|
Contig
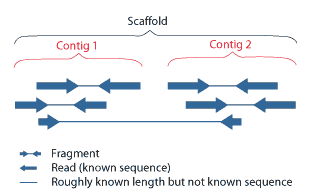
A contig (from ''contiguous'') is a set of overlapping DNA segments that together represent a consensus region of DNA.Gregory, S. ''Contig Assembly''. Encyclopedia of Life Sciences, 2005. In bottom-up sequencing projects, a contig refers to overlapping sequence data ( reads); in top-down sequencing projects, contig refers to the overlapping clones that form a physical map of the genome that is used to guide sequencing and assembly.Dear, P. H. ''Genome Mapping''. Encyclopedia of Life Sciences, 2005. . Contigs can thus refer both to overlapping DNA sequences and to overlapping physical segments (fragments) contained in clones depending on the context. Original definition of contig In 1980, Staden wrote: ''In order to make it easier to talk about our data gained by the shotgun method of sequencing we have invented the word "contig". A contig is a set of gel readings that are related to one another by overlap of their sequences. All gel readings belong to one and only one cont ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Scaffolding (bioinformatics)
Scaffolding is a technique used in bioinformatics. It is defined as follows: Link together a non-contiguous series of genomic sequences into a scaffold, consisting of sequences separated by gaps of known length. The sequences that are linked are typically contiguous sequences corresponding to read overlaps. When creating a draft genome, individual reads of DNA are second assembled into contigs, which, by the nature of their assembly, have gaps between them. The next step is to then bridge the gaps between these contigs to create a scaffold. This can be done using either optical mapping or mate-pair sequencing. Assembly software The sequencing of the ''Haemophilus influenzae'' genome marked the advent of scaffolding. That project generated a total of 140 contigs, which were oriented and linked using paired-end reads. The success of this strategy prompted The Institute for Genomic Research to develop the scaffolding program Grouper for their other sequencing projects. Until 200 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Shotgun Sequencing
In genetics, shotgun sequencing is a method used for sequencing random DNA strands. It is named by analogy with the rapidly expanding, quasi-random shot grouping of a shotgun. The Sanger sequencing#Method, chain-termination method of DNA sequencing ("Sanger sequencing") can only be used for short DNA strands of 100 to 1000 base pairs. Due to this size limit, longer sequences are subdivided into smaller fragments that can be sequenced separately, and these sequences are sequence assembly, assembled to give the overall sequence. In shotgun sequencing, DNA is broken up randomly into numerous small segments, which are sequenced using the chain termination method to obtain ''reads''. Multiple overlapping reads for the target DNA are obtained by performing several rounds of this fragmentation and sequencing. Computer programs then use the overlapping ends of different reads to assemble them into a continuous sequence. Shotgun sequencing was one of the precursor technologies that was resp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Sequencing Theory
DNA sequencing theory is the broad body of work that attempts to lay analytical foundations for determining the order of specific nucleotides in a sequence of DNA, otherwise known as DNA sequencing. The practical aspects revolve around designing and optimizing sequencing projects (known as "strategic genomics"), predicting project performance, troubleshooting experimental results, characterizing factors such as sequence bias and the effects of software processing algorithms, and comparing various sequencing methods to one another. In this sense, it could be considered a branch of systems engineering or operations research. The permanent archive of work is primarily mathematical, although numerical calculations are often conducted for particular problems too. DNA sequencing theory addresses ''physical processes'' related to sequencing DNA and should not be confused with theories of analyzing resultant DNA sequences, e.g. sequence alignment. Publications sometimes do not make a careful ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Assembly
In bioinformatics, sequence assembly refers to aligning and merging fragments from a longer DNA sequence in order to reconstruct the original sequence. This is needed as DNA sequencing technology might not be able to 'read' whole genomes in one go, but rather reads small pieces of between 20 and 30,000 bases, depending on the technology used. Typically, the short fragments (reads) result from shotgun sequencing genomic DNA, or gene transcript ( ESTs). The problem of sequence assembly can be compared to taking many copies of a book, passing each of them through a shredder with a different cutter, and piecing the text of the book back together just by looking at the shredded pieces. Besides the obvious difficulty of this task, there are some extra practical issues: the original may have many repeated paragraphs, and some shreds may be modified during shredding to have typos. Excerpts from another book may also be added in, and some shreds may be completely unrecognizable. Types ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Mapping
Gene mapping or genome mapping describes the methods used to identify the location of a gene on a chromosome and the distances between genes. Gene mapping can also describe the distances between different sites within a gene. The essence of all genome mapping is to place a collection of molecular markers onto their respective positions on the genome. Molecular markers come in all forms. Genes can be viewed as one special type of genetic markers in the construction of genome maps, and mapped the same way as any other markers. In some areas of study, gene mapping contributes to the creation of new recombinants within an organism. Gene maps help describe the spatial arrangement of genes on a chromosome. Genes are designated to a specific location on a chromosome known as the locus and can be used as molecular markers to find the distance between other genes on a chromosome. Maps provide researchers with the opportunity to predict the inheritance patterns of specific traits, which ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Sequencing
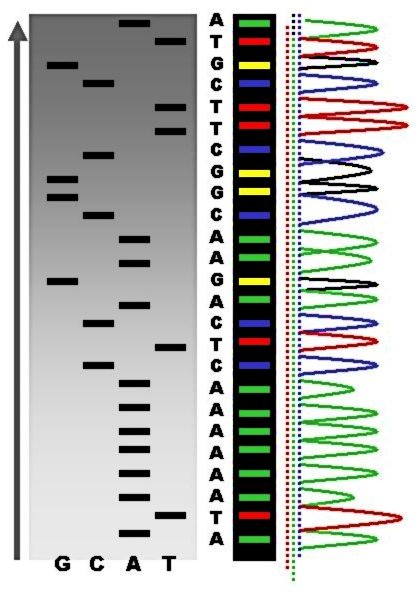
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, thymine, cytosine, and guanine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, Genographic Project, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid advancements in DNA seque ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Primer Walking
Primer walking is a technique used to clone a gene (e.g., disease gene) from its known closest markers (e.g., known gene). As a result, it is employed in cloning and sequencing efforts in plants, fungi, and mammals with minor alterations. This technique, also known as "directed sequencing," employs a series of Sanger sequencing reactions to either confirm the reference sequence of a known plasmid or PCR product based on the reference sequence (sequence confirmation service) or to discover the unknown sequence of a full plasmid or PCR product by designing primers to sequence overlapping sections (sequence discovery service). Primer walking: a DNA sequencing method Primer walking is a method to determine the sequence of DNA up to the 1.3–7.0 kb range whereas chromosome walking is used to produce the clones of already known sequences of the gene. Too long fragments cannot be sequenced in a single sequence read using the chain termination method. This method works by dividing th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Read (biology)
In DNA sequencing, a read is an inferred sequence of base pairs (or base pair probabilities) corresponding to all or part of a single DNA fragment. A typical sequencing experiment involves DNA fragmentation, fragmentation of the genome into millions of molecules, which are size-selected and ligation (molecular biology), ligated to adapter (genetics), adapters. The set of fragments is referred to as a sequencing library, which is sequenced to produce a set of reads. Read length Sequencing technologies vary in the length of reads produced. Reads of length 20-40 base pairs (bp) are referred to as ultra-short. Typical sequencers produce read lengths in the range of 100-500 bp. However, Pacific Biosciences platforms produce read lengths of approximately 1500 bp. Read length is a factor which can affect the results of biological studies. For example, longer read lengths improve the resolution of ''de novo'' genome assembly and detection of structural variants. It is estimated that read ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Staden Package
The Staden Package is computer software, a set of tools for DNA sequence assembly, editing, and sequence analysis. It is open-source software, released under a BSD 3-clause license. Package components The Staden package consists of several different programs. The main components are: * pregap4 – base calling with Phred, end clipping, and vector trimming * trev – trace viewing and editing * gap4 – sequence assembly, contig editing, and finishing * gap5 – assembly visualising, editing, and finishing of NGS data * Spin – DNA and protein sequence analysis History The Staden Package was developed by Rodger Staden's group at the Medical Research Council (MRC) Laboratory of Molecular Biology, Cambridge, England, since 1977. The package was available free to academic users, with 2,500 licenses issued in 2003 and an estimated 10,000 users, when funding for further development ended. The package was converted to open-source in 2004, and several new versions have been rele ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome Project
Genome projects are scientific endeavours that ultimately aim to determine the complete genome sequence of an organism (be it an animal, a plant, a fungus, a bacterium, an archaean, a protist or a virus) and to annotate protein-coding genes and other important genome-encoded features. The genome sequence of an organism includes the collective DNA sequences of each chromosome in the organism. For a bacterium containing a single chromosome, a genome project will aim to map the sequence of that chromosome. For the human species, whose genome includes 22 pairs of autosomes and 2 sex chromosomes, a complete genome sequence will involve 46 separate chromosome sequences. The Human Genome Project is a well known example of a genome project. Genome assembly Genome assembly refers to the process of taking a large number of short DNA sequences and reassembling them to create a representation of the original chromosomes from which the DNA originated. In a shotgun sequencing project, all th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacterial Artificial Chromosome
A bacterial artificial chromosome (BAC) is a DNA construct, based on a functional fertility plasmid (or F-plasmid), used for transforming and cloning in bacteria, usually '' E. coli''. F-plasmids play a crucial role because they contain partition genes that promote the even distribution of plasmids after bacterial cell division. The bacterial artificial chromosome's usual insert size is 150–350 kbp. A similar cloning vector called a PAC has also been produced from the DNA of P1 bacteriophage. BACs were often used to sequence the genomes of organisms in genome projects, for example the Human Genome Project, though they have been replaced by more modern technologies. In BAC sequencing, short piece of the organism's DNA is amplified as an insert in BACs, and then sequenced. Finally, the sequenced parts are rearranged '' in silico'', resulting in the genomic sequence of the organism. BACs were replaced with faster and less laborious sequencing methods like whole genome shot ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |