constitution of the World Health Organiza ...
, but their composition and the mechanism by which they do so remains mysterious. Metagenomic sequencing is being used to characterize the microbial communities from 15–18 body sites from at least 250 individuals. This is part of the Human Microbiome initiative with primary goals to determine if there is a core human microbiome
The human microbiome is the aggregate of all microbiota that reside on or within human tissues and biofluids along with the corresponding anatomical sites in which they reside, including the skin, mammary glands, seminal fluid, uterus, ovarian ...
, to understand the changes in the human microbiome that can be correlated with human health, and to develop new technological and bioinformatics
Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combi ...
tools to support these goals.
Another medical study as part of the MetaHit (Metagenomics of the Human Intestinal Tract) project consisted of 124 individuals from Denmark and Spain consisting of healthy, overweight, and irritable bowel disease patients. The study attempted to categorize the depth and phylogenetic diversity of gastrointestinal bacteria. Using Illumina GA sequence data and SOAPdenovo, a de Bruijn graph-based tool specifically designed for assembly short reads, they were able to generate 6.58 million contigs greater than 500 bp for a total contig length of 10.3 Gb and a N50 length of 2.2 kb.
The study demonstrated that two bacterial divisions, Bacteroidetes and Firmicutes, constitute over 90% of the known phylogenetic categories that dominate distal gut bacteria. Using the relative gene frequencies found within the gut these researchers identified 1,244 metagenomic clusters that are critically important for the health of the intestinal tract. There are two types of functions in these range clusters: housekeeping and those specific to the intestine. The housekeeping gene clusters are required in all bacteria and are often major players in the main metabolic pathways including central carbon metabolism and amino acid synthesis. The gut-specific functions include adhesion to host proteins and the harvesting of sugars from globoseries glycolipids. Patients with irritable bowel syndrome were shown to exhibit 25% fewer genes and lower bacterial diversity than individuals not suffering from irritable bowel syndrome indicating that changes in patients' gut biome diversity may be associated with this condition.
While these studies highlight some potentially valuable medical applications, only 31–48.8% of the reads could be aligned to 194 public human gut bacterial genomes and 7.6–21.2% to bacterial genomes available in GenBank which indicates that there is still far more research necessary to capture novel bacterial genomes.
In the Human Microbiome Project
The Human Microbiome Project (HMP) was a United States National Institutes of Health (NIH) research initiative to improve understanding of the microbiota involved in human health and disease. Launched in 2007, the first phase (HMP1) focused on i ...
(HMP), gut microbial communities were assayed using high-throughput DNA sequencing. HMP showed that, unlike individual microbial species, many metabolic processes were present among all body habitats with varying frequencies. Microbial communities of 649 metagenomes drawn from seven primary body sites on 102 individuals were studied as part of the human microbiome
The human microbiome is the aggregate of all microbiota that reside on or within human tissues and biofluids along with the corresponding anatomical sites in which they reside, including the skin, mammary glands, seminal fluid, uterus, ovarian ...
project. The metagenomic analysis revealed variations in niche specific abundance among 168 functional modules and 196 metabolic pathways within the microbiome. These included glycosaminoglycan degradation in the gut, as well as phosphate and amino acid transport linked to host phenotype (vaginal pH) in the posterior fornix. The HMP has brought to light the utility of metagenomics in diagnostics and evidence-based medicine
Evidence-based medicine (EBM) is "the conscientious, explicit and judicious use of current best evidence in making decisions about the care of individual patients". The aim of EBM is to integrate the experience of the clinician, the values of t ...
. Thus metagenomics is a powerful tool to address many of the pressing issues in the field of Personalized medicine
Personalized medicine, also referred to as precision medicine, is a medical model that separates people into different groups—with medical decisions, practices, interventions and/or products being tailored to the individual patient based on the ...
.
In animals, metagenomics can be used to profile their gut microbiomes and enable detection of antibiotic-resistant bacteria. This can have implications in monitoring the spread of diseases from wildlife to farmed animals and humans.
Infectious disease diagnosis
Differentiating between infectious and non-infectious illness, and identifying the underlying etiology of infection, can be challenging. For example, more than half of cases ofencephalitis
Encephalitis is inflammation of the brain. The severity can be variable with symptoms including reduction or alteration in consciousness, headache, fever, confusion, a stiff neck, and vomiting. Complications may include seizures, hallucinations, ...
remain undiagnosed, despite extensive testing using state-of-the-art clinical laboratory methods. Clinical metagenomic sequencing shows promise as a sensitive and rapid method to diagnose infection by comparing genetic material found in a patient's sample to databases of all known microscopic human pathogens and thousands of other bacterial, viral, fungal, and parasitic organisms and databases on antimicrobial resistances gene sequences with associated clinical phenotypes.
Arbovirus surveillance
Metagenomics has been an invaluable tool to help characterise the diversity and ecology of pathogens that are vectored by hematophagous (blood-feeding) insects such as mosquitoes and ticks. Metagenomics is routinely used by public health officials and organisations for the surveillance of arboviruses.See also
* Binning * Epidemiology and sewage *Metaproteomics Metaproteomics (also Community Proteomics, Environmental Proteomics, or Community Proteogenomics) is an umbrella term for experimental approaches to study all proteins in microbial communities and microbiomes from environmental sources. Metaproteomi ...
*Microbial ecology
Microbial ecology (or environmental microbiology) is the ecology of microorganisms: their relationship with one another and with their environment. It concerns the three major domains of life—Eukaryota, Archaea, and Bacteria—as well as viru ...
*Pathogenomics Pathogenomics is a field which uses high-throughput screening technology and bioinformatics to study encoded microbe resistance, as well as virulence factors (VFs), which enable a microorganism to infect a host and possibly cause disease. This inclu ...
References
External links
Focus on Metagenomics
at ''
Nature Reviews Microbiology
''Nature Reviews Microbiology'' is a monthly peer-reviewed review journal published by Nature Portfolio. It was established in 2003. The journal publishes reviews and perspectives on microbiology, bridging fundamental research and its clinical, in ...
'' journal websiteThe “Critical Assessment of Metagenome Interpretation” (CAMI) initiative
to evaluate methods in metagenomics {{Portal bar, Biology, Medicine Bioinformatics Genomics Environmental microbiology Microbiology techniques