Archaea on:
[Wikipedia]
[Google]
[Amazon]
Archaea ( ; singular archaeon ) is a
 For much of the 20th century, prokaryotes were regarded as a single group of organisms and classified based on their
For much of the 20th century, prokaryotes were regarded as a single group of organisms and classified based on their
 The classification of archaea, and of prokaryotes in general, is a rapidly moving and contentious field. Current classification systems aim to organize archaea into groups of organisms that share structural features and common ancestors. These classifications rely heavily on the use of the sequence of
The classification of archaea, and of prokaryotes in general, is a rapidly moving and contentious field. Current classification systems aim to organize archaea into groups of organisms that share structural features and common ancestors. These classifications rely heavily on the use of the sequence of
 Archaeal membranes are made of molecules that are distinctly different from those in all other life forms, showing that archaea are related only distantly to bacteria and eukaryotes. In all organisms,
Archaeal membranes are made of molecules that are distinctly different from those in all other life forms, showing that archaea are related only distantly to bacteria and eukaryotes. In all organisms,
 Other archaea use in the atmosphere as a source of carbon, in a process called carbon fixation (they are
Other archaea use in the atmosphere as a source of carbon, in a process called carbon fixation (they are
 Archaea are genetically distinct from bacteria and eukaryotes, with up to 15% of the proteins encoded by any one archaeal genome being unique to the domain, although most of these unique genes have no known function. Of the remainder of the unique proteins that have an identified function, most belong to the Euryarchaeota and are involved in methanogenesis. The proteins that archaea, bacteria and eukaryotes share form a common core of cell function, relating mostly to
Archaea are genetically distinct from bacteria and eukaryotes, with up to 15% of the proteins encoded by any one archaeal genome being unique to the domain, although most of these unique genes have no known function. Of the remainder of the unique proteins that have an identified function, most belong to the Euryarchaeota and are involved in methanogenesis. The proteins that archaea, bacteria and eukaryotes share form a common core of cell function, relating mostly to
 Archaea exist in a broad range of
Archaea exist in a broad range of
 The well-characterized interactions between archaea and other organisms are either mutual or
The well-characterized interactions between archaea and other organisms are either mutual or
Introduction to the Archaea, ecology, systematics and morphology
Oceans of Archaea
nbsp;– E.F. DeLong, ''ASM News'', 2003
NCBI taxonomy page on Archaea
nbsp;– list of Prokaryotic names with Standing in Nomenclature
Shotgun sequencing finds nanoorganisms
nbsp;– discovery of the ARMAN group of archaea
Browse any completed archaeal genome at UCSC
Comparative Analysis of Archaeal Genomes
(at DOE's IMG system) {{Authority control Extremophiles Domains (biology) Systems of bacterial taxonomy
domain
Domain may refer to:
Mathematics
*Domain of a function, the set of input values for which the (total) function is defined
**Domain of definition of a partial function
**Natural domain of a partial function
**Domain of holomorphy of a function
* Do ...
of single-celled organisms. These microorganism
A microorganism, or microbe,, ''mikros'', "small") and ''organism'' from the el, ὀργανισμός, ''organismós'', "organism"). It is usually written as a single word but is sometimes hyphenated (''micro-organism''), especially in olde ...
s lack cell nuclei
The cell nucleus (pl. nuclei; from Latin or , meaning ''kernel'' or ''seed'') is a membrane-bound organelle found in eukaryotic cells. Eukaryotic cells usually have a single nucleus, but a few cell types, such as mammalian red blood cells, ha ...
and are therefore prokaryotes. Archaea were initially classified as bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
, receiving the name archaebacteria (in the Archaebacteria kingdom
Kingdom commonly refers to:
* A monarchy ruled by a king or queen
* Kingdom (biology), a category in biological taxonomy
Kingdom may also refer to:
Arts and media Television
* ''Kingdom'' (British TV series), a 2007 British television drama s ...
), but this term has fallen out of use.
Archaeal cells have unique properties separating them from the other two domains, Bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
and Eukaryota. Archaea are further divided into multiple recognized phyla. Classification is difficult because most have not been isolated in a laboratory and have been detected only by their gene sequences in environmental samples.
Archaea and bacteria are generally similar in size and shape, although a few archaea have very different shapes, such as the flat, square cells of ''Haloquadratum walsbyi
''Haloquadratum walsbyi'' is of the genus ''Haloquadratum,'' within the archaea domain known for its square halophilic nature. First discovered in a brine pool in the Sinai peninsula of Egypt, ''H. walsbyi'' is noted for its flat, square-shaped ...
''. Despite this morphological similarity to bacteria, archaea possess gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a ba ...
s and several metabolic pathway
In biochemistry, a metabolic pathway is a linked series of chemical reactions occurring within a cell. The reactants, products, and intermediates of an enzymatic reaction are known as metabolites, which are modified by a sequence of chemical reac ...
s that are more closely related to those of eukaryotes, notably for the enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. A ...
s involved in transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
and translation
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transla ...
. Other aspects of archaeal biochemistry are unique, such as their reliance on ether lipid
In organic chemistry, ethers are a class of compounds that contain an ether group—an oxygen atom connected to two alkyl or aryl groups. They have the general formula , where R and R′ represent the alkyl or aryl groups. Ethers can again be ...
s in their cell membrane
The cell membrane (also known as the plasma membrane (PM) or cytoplasmic membrane, and historically referred to as the plasmalemma) is a biological membrane that separates and protects the interior of all cells from the outside environment ( ...
s, including archaeol
Archaeol is composed of two phytanyl chains linked to the sn-2 and sn-3 positions of glycerol. As its phosphate ester, it is a common component of the membranes of archaea.
Structure and contrast with other lipids
Archaeol is a diether.
The 2 ...
s. Archaea use more diverse energy sources than eukaryotes, ranging from organic compounds
In chemistry, organic compounds are generally any chemical compounds that contain carbon-hydrogen or carbon-carbon bonds. Due to carbon's ability to catenate (form chains with other carbon atoms), millions of organic compounds are known. The s ...
such as sugars, to ammonia
Ammonia is an inorganic compound of nitrogen and hydrogen with the formula . A stable binary hydride, and the simplest pnictogen hydride, ammonia is a colourless gas with a distinct pungent smell. Biologically, it is a common nitrogenous wa ...
, metal ions
A metal (from Greek μέταλλον ''métallon'', "mine, quarry, metal") is a material that, when freshly prepared, polished, or fractured, shows a lustrous appearance, and conducts electricity and heat relatively well. Metals are typicall ...
or even hydrogen gas
Hydrogen is the chemical element with the symbol H and atomic number 1. Hydrogen is the lightest element. At standard conditions hydrogen is a gas of diatomic molecules having the formula . It is colorless, odorless, tasteless, non-toxic, a ...
. The salt-tolerant
Halotolerance is the adaptation of living organisms to conditions of high salinity. Halotolerant species tend to live in areas such as hypersaline lakes, coastal dunes, saline deserts, salt marshes, and inland salt seas and springs. Halophiles a ...
Haloarchaea
Haloarchaea (halophilic archaea, halophilic archaebacteria, halobacteria) are a class of the Euryarchaeota, found in water saturated or nearly saturated with salt. Halobacteria are now recognized as archaea rather than bacteria and are one of t ...
use sunlight as an energy source, and other species of archaea fix carbon (autotrophy), but unlike plants and cyanobacteria, no known species of archaea does both. Archaea reproduce asexually by binary fission
Binary may refer to:
Science and technology Mathematics
* Binary number, a representation of numbers using only two digits (0 and 1)
* Binary function, a function that takes two arguments
* Binary operation, a mathematical operation that ta ...
, fragmentation, or budding
Budding or blastogenesis is a type of asexual reproduction in which a new organism develops from an outgrowth or bud due to cell division at one particular site. For example, the small bulb-like projection coming out from the yeast cell is kno ...
; unlike bacteria, no known species of Archaea form endospores.
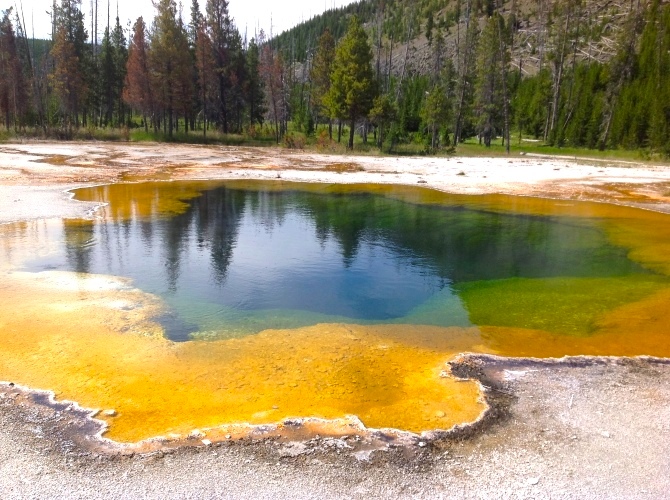
The first observed archaea were extremophile
An extremophile (from Latin ' meaning "extreme" and Greek ' () meaning "love") is an organism that is able to live (or in some cases thrive) in extreme environments, i.e. environments that make survival challenging such as due to extreme temper ...
s, living in extreme environments such as hot spring
A hot spring, hydrothermal spring, or geothermal spring is a spring produced by the emergence of geothermally heated groundwater onto the surface of the Earth. The groundwater is heated either by shallow bodies of magma (molten rock) or by c ...
s and salt lakes with no other organisms. Improved molecular detection tools led to the discovery of archaea in almost every habitat
In ecology, the term habitat summarises the array of resources, physical and biotic factors that are present in an area, such as to support the survival and reproduction of a particular species. A species habitat can be seen as the physical ...
, including soil, oceans, and marshland
A marsh is a wetland that is dominated by herbaceous rather than woody plant species.Keddy, P.A. 2010. Wetland Ecology: Principles and Conservation (2nd edition). Cambridge University Press, Cambridge, UK. 497 p Marshes can often be found at ...
s. Archaea are particularly numerous in the oceans, and the archaea in plankton
Plankton are the diverse collection of organisms found in water (or air) that are unable to propel themselves against a current (or wind). The individual organisms constituting plankton are called plankters. In the ocean, they provide a crucia ...
may be one of the most abundant groups of organisms on the planet.
Archaea are a major part of Earth's life. They are part of the microbiota of all organisms. In the human microbiome, they are important in the gut, mouth, and on the skin. Their morphological, metabolic, and geographical diversity permits them to play multiple ecological roles: carbon fixation; nitrogen cycling
The nitrogen cycle is the biogeochemical cycle by which nitrogen is converted into multiple chemical forms as it circulates among atmospheric, terrestrial, and marine ecosystems. The conversion of nitrogen can be carried out through both biolog ...
; organic compound turnover; and maintaining microbial symbiotic and syntrophic communities, for example.
No clear examples of archaeal pathogen
In biology, a pathogen ( el, πάθος, "suffering", "passion" and , "producer of") in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ ...
s or parasite
Parasitism is a close relationship between species, where one organism, the parasite, lives on or inside another organism, the host, causing it some harm, and is adapted structurally to this way of life. The entomologist E. O. Wilson has ...
s are known. Instead they are often mutualists
Mutualism describes the ecological Biological interaction, interaction between two or more species where each species has a net benefit. Mutualism is a common type of ecological interaction. Prominent examples include most vascular plants engag ...
or commensals, such as the methanogens (methane-producing strains) that inhabit the gastrointestinal tract in humans and ruminant
Ruminants (suborder Ruminantia) are hoofed herbivorous grazing or browsing mammals that are able to acquire nutrients from plant-based food by fermenting it in a specialized stomach prior to digestion, principally through microbial actions. The ...
s, where their vast numbers facilitate digestion
Digestion is the breakdown of large insoluble food molecules into small water-soluble food molecules so that they can be absorbed into the watery blood plasma. In certain organisms, these smaller substances are absorbed through the small intest ...
. Methanogens are also used in biogas
Biogas is a mixture of gases, primarily consisting of methane, carbon dioxide and hydrogen sulphide, produced from raw materials such as agricultural waste, manure, municipal waste, plant material, sewage, green waste and food waste. It is a ...
production and sewage treatment
Sewage treatment (or domestic wastewater treatment, municipal wastewater treatment) is a type of wastewater treatment which aims to remove contaminants from sewage to produce an effluent that is suitable for discharge to the surrounding e ...
, and biotechnology
Biotechnology is the integration of natural sciences and engineering sciences in order to achieve the application of organisms, cells, parts thereof and molecular analogues for products and services. The term ''biotechnology'' was first used ...
exploits enzymes from extremophile archaea that can endure high temperatures and organic solvents.
Classification
Early concept
 For much of the 20th century, prokaryotes were regarded as a single group of organisms and classified based on their
For much of the 20th century, prokaryotes were regarded as a single group of organisms and classified based on their biochemistry
Biochemistry or biological chemistry is the study of chemical processes within and relating to living organisms. A sub-discipline of both chemistry and biology, biochemistry may be divided into three fields: structural biology, enzymology and ...
, morphology
Morphology, from the Greek and meaning "study of shape", may refer to:
Disciplines
* Morphology (archaeology), study of the shapes or forms of artifacts
* Morphology (astronomy), study of the shape of astronomical objects such as nebulae, galaxies ...
and metabolism
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of life-sustaining chemical reactions in organisms. The three main functions of metabolism are: the conversion of the energy in food to energy available to run c ...
. Microbiologists tried to classify microorganisms based on the structures of their cell walls, their shapes, and the substances they consume. In 1965, Emile Zuckerkandl
Émile Zuckerkandl (July 4, 1922 – November 9, 2013) was an Austrian-born French biologist considered one of the founders of the field of molecular evolution. He introduced, with Linus Pauling, the concept of the "molecular clock", which enab ...
and Linus Pauling instead proposed using the sequences of the gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a ba ...
s in different prokaryotes to work out how they are related to each other. This phylogenetic
In biology, phylogenetics (; from Greek φυλή/ φῦλον [] "tribe, clan, race", and wikt:γενετικός, γενετικός [] "origin, source, birth") is the study of the evolutionary history and relationships among or within groups o ...
approach is the main method used today.
Archaea – at that time only the methanogens were known – were first classified separately from bacteria in 1977 by Carl Woese
Carl Richard Woese (; July 15, 1928 – December 30, 2012) was an American microbiologist and biophysicist. Woese is famous for defining the Archaea (a new domain of life) in 1977 through a pioneering phylogenetic taxonomy of 16S ribosomal RNA, ...
and George E. Fox
George Edward Fox (born December 17, 1945) is an astrobiologist, a Professor Emeritus and researcher at the University of Houston. He is an elected fellow of the American Academy of Microbiology, the American Association for the Advancement of Sc ...
based on their ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosom ...
(rRNA) genes. They called these groups the ''Urkingdoms'' of Archaebacteria and Eubacteria, though other researchers treated them as kingdoms
Kingdom commonly refers to:
* A monarchy ruled by a king or queen
* Kingdom (biology), a category in biological taxonomy
Kingdom may also refer to:
Arts and media Television
* ''Kingdom'' (British TV series), a 2007 British television drama s ...
or subkingdoms. Woese and Fox gave the first evidence for Archaebacteria as a separate "line of descent": 1. lack of peptidoglycan
Peptidoglycan or murein is a unique large macromolecule, a polysaccharide, consisting of sugars and amino acids that forms a mesh-like peptidoglycan layer outside the plasma membrane, the rigid cell wall (murein sacculus) characteristic of most ba ...
in their cell walls, 2. two unusual coenzymes, 3. results of 16S ribosomal RNA gene sequencing. To emphasize this difference, Woese, Otto Kandler
Otto Kandler (23 October 1920 in Deggendorf – 29 August 2017 in Munich, Bavaria)
was a German botanist and microbiologist. Until his retirement in 1986 he was professor of botany at the Ludwig Maximilian University of Munich.
His most importa ...
and Mark Wheelis Mark L. Wheelis is an American microbiologist. Wheelis is currently a professor in the College of Biological Sciences, University of California, Davis. Carl Woese and Otto Kandler with Wheelis wrote the important paper '' Towards a natural system o ...
later proposed reclassifying organisms into three natural domains known as the three-domain system
The three-domain system is a biological classification introduced by Carl Woese, Otto Kandler, and Mark Wheelis in 1990 that divides cellular life forms into three domains, namely Archaea, Bacteria, and Eukaryota or Eukarya. The key difference ...
: the Eukarya
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bact ...
, the Bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
and the Archaea, in what is now known as the Woesian Revolution.
The word ''archaea'' comes from the Ancient Greek
Ancient Greek includes the forms of the Greek language used in ancient Greece and the ancient world from around 1500 BC to 300 BC. It is often roughly divided into the following periods: Mycenaean Greek (), Dark Ages (), the Archaic peri ...
, meaning "ancient things", as the first representatives of the domain Archaea were methanogens and it was assumed that their metabolism reflected Earth's primitive atmosphere and the organisms' antiquity, but as new habitats were studied, more organisms were discovered. Extreme halophilic
The halophiles, named after the Greek word for "salt-loving", are extremophiles that thrive in high salt concentrations. While most halophiles are classified into the domain Archaea, there are also bacterial halophiles and some eukaryotic species, ...
and hyperthermophilic
A hyperthermophile is an organism that thrives in extremely hot environments—from 60 °C (140 °F) upwards. An optimal temperature for the existence of hyperthermophiles is often above 80 °C (176 °F). Hyperthermophiles are often within the doma ...
microbes were also included in Archaea. For a long time, archaea were seen as extremophiles that exist only in extreme habitats such as hot spring
A hot spring, hydrothermal spring, or geothermal spring is a spring produced by the emergence of geothermally heated groundwater onto the surface of the Earth. The groundwater is heated either by shallow bodies of magma (molten rock) or by c ...
s and salt lakes, but by the end of the 20th century, archaea had been identified in non-extreme environments as well. Today, they are known to be a large and diverse group of organisms abundantly distributed throughout nature. This new appreciation of the importance and ubiquity of archaea came from using polymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) ...
(PCR) to detect prokaryotes from environmental samples (such as water or soil) by multiplying their ribosomal genes. This allows the detection and identification of organisms that have not been cultured in the laboratory.
Classification
 The classification of archaea, and of prokaryotes in general, is a rapidly moving and contentious field. Current classification systems aim to organize archaea into groups of organisms that share structural features and common ancestors. These classifications rely heavily on the use of the sequence of
The classification of archaea, and of prokaryotes in general, is a rapidly moving and contentious field. Current classification systems aim to organize archaea into groups of organisms that share structural features and common ancestors. These classifications rely heavily on the use of the sequence of ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosom ...
genes to reveal relationships among organisms (molecular phylogenetics
Molecular phylogenetics () is the branch of phylogeny that analyzes genetic, hereditary molecular differences, predominantly in DNA sequences, to gain information on an organism's evolutionary relationships. From these analyses, it is possible to ...
). Most of the culturable and well-investigated species of archaea are members of two main phyla, the "Euryarchaeota
Euryarchaeota (from Ancient Greek ''εὐρύς'' eurús, "broad, wide") is a phylum of archaea. Euryarchaeota are highly diverse and include methanogens, which produce methane and are often found in intestines, halobacteria, which survive extr ...
" and the Thermoproteota
The Thermoproteota (also known as crenarchaea) are archaea that have been classified as a phylum of the Archaea domain. Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteris ...
(formerly Crenarchaeota). Other groups have been tentatively created, like the peculiar species ''Nanoarchaeum equitans
''Nanoarchaeum equitans'' is a species of marine archaea that was discovered in 2002 in a hydrothermal vent off the coast of Iceland on the Kolbeinsey Ridge by Karl Stetter. It has been proposed as the first species in a new phylum. Strains of ...
'', which was discovered in 2003 and has been given its own phylum, the "Nanoarchaeota
Nanoarchaeota (Greek, "dwarf or tiny ancient one") are a phylum of the Archaea. This phylum currently has only one representative, ''Nanoarchaeum equitans''.
Taxonomy
53 marker proteins based GTDB 07-RS207 phylogeny.
The currently accepted ...
". A new phylum "Korarchaeota
In taxonomy, the Korarchaeota are a phylum of the Archaea. The name is derived from the Greek noun koros or kore, meaning ''young man'' or ''young woman,'' and the Greek adjective archaios which means ''ancient.'' They are also known as Xenarchae ...
" has also been proposed. It contains a small group of unusual thermophilic species that shares features of both of the main phyla, but is most closely related to the Thermoproteota. Other recently detected species of archaea are only distantly related to any of these groups, such as the Archaeal Richmond Mine acidophilic nanoorganisms (ARMAN, comprising Micrarchaeota
DPANN is a super phylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared ...
and Parvarchaeota), which were discovered in 2006 and are some of the smallest organisms known.
A superphylum – TACK – which includes the Thaumarchaeota (now Nitrososphaerota
The Nitrososphaerota (syn. Thaumarchaeota) are a phylum of the Archaea proposed in 2008 after the genome of ''Cenarchaeum symbiosum'' was sequenced and found to differ significantly from other members of the hyperthermophilic phylum Thermoproteo ...
), "Aigarchaeota
The "Aigarchaeota" are a proposed archaeal phylum of which the main representative is '' Caldiarchaeum subterraneum''.. It is not yet clear if this represents a new phylum or a and order of the Nitrososphaerota, since the genome of ''Caldiarchae ...
", Crenarchaeota (now Thermoproteota
The Thermoproteota (also known as crenarchaea) are archaea that have been classified as a phylum of the Archaea domain. Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteris ...
), and "Korarchaeota
In taxonomy, the Korarchaeota are a phylum of the Archaea. The name is derived from the Greek noun koros or kore, meaning ''young man'' or ''young woman,'' and the Greek adjective archaios which means ''ancient.'' They are also known as Xenarchae ...
" was proposed in 2011 to be related to the origin of eukaryotes. In 2017, the newly discovered and newly named Asgard
In Nordic mythology, Asgard (Old Norse: ''Ásgarðr'' ; "enclosure of the Æsir") is a location associated with the gods. It appears in a multitude of Old Norse sagas and mythological texts. It is described as the fortified home of the Æsir ...
superphylum was proposed to be more closely related to the original eukaryote and a sister group to TACK.
In 2013 the superphylum DPANN was proposed to group "Nanoarchaeota
Nanoarchaeota (Greek, "dwarf or tiny ancient one") are a phylum of the Archaea. This phylum currently has only one representative, ''Nanoarchaeum equitans''.
Taxonomy
53 marker proteins based GTDB 07-RS207 phylogeny.
The currently accepted ...
", "Nanohaloarchaeota
Nanohaloarchaea is a clade of diminutive archaea with small genomes and limited metabolic capabilities, belonging to the DPANN archaea. They are ubiquitous in hypersaline habitats, which they share with the extremely halophilic haloarchaea.
Nan ...
", Archaeal Richmond Mine acidophilic nanoorganisms (ARMAN, comprising "Micrarchaeota
DPANN is a super phylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared ...
" and "Parvarchaeota
Parvarchaeota is a phylum of archaea belonging to the DPANN archaea. They have been discovered in acid mine drainage waters and later in marine sediments. The cells of these organisms are extremely small consistent with small genomes. Metageno ...
"), and other similar archaea. This archaeal superphylum encompasses at least 10 different lineages and includes organisms with extremely small cell and genome sizes and limited metabolic capabilities. Therefore, many members of DPANN may be obligately dependent on symbiotic interactions with other organisms and may even include novel parasites. However, in other phylogenetic analyses it was found that DPANN does not form a monophyletic group and that it is caused by the long branch attraction
In phylogenetics, long branch attraction (LBA) is a form of systematic error whereby distantly related lineages are incorrectly inferred to be closely related. LBA arises when the amount of molecular or morphological change accumulated within a lin ...
(LBA), suggesting that all these lineages belong to "Euryarchaeota".
Cladogram
According to Tom A. Williams ''et al.'' (2017), Castelle & Banfield (2018) andGTDB
The Genome Taxonomy Database (GTDB) is an online database that maintains information on a proposed nomenclature of prokaryotes, following a phylogenomic approach based on a set of conserved single-copy proteins. In addition to breaking up parap ...
release 07-RS207 (8th April 2022):
Concept of species
The classification of archaea into species is also controversial. Biology defines aspecies
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriate s ...
as a group of related organisms. The familiar exclusive breeding criterion (organisms that can breed with each other but not with others) is of no help since archaea only reproduce asexually.
Archaea show high levels of horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). H ...
between lineages. Some researchers suggest that individuals can be grouped into species-like populations given highly similar genomes and infrequent gene transfer to/from cells with less-related genomes, as in the genus ''Ferroplasma
''Ferroplasma'' is a genus of Archaea that belong to the family Ferroplasmaceae. Members of the ''Ferroplasma'' are typically acidophillic, pleomorphic, irregularly shaped cocci.
The archaean family Ferroplasmaceae was first described in the ea ...
''. On the other hand, studies in ''Halorubrum
''Halorubrum'' is a genus in the family Halorubraceae. ''Halorubrum'' species areusually halophilic and can be found in waters with high salt concentration such as the Dead Sea or Lake Zabuye.
Genetic exchange
A population of the haloarchaea '' ...
'' found significant genetic transfer to/from less-related populations, limiting the criterion's applicability. Some researchers question whether such species designations have practical meaning.
Current knowledge on genetic diversity is fragmentary and the total number of archaeal species cannot be estimated with any accuracy. Estimates of the number of phyla range from 18 to 23, of which only 8 have representatives that have been cultured and studied directly. Many of these hypothesized groups are known from a single rRNA sequence, indicating that the diversity among these organisms remains obscure. The Bacteria also include many uncultured microbes with similar implications for characterization.
Phyla
Valid Phyla
The following phyla have been validly published according to theBacteriological Code
The International Code of Nomenclature of Prokaryotes (ICNP) formerly the International Code of Nomenclature of Bacteria (ICNB) or Bacteriological Code (BC) governs the scientific names for Bacteria and Archaea.P. H. A. Sneath, 2003. A short histor ...
:
* Nitrososphaerota
The Nitrososphaerota (syn. Thaumarchaeota) are a phylum of the Archaea proposed in 2008 after the genome of ''Cenarchaeum symbiosum'' was sequenced and found to differ significantly from other members of the hyperthermophilic phylum Thermoproteo ...
* Thermoproteota
The Thermoproteota (also known as crenarchaea) are archaea that have been classified as a phylum of the Archaea domain. Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteris ...
Provisional Phyla
The following phyla have been proposed, but have not been validly published according to the Bacteriological Code (including those that have ''candidatus
In prokaryote nomenclature, ''Candidatus'' (Latin for candidate of Roman office) is used to name prokaryotic phyla that are well characterized but yet-uncultured. Contemporary sequencing approaches, such as 16S sequencing or metagenomics, provide m ...
'' status):
* "''Candidatus'' Aenigmarchaeota
DPANN is a super phylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared ...
"
* "''Candidatus'' Aigarchaeota
The "Aigarchaeota" are a proposed archaeal phylum of which the main representative is '' Caldiarchaeum subterraneum''.. It is not yet clear if this represents a new phylum or a and order of the Nitrososphaerota, since the genome of ''Caldiarchae ...
"
* "''Candidatus'' Altiarchaeota"
* "''Candidatus'' Asgardaeota
Asgard or Asgardarchaeota is a proposed superphylum consisting of a group of archaea that includes Lokiarchaeota, Thorarchaeota, Odinarchaeota, and Heimdallarchaeota. It appears the eukaryotes emerged within the Asgard, in a branch containing ...
"
* "''Candidatus'' Bathyarchaeota"
* "''Candidatus'' Brockarchaeota"
* "''Candidatus'' Diapherotrites
DPANN is a superphylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared t ...
"
* "''Euryarchaeota
Euryarchaeota (from Ancient Greek ''εὐρύς'' eurús, "broad, wide") is a phylum of archaea. Euryarchaeota are highly diverse and include methanogens, which produce methane and are often found in intestines, halobacteria, which survive extr ...
''"
* "''Candidatus'' Geoarchaeota
TACK is a group of archaea acronym for Thaumarchaeota (now Nitrososphaerota), Aigarchaeota, Crenarchaeota (now Thermoproteota), and Korarchaeota, the first groups discovered. They are found in different environments ranging from acidophilic th ...
"
* "''Candidatus'' Hadarchaeota"
* "''Candidatus'' Hadesarchaeota"
* "''Candidatus'' Halobacterota"
* "''Candidatus'' Heimdallarchaeota
Asgard or Asgardarchaeota is a proposed superphylum consisting of a group of archaea that includes Lokiarchaeota, Thorarchaeota, Odinarchaeota, and Heimdallarchaeota. It appears the eukaryotes emerged within the Asgard, in a branch containin ...
"
* "''Candidatus'' Helarchaeota"
* "''Candidatus'' Huberarchaeota"
* "''Candidatus'' Hydrothermarchaeota"
* "''Candidatus'' Korarchaeota
In taxonomy, the Korarchaeota are a phylum of the Archaea. The name is derived from the Greek noun koros or kore, meaning ''young man'' or ''young woman,'' and the Greek adjective archaios which means ''ancient.'' They are also known as Xenarchae ...
"
* "''Candidatus'' Lokiarchaeia"
* "''Candidatus'' Lokiarchaeota
Lokiarchaeota is a proposed phylum of the Archaea. The phylum includes all members of the group previously named Deep Sea Archaeal Group (DSAG), also known as Marine Benthic Group B (MBG-B). Lokiarchaeota is part of the superphylum Asgard contai ...
"
* "''Candidatus'' Mamarchaeota"
* "''Candidatus'' Marsarchaeota"
* "''Candidatus'' Methanobacteriota"
* "''Candidatus'' Micrarchaeota
DPANN is a super phylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared ...
"
* "''Candidatus'' Nanoarchaeota
Nanoarchaeota (Greek, "dwarf or tiny ancient one") are a phylum of the Archaea. This phylum currently has only one representative, ''Nanoarchaeum equitans''.
Taxonomy
53 marker proteins based GTDB 07-RS207 phylogeny.
The currently accepted ...
"
* "''Candidatus'' Nanohaloarchaeota
Nanohaloarchaea is a clade of diminutive archaea with small genomes and limited metabolic capabilities, belonging to the DPANN archaea. They are ubiquitous in hypersaline habitats, which they share with the extremely halophilic haloarchaea.
Nan ...
"
* "''Candidatus'' Nezhaarchaeota"
* "''Candidatus'' Odinarchaeota
Asgard or Asgardarchaeota is a proposed superphylum consisting of a group of archaea that includes Lokiarchaeota, Thorarchaeota, Odinarchaeota, and Heimdallarchaeota. It appears the eukaryotes emerged within the Asgard, in a branch containin ...
"
* "''Candidatus'' Pacearchaeota
DPANN is a superphylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared t ...
"
* "''Candidatus'' Parvarchaeota
Parvarchaeota is a phylum of archaea belonging to the DPANN archaea. They have been discovered in acid mine drainage waters and later in marine sediments. The cells of these organisms are extremely small consistent with small genomes. Metageno ...
"
* "''Candidatus'' Thermoplasmatota"
* "''Candidatus'' Thorarchaeota
"''Candidatus'' Thorarchaeota", or simply Thorarchaeota, is a phylum within the superphylum Asgard archaea. The Asgard superphylum represents the closest prokaryotic relatives of eukaryotes. Since there is such a close relation between the tw ...
"
* "''Candidatus'' Undinarchaeota"
* "''Candidatus'' Verstraetearchaeota"
* "''Candidatus'' Woesearchaeota
DPANN is a superphylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared t ...
"
Origin and evolution
The age of the Earth is about 4.54 billion years. Scientific evidence suggests that life began on Earth at least 3.5billion years ago
bya or b.y.a. is an abbreviation for "billion years ago". It is commonly used as a unit of time to denote length of time before the present in 109 years. This initialism is often used in the sciences of astronomy, geology, and paleontology.
The " ...
. The earliest evidence for life on Earth Life on Earth may refer to:
Science
* Life
* Earliest known life forms
* Evolutionary history of life
** Abiogenesis
Film and television
* ''Life on Earth'' (film) (''La Vie Sur Terre''), a 1998 Malian film
* ''Life on Earth'' (TV series), a 197 ...
is graphite
Graphite () is a crystalline form of the element carbon. It consists of stacked layers of graphene. Graphite occurs naturally and is the most stable form of carbon under standard conditions. Synthetic and natural graphite are consumed on lar ...
found to be biogenic in 3.7-billion-year-old metasedimentary rocks
In geology, metasedimentary rock is a type of metamorphic rock. Such a rock was first formed through the deposition and solidification of sediment
Sediment is a naturally occurring material that is broken down by processes of weathering and e ...
discovered in Western Greenland
Kitaa, originally Vestgrønland ("West Greenland"), is a former administrative division of Greenland. It was by far the most populated of the divisions, being home to almost 90% of the total population. The divisions were de facto replaced by st ...
and microbial mat
A microbial mat is a multi-layered sheet of microorganisms, mainly bacteria and archaea, or bacteria alone. Microbial mats grow at interfaces between different types of material, mostly on submerged or moist surfaces, but a few survive in deserts ...
fossils found in 3.48-billion-year-old sandstone
Sandstone is a clastic sedimentary rock composed mainly of sand-sized (0.0625 to 2 mm) silicate grains. Sandstones comprise about 20–25% of all sedimentary rocks.
Most sandstone is composed of quartz or feldspar (both silicates ...
discovered in Western Australia
Western Australia (commonly abbreviated as WA) is a state of Australia occupying the western percent of the land area of Australia excluding external territories. It is bounded by the Indian Ocean to the north and west, the Southern Ocean to th ...
. In 2015, possible remains of biotic matter were found in 4.1-billion-year-old rocks in Western Australia.
Although probable prokaryotic cell fossils date to almost 3.5 billion years ago
bya or b.y.a. is an abbreviation for "billion years ago". It is commonly used as a unit of time to denote length of time before the present in 109 years. This initialism is often used in the sciences of astronomy, geology, and paleontology.
The " ...
, most prokaryotes do not have distinctive morphologies, and fossil shapes cannot be used to identify them as archaea. Instead, chemical fossils of unique lipid
Lipids are a broad group of naturally-occurring molecules which includes fats, waxes, sterols, fat-soluble vitamins (such as vitamins A, D, E and K), monoglycerides, diglycerides, phospholipids, and others. The functions of lipids includ ...
s are more informative because such compounds do not occur in other organisms. Some publications suggest that archaeal or eukaryotic lipid remains are present in shales dating from 2.7 billion years ago, though such data have since been questioned. These lipids have also been detected in even older rocks from west Greenland
Greenland ( kl, Kalaallit Nunaat, ; da, Grønland, ) is an island country in North America that is part of the Kingdom of Denmark. It is located between the Arctic and Atlantic oceans, east of the Canadian Arctic Archipelago. Greenland is t ...
. The oldest such traces come from the Isua district, which includes Earth's oldest known sediments, formed 3.8 billion years ago. The archaeal lineage may be the most ancient that exists on Earth.
Woese argued that the Bacteria, Archaea, and Eukaryotes represent separate lines of descent that diverged early on from an ancestral colony of organisms. One possibility is that this occurred before the evolution of cells, when the lack of a typical cell membrane allowed unrestricted lateral gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring ( reproduction). ...
, and that the common ancestors of the three domains arose by fixation of specific subsets of genes. It is possible that the last common ancestor of bacteria and archaea was a thermophile
A thermophile is an organism—a type of extremophile—that thrives at relatively high temperatures, between . Many thermophiles are archaea, though they can be bacteria or fungi. Thermophilic eubacteria are suggested to have been among the earl ...
, which raises the possibility that lower temperatures are "extreme environments" for archaea, and organisms that live in cooler environments appeared only later. Since archaea and bacteria are no more related to each other than they are to eukaryotes, the term ''prokaryote'' may suggest a false similarity between them. However, structural and functional similarities between lineages often occur because of shared ancestral traits or evolutionary convergence
Convergent evolution is the independent evolution of similar features in species of different periods or epochs in time. Convergent evolution creates analogous structures that have similar form or function but were not present in the last com ...
. These similarities are known as a ''grade'', and prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Conne ...
s are best thought of as a grade of life, characterized by such features as an absence of membrane-bound organelles.
Comparison with other domains
The following table compares some major characteristics of the three domains, to illustrate their similarities and differences. Archaea were split off as a third domain because of the large differences in their ribosomal RNA structure. The particular molecule 16S rRNA is key to the production of proteins in all organisms. Because this function is so central to life, organisms with mutations in their 16S rRNA are unlikely to survive, leading to great (but not absolute) stability in the structure of this polynucleotide over generations. 16S rRNA is large enough to show organism-specific variations, but still small enough to be compared quickly. In 1977, Carl Woese, a microbiologist studying the genetic sequences of organisms, developed a new comparison method that involved splitting the RNA into fragments that could be sorted and compared with other fragments from other organisms. The more similar the patterns between species, the more closely they are related. Woese used his new rRNA comparison method to categorize and contrast different organisms. He compared a variety of species and happened upon a group of methanogens with rRNA vastly different from any known prokaryotes or eukaryotes. These methanogens were much more similar to each other than to other organisms, leading Woese to propose the new domain of Archaea. His experiments showed that the archaea were genetically more similar to eukaryotes than prokaryotes, even though they were more similar to prokaryotes in structure. This led to the conclusion that Archaea and Eukarya shared a common ancestor more recent than Eukarya and Bacteria. The development of the nucleus occurred after the split between Bacteria and this common ancestor. One property unique to archaea is the abundant use of ether-linked lipids in their cell membranes. Ether linkages are more chemically stable than the ester linkages found in bacteria and eukarya, which may be a contributing factor to the ability of many archaea to survive in extreme environments that place heavy stress on cell membranes, such as extreme heat and salinity. Comparative analysis of archaeal genomes has also identified several molecularconserved signature indels Conserved signature inserts and deletions (CSIs) in protein sequences provide an important category of molecular markers for understanding phylogenetic relationships. CSIs, brought about by rare genetic changes, provide useful phylogenetic markers ...
and signature proteins uniquely present in either all archaea or different main groups within archaea. Another unique feature of archaea, found in no other organisms, is methanogenesis (the metabolic production of methane). Methanogenic archaea play a pivotal role in ecosystems with organisms that derive energy from oxidation of methane, many of which are bacteria, as they are often a major source of methane in such environments and can play a role as primary producers. Methanogens also play a critical role in the carbon cycle
The carbon cycle is the biogeochemical cycle by which carbon is exchanged among the biosphere, pedosphere, geosphere, hydrosphere, and atmosphere of the Earth. Carbon is the main component of biological compounds as well as a major componen ...
, breaking down organic carbon into methane, which is also a major greenhouse gas.
This difference in biochemical structure of Bacteria and Archaea has been explained by researchers that they originated at deep sea alkaline hydrothermal vents, where they independently developed lipid biosynthesis and cell wall biochemistry during their transition to Archaea and Bacteria. It has been suggested that the last universal common ancestor was a not free-living organism. However this view has been challenged by other researchers and is currently in dispute.
Relationship to bacteria
The relationships among the three domains are of central importance for understanding the origin of life. Most of themetabolic pathway
In biochemistry, a metabolic pathway is a linked series of chemical reactions occurring within a cell. The reactants, products, and intermediates of an enzymatic reaction are known as metabolites, which are modified by a sequence of chemical reac ...
s, which are the object of the majority of an organism's genes, are common between Archaea and Bacteria, while most genes involved in genome expression are common between Archaea and Eukarya. Within prokaryotes, archaeal cell structure is most similar to that of gram-positive
In bacteriology, gram-positive bacteria are bacteria that give a positive result in the Gram stain test, which is traditionally used to quickly classify bacteria into two broad categories according to their type of cell wall.
Gram-positive bact ...
bacteria, largely because both have a single lipid bilayer and usually contain a thick sacculus (exoskeleton) of varying chemical composition. In some phylogenetic trees based upon different gene/protein sequences of prokaryotic homologs, the archaeal homologs are more closely related to those of gram-positive bacteria. Archaea and gram-positive bacteria also share conserved indels
Indel is a molecular biology term for an insertion or deletion of bases in the genome of an organism. It is classified among small genetic variations, measuring from 1 to 10 000 base pairs in length, including insertion and deletion events that ...
in a number of important proteins, such as Hsp70
The 70 kilodalton heat shock proteins (Hsp70s or DnaK) are a family of conserved ubiquitously expressed heat shock proteins. Proteins with similar structure exist in virtually all living organisms. Intracellularly localized Hsp70s are an import ...
and glutamine synthetase
Glutamine synthetase (GS) () is an enzyme that plays an essential role in the metabolism of nitrogen by catalyzing the condensation of glutamate and ammonia to form glutamine:
Glutamate + ATP + NH3 → Glutamine + ADP + phosphate
Glutam ...
I; but the phylogeny of these genes was interpreted to reveal interdomain gene transfer, and might not reflect the organismal relationship(s).
It has been proposed that the archaea evolved from gram-positive bacteria in response to antibiotic selection pressure
Any cause that reduces or increases reproductive success in a portion of a population potentially exerts evolutionary pressure, selective pressure or selection pressure, driving natural selection. It is a quantitative description of the amount of ...
. This is suggested by the observation that archaea are resistant to a wide variety of antibiotics that are produced primarily by gram-positive bacteria, and that these antibiotics act primarily on the genes that distinguish archaea from bacteria. The proposal is that the selective pressure towards resistance generated by the gram-positive antibiotics was eventually sufficient to cause extensive changes in many of the antibiotics' target genes, and that these strains represented the common ancestors of present-day Archaea. The evolution of Archaea in response to antibiotic selection, or any other competitive selective pressure, could also explain their adaptation to extreme environments (such as high temperature or acidity) as the result of a search for unoccupied niches to escape from antibiotic-producing organisms; Cavalier-Smith
Thomas (Tom) Cavalier-Smith, FRS, FRSC, NERC Professorial Fellow (21 October 1942 – 19 March 2021), was a professor of evolutionary biology in the Department of Zoology, at the University of Oxford.
His research has led to disc ...
has made a similar suggestion. This proposal is also supported by other work investigating protein structural relationships and studies that suggest that gram-positive bacteria may constitute the earliest branching lineages within the prokaryotes.
Relation to eukaryotes
The evolutionary relationship between archaea and eukaryotes remains unclear. Aside from the similarities in cell structure and function that are discussed below, many genetic trees group the two. Complicating factors include claims that the relationship between eukaryotes and the archaeal phylumThermoproteota
The Thermoproteota (also known as crenarchaea) are archaea that have been classified as a phylum of the Archaea domain. Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteris ...
is closer than the relationship between the "Euryarchaeota
Euryarchaeota (from Ancient Greek ''εὐρύς'' eurús, "broad, wide") is a phylum of archaea. Euryarchaeota are highly diverse and include methanogens, which produce methane and are often found in intestines, halobacteria, which survive extr ...
" and the phylum Thermoproteota and the presence of archaea-like genes in certain bacteria, such as ''Thermotoga maritima
''Thermotoga maritima'' is a hyperthermophilic, anaerobic organism that is a member of the order Thermotogales. ''T. maritima'' is well known for its ability to produce hydrogen (clean energy) and it is the only fermentative bacterium that has b ...
'', from horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). H ...
. The standard hypothesis states that the ancestor of the eukaryotes diverged early from the Archaea, and that eukaryotes arose through fusion of an archaean and eubacterium, which became the nucleus and cytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
; this hypothesis explains the genetic similarities between the groups. The eocyte hypothesis
The eocyte hypothesis in evolutionary biology proposes the origin of eukaryotes from a group of prokaryotes called eocytes (later classified as Thermoproteota, a group of archaea). After his team at the University of California, Los Angeles disc ...
instead posits that Eukaryota emerged relatively late from the Archaea.
A lineage of archaea discovered in 2015, '' Lokiarchaeum'' (of proposed new Phylum "Lokiarchaeota
Lokiarchaeota is a proposed phylum of the Archaea. The phylum includes all members of the group previously named Deep Sea Archaeal Group (DSAG), also known as Marine Benthic Group B (MBG-B). Lokiarchaeota is part of the superphylum Asgard contai ...
"), named for a hydrothermal vent called Loki's Castle
Loki's Castle is a field of five active hydrothermal vents in the mid-Atlantic Ocean, located at 73 degrees north on the Mid-Atlantic Ridge between Greenland and Norway at a depth of . The vents were discovered in mid-July 2008 and are the most ...
in the Arctic Ocean, was found to be the most closely related to eukaryotes known at that time. It has been called a transitional organism between prokaryotes and eukaryotes.
Several sister phyla of "Lokiarchaeota" have since been found ("Thorarchaeota
"''Candidatus'' Thorarchaeota", or simply Thorarchaeota, is a phylum within the superphylum Asgard archaea. The Asgard superphylum represents the closest prokaryotic relatives of eukaryotes. Since there is such a close relation between the tw ...
", "Odinarchaeota
Asgard or Asgardarchaeota is a proposed superphylum consisting of a group of archaea that includes Lokiarchaeota, Thorarchaeota, Odinarchaeota, and Heimdallarchaeota. It appears the eukaryotes emerged within the Asgard, in a branch containin ...
", "Heimdallarchaeota
Asgard or Asgardarchaeota is a proposed superphylum consisting of a group of archaea that includes Lokiarchaeota, Thorarchaeota, Odinarchaeota, and Heimdallarchaeota. It appears the eukaryotes emerged within the Asgard, in a branch containin ...
"), all together comprising a newly proposed supergroup Asgard
In Nordic mythology, Asgard (Old Norse: ''Ásgarðr'' ; "enclosure of the Æsir") is a location associated with the gods. It appears in a multitude of Old Norse sagas and mythological texts. It is described as the fortified home of the Æsir ...
, which may appear as a sister taxon to Proteoarchaeota
"Proteoarchaeota" are a proposed archaeal kingdom thought to be closely related to the Eukaryotes.Approximately the same group is sometimes referred to as TACK after the initial letters of its early-found daughter clades: Thaumarchaeota (now Ni ...
.
Details of the relation of Asgard members and eukaryotes are still under consideration, although, in January 2020, scientists reported that ''Candidatus Prometheoarchaeum syntrophicum
Lokiarchaeota is a proposed phylum of the Archaea. The phylum includes all members of the group previously named Deep Sea Archaeal Group (DSAG), also known as Marine Benthic Group B (MBG-B). Lokiarchaeota is part of the superphylum Asgard contai ...
'', a type of Asgard archaea, may be a possible link between simple prokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
and complex eukaryotic
Eukaryotes () are organisms whose Cell (biology), cells have a cell nucleus, nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the ...
microorganisms about two billion years ago.
Morphology
Individual archaea range from 0.1micrometers
The micrometre ( international spelling as used by the International Bureau of Weights and Measures; SI symbol: μm) or micrometer (American spelling), also commonly known as a micron, is a unit of length in the International System of Unit ...
(μm) to over 15 μm in diameter, and occur in various shapes, commonly as spheres, rods, spirals or plates. Other morphologies in the Thermoproteota
The Thermoproteota (also known as crenarchaea) are archaea that have been classified as a phylum of the Archaea domain. Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteris ...
include irregularly shaped lobed cells in '' Sulfolobus'', needle-like filaments that are less than half a micrometer in diameter in '' Thermofilum'', and almost perfectly rectangular rods in ''Thermoproteus
In alpha taxonomy, taxonomy, ''Thermoproteus'' is a genus (biology), genus of the Thermoproteaceae. These prokaryotes are thermophile, thermophilic sulphur-dependent organisms related to the genera ''Sulfolobus'', ''Pyrodictium'' and ''Desulfuroc ...
'' and ''Pyrobaculum
''Pyrobaculum'' is a genus of the Thermoproteaceae.
Description and significance
As its Latin name ''Pyrobaculum'' (the "fire stick") suggests, the archaeon is rod-shaped and isolated from locations with high temperatures. It is Gram-negative ...
''. Archaea in the genus ''Haloquadratum
''Haloquadratum'' (common abbreviation: ''Hqr.'') is a genus of archaean, belonging to the family Haloferacaceae. The first species to be identified in this group, ''Haloquadratum walsbyi'', is unusual in that its cells are shaped like square, ...
'' such as ''Haloquadratum walsbyi
''Haloquadratum walsbyi'' is of the genus ''Haloquadratum,'' within the archaea domain known for its square halophilic nature. First discovered in a brine pool in the Sinai peninsula of Egypt, ''H. walsbyi'' is noted for its flat, square-shaped ...
'' are flat, square specimens that live in hypersaline pools. These unusual shapes are probably maintained by both their cell walls and a prokaryotic cytoskeleton
The prokaryotic cytoskeleton is the collective name for all structural filaments in prokaryotes. It was once thought that prokaryotic cells did not possess cytoskeletons, but advances in visualization technology and structure determination led t ...
. Proteins related to the cytoskeleton components of other organisms exist in archaea, and filaments form within their cells, but in contrast with other organisms, these cellular structures are poorly understood. In ''Thermoplasma
In taxonomy, ''Thermoplasma'' is a genus of the Thermoplasmataceae.See the NCBIbr>webpage on Thermoplasma Data extracted from the
''Thermoplasma'' is a genus of archaea. It belongs to the Thermoplasmata, which thrive in acidic and high-tempe ...
'' and ''Ferroplasma
''Ferroplasma'' is a genus of Archaea that belong to the family Ferroplasmaceae. Members of the ''Ferroplasma'' are typically acidophillic, pleomorphic, irregularly shaped cocci.
The archaean family Ferroplasmaceae was first described in the ea ...
'' the lack of a cell wall means that the cells have irregular shapes, and can resemble amoebae
An amoeba (; less commonly spelled ameba or amœba; plural ''am(o)ebas'' or ''am(o)ebae'' ), often called an amoeboid, is a type of cell or unicellular organism with the ability to alter its shape, primarily by extending and retracting pseudopo ...
.
Some species form aggregates or filaments of cells up to 200 μm long. These organisms can be prominent in biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
s. Notably, aggregates of '' Thermococcus coalescens'' cells fuse together in culture, forming single giant cells. Archaea in the genus '' Pyrodictium'' produce an elaborate multicell colony involving arrays of long, thin hollow tubes called ''cannulae'' that stick out from the cells' surfaces and connect them into a dense bush-like agglomeration. The function of these cannulae is not settled, but they may allow communication or nutrient exchange with neighbors. Multi-species colonies exist, such as the "string-of-pearls" community that was discovered in 2001 in a German swamp. Round whitish colonies of a novel Euryarchaeota species are spaced along thin filaments that can range up to long; these filaments are made of a particular bacteria species.
Structure, composition development, and operation
Archaea and bacteria have generally similarcell
Cell most often refers to:
* Cell (biology), the functional basic unit of life
Cell may also refer to:
Locations
* Monastic cell, a small room, hut, or cave in which a religious recluse lives, alternatively the small precursor of a monastery ...
structure, but cell composition and organization set the archaea apart. Like bacteria, archaea lack interior membranes and organelles. Like bacteria, the cell membrane
The cell membrane (also known as the plasma membrane (PM) or cytoplasmic membrane, and historically referred to as the plasmalemma) is a biological membrane that separates and protects the interior of all cells from the outside environment ( ...
s of archaea are usually bounded by a cell wall and they swim using one or more flagella. Structurally, archaea are most similar to gram-positive bacteria. Most have a single plasma membrane and cell wall, and lack a periplasmic space
The periplasm is a concentrated gel-like matrix in the space between the inner cytoplasmic membrane and the bacterial outer membrane called the ''periplasmic space'' in gram-negative bacteria. Using cryo-electron microscopy it has been found tha ...
; the exception to this general rule is ''Ignicoccus
''Ignicoccus'' is a genus of hyperthermophillic Archaea living in marine hydrothermal vents. They were discovered in samples taken at the Kolbeinsey Ridge north of Iceland, as well as at the East Pacific Rise (at 9 degrees N, 104 degrees W) ...
'', which possess a particularly large periplasm that contains membrane-bound vesicles
Vesicle may refer to:
; In cellular biology or chemistry
* Vesicle (biology and chemistry), a supramolecular assembly of lipid molecules, like a cell membrane
* Synaptic vesicle
; In human embryology
* Vesicle (embryology), bulge-like features o ...
and is enclosed by an outer membrane.
Cell wall and archaella
Most archaea (but not ''Thermoplasma
In taxonomy, ''Thermoplasma'' is a genus of the Thermoplasmataceae.See the NCBIbr>webpage on Thermoplasma Data extracted from the
''Thermoplasma'' is a genus of archaea. It belongs to the Thermoplasmata, which thrive in acidic and high-tempe ...
'' and ''Ferroplasma
''Ferroplasma'' is a genus of Archaea that belong to the family Ferroplasmaceae. Members of the ''Ferroplasma'' are typically acidophillic, pleomorphic, irregularly shaped cocci.
The archaean family Ferroplasmaceae was first described in the ea ...
'') possess a cell wall. In most archaea the wall is assembled from surface-layer proteins, which form an S-layer An S-layer (surface layer) is a part of the cell envelope found in almost all archaea, as well as in many types of bacteria.
The S-layers of both archaea and bacteria consists of a monomolecular layer composed of only one (or, in a few cases, two) ...
. An S-layer is a rigid array of protein molecules that cover the outside of the cell (like chain mail
Chain mail (properly called mail or maille but usually called chain mail or chainmail) is a type of armour consisting of small metal rings linked together in a pattern to form a mesh. It was in common military use between the 3rd century BC and ...
). This layer provides both chemical and physical protection, and can prevent macromolecules from contacting the cell membrane. Unlike bacteria, archaea lack peptidoglycan
Peptidoglycan or murein is a unique large macromolecule, a polysaccharide, consisting of sugars and amino acids that forms a mesh-like peptidoglycan layer outside the plasma membrane, the rigid cell wall (murein sacculus) characteristic of most ba ...
in their cell walls. Methanobacteriales
In taxonomy, the Methanobacteriales are an order of the Methanobacteria. Species within this order differ from other methanogens in that they can use fewer catabolic substrates and have distinct morphological characteristics, lipid compositi ...
do have cell walls containing pseudopeptidoglycan
Pseudopeptidoglycan (also known as pseudomurein;White, David. (1995) ''The Physiology and Biochemistry of Prokaryotes'', pages 6, 12-21. (Oxford: Oxford University Press). . PPG hereafter) is a major cell wall component of some Archaea that differs ...
, which resembles eubacterial peptidoglycan in morphology, function, and physical structure, but pseudopeptidoglycan is distinct in chemical structure; it lacks D-amino acids and N-acetylmuramic acid
''N''-Acetylmuramic acid (NAM or MurNAc) is an organic compound with the chemical formula . It is a monomer of peptidoglycan in most bacterial cell walls, which is built from alternating units of ''N''-acetylglucosamine (GlcNAc) and ''N''-acet ...
, substituting the latter with N-Acetyltalosaminuronic acid.
Archaeal flagella are known as archaella, that operate like bacterial flagella – their long stalks are driven by rotatory motors at the base. These motors are powered by a proton gradient
An electrochemical gradient is a gradient of electrochemical potential, usually for an ion that can move across a membrane. The gradient consists of two parts, the chemical gradient, or difference in solute concentration across a membrane, and th ...
across the membrane, but archaella are notably different in composition and development. The two types of flagella evolved from different ancestors. The bacterial flagellum shares a common ancestor with the type III secretion system
The type III secretion system (T3SS or TTSS), also called the injectisome, is one of the bacterial secretion systems used by bacteria to secrete their effector proteins into the host's cells to promote virulence and colonisation. The T3SS is a ...
, while archaeal flagella appear to have evolved from bacterial type IV pili
A pilus (Latin for 'hair'; plural: ''pili'') is a hair-like appendage found on the surface of many bacteria and archaea. The terms ''pilus'' and '' fimbria'' (Latin for 'fringe'; plural: ''fimbriae'') can be used interchangeably, although some r ...
. In contrast with the bacterial flagellum, which is hollow and assembled by subunits moving up the central pore to the tip of the flagella, archaeal flagella are synthesized by adding subunits at the base.
Membranes
cell membrane
The cell membrane (also known as the plasma membrane (PM) or cytoplasmic membrane, and historically referred to as the plasmalemma) is a biological membrane that separates and protects the interior of all cells from the outside environment ( ...
s are made of molecules known as phospholipids. These molecules possess both a polar
Polar may refer to:
Geography
Polar may refer to:
* Geographical pole, either of two fixed points on the surface of a rotating body or planet, at 90 degrees from the equator, based on the axis around which a body rotates
* Polar climate, the c ...
part that dissolves in water (the phosphate
In chemistry, a phosphate is an anion, salt, functional group or ester derived from a phosphoric acid. It most commonly means orthophosphate, a derivative of orthophosphoric acid .
The phosphate or orthophosphate ion is derived from phosph ...
"head"), and a "greasy" non-polar part that does not (the lipid tail). These dissimilar parts are connected by a glycerol
Glycerol (), also called glycerine in British English and glycerin in American English, is a simple triol compound. It is a colorless, odorless, viscous liquid that is sweet-tasting and non-toxic. The glycerol backbone is found in lipids known ...
moiety. In water, phospholipids cluster, with the heads facing the water and the tails facing away from it. The major structure in cell membranes is a double layer of these phospholipids, which is called a lipid bilayer
The lipid bilayer (or phospholipid bilayer) is a thin polar membrane made of two layers of lipid molecules. These membranes are flat sheets that form a continuous barrier around all cells. The cell membranes of almost all organisms and many vir ...
.
The phospholipids of archaea are unusual in four ways:
* They have membranes composed of glycerol-ether lipid
In organic chemistry, ethers are a class of compounds that contain an ether group—an oxygen atom connected to two alkyl or aryl groups. They have the general formula , where R and R′ represent the alkyl or aryl groups. Ethers can again be ...
s, whereas bacteria and eukaryotes have membranes composed mainly of glycerol-ester
In chemistry, an ester is a compound derived from an oxoacid (organic or inorganic) in which at least one hydroxyl group () is replaced by an alkoxy group (), as in the substitution reaction of a carboxylic acid and an alcohol. Glycerides a ...
lipid
Lipids are a broad group of naturally-occurring molecules which includes fats, waxes, sterols, fat-soluble vitamins (such as vitamins A, D, E and K), monoglycerides, diglycerides, phospholipids, and others. The functions of lipids includ ...
s. The difference is the type of bond that joins the lipids to the glycerol moiety; the two types are shown in yellow in the figure at the right. In ester lipids this is an ester bond
In chemistry, an ester is a compound derived from an oxoacid (organic or inorganic) in which at least one hydroxyl group () is replaced by an alkoxy group (), as in the substitution reaction of a carboxylic acid and an alcohol. Glycerides ar ...
, whereas in ether lipids this is an ether bond.
* The stereochemistry of the archaeal glycerol moiety is the mirror image of that found in other organisms. The glycerol moiety can occur in two forms that are mirror images of one another, called '' enantiomers''. Just as a right hand does not fit easily into a left-handed glove, enantiomers of one type generally cannot be used or made by enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. A ...
s adapted for the other. The archaeal phospholipids are built on a backbone of ''sn''-glycerol-1-phosphate, which is an enantiomer of ''sn''-glycerol-3-phosphate, the phospholipid backbone found in bacteria and eukaryotes. This suggests that archaea use entirely different enzymes for synthesizing phospholipids as compared to bacteria and eukaryotes. Such enzymes developed very early in life's history, indicating an early split from the other two domains.
* Archaeal lipid tails differ from those of other organisms in that they are based upon long isoprenoid chains with multiple side-branches, sometimes with cyclopropane or cyclohexane rings. By contrast, the fatty acid
In chemistry, particularly in biochemistry, a fatty acid is a carboxylic acid with an aliphatic chain, which is either saturated or unsaturated. Most naturally occurring fatty acids have an unbranched chain of an even number of carbon atoms, ...
s in the membranes of other organisms have straight chains without side branches or rings. Although isoprenoids play an important role in the biochemistry of many organisms, only the archaea use them to make phospholipids. These branched chains may help prevent archaeal membranes from leaking at high temperatures.
* In some archaea, the lipid bilayer is replaced by a monolayer. In effect, the archaea fuse the tails of two phospholipid molecules into a single molecule with two polar heads (a bolaamphiphile Bolaamphiphiles (also known as ''bolaform surfactants'',
''bolaphiles'', or ''alpha-omega-type surfactants'') are amphiphilic molecules that have hydrophilic groups at both ends of a sufficiently long hydrophobic hydrocarbon chain. Compared to singl ...
); this fusion may make their membranes more rigid and better able to resist harsh environments. For example, the lipids in ''Ferroplasma
''Ferroplasma'' is a genus of Archaea that belong to the family Ferroplasmaceae. Members of the ''Ferroplasma'' are typically acidophillic, pleomorphic, irregularly shaped cocci.
The archaean family Ferroplasmaceae was first described in the ea ...
'' are of this type, which is thought to aid this organism's survival in its highly acidic habitat.
Metabolism
Archaea exhibit a great variety of chemical reactions in theirmetabolism
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of life-sustaining chemical reactions in organisms. The three main functions of metabolism are: the conversion of the energy in food to energy available to run c ...
and use many sources of energy. These reactions are classified into nutritional groups, depending on energy and carbon sources. Some archaea obtain energy from inorganic compounds such as sulfur
Sulfur (or sulphur in British English) is a chemical element with the symbol S and atomic number 16. It is abundant, multivalent and nonmetallic. Under normal conditions, sulfur atoms form cyclic octatomic molecules with a chemical formula ...
or ammonia
Ammonia is an inorganic compound of nitrogen and hydrogen with the formula . A stable binary hydride, and the simplest pnictogen hydride, ammonia is a colourless gas with a distinct pungent smell. Biologically, it is a common nitrogenous wa ...
(they are chemotroph
A Chemotroph is an organism that obtains energy by the oxidation of electron donors in their environments. These molecules can be organic (chemoorganotrophs) or inorganic ( chemolithotrophs). The chemotroph designation is in contrast to phototr ...
s). These include nitrifiers, methanogens and anaerobic
Anaerobic means "living, active, occurring, or existing in the absence of free oxygen", as opposed to aerobic which means "living, active, or occurring only in the presence of oxygen." Anaerobic may also refer to:
* Anaerobic adhesive, a bonding a ...
methane
Methane ( , ) is a chemical compound with the chemical formula (one carbon atom bonded to four hydrogen atoms). It is a group-14 hydride, the simplest alkane, and the main constituent of natural gas. The relative abundance of methane on Ea ...
oxidiser
An oxidizing agent (also known as an oxidant, oxidizer, electron recipient, or electron acceptor) is a substance in a redox chemical reaction that gains or " accepts"/"receives" an electron from a (called the , , or ). In other words, an oxid ...
s. In these reactions one compound passes electrons to another (in a redox
Redox (reduction–oxidation, , ) is a type of chemical reaction in which the oxidation states of substrate change. Oxidation is the loss of electrons or an increase in the oxidation state, while reduction is the gain of electrons or a ...
reaction), releasing energy to fuel the cell's activities. One compound acts as an electron donor
In chemistry, an electron donor is a chemical entity that donates electrons to another compound. It is a reducing agent that, by virtue of its donating electrons, is itself oxidized in the process.
Typical reducing agents undergo permanent chemi ...
and one as an electron acceptor. The energy released is used to generate adenosine triphosphate
Adenosine triphosphate (ATP) is an organic compound that provides energy to drive many processes in living cells, such as muscle contraction, nerve impulse propagation, condensate dissolution, and chemical synthesis. Found in all known forms o ...
(ATP) through chemiosmosis
Chemiosmosis is the movement of ions across a semipermeable membrane bound structure, down their electrochemical gradient. An important example is the formation of adenosine triphosphate (ATP) by the movement of hydrogen ions (H+) across a memb ...
, the same basic process that happens in the mitochondrion of eukaryotic cells.
Other groups of archaea use sunlight as a source of energy (they are phototroph
Phototrophs () are organisms that carry out photon capture to produce complex organic compounds (e.g. carbohydrates) and acquire energy. They use the energy from light to carry out various cellular metabolic processes. It is a common misconcep ...
s), but oxygen–generating photosynthesis
Photosynthesis is a process used by plants and other organisms to convert light energy into chemical energy that, through cellular respiration, can later be released to fuel the organism's activities. Some of this chemical energy is stored i ...
does not occur in any of these organisms. Many basic metabolic pathway
In biochemistry, a metabolic pathway is a linked series of chemical reactions occurring within a cell. The reactants, products, and intermediates of an enzymatic reaction are known as metabolites, which are modified by a sequence of chemical reac ...
s are shared among all forms of life; for example, archaea use a modified form of glycolysis (the Entner–Doudoroff pathway
The Entner–Doudoroff pathway (ED Pathway) is a metabolic pathway that is most notable in Gram-negative bacteria, certain Gram-positive bacteria and archaea. Glucose is the substrate in the ED pathway and through a series of enzyme assisted chem ...
) and either a complete or partial citric acid cycle
The citric acid cycle (CAC)—also known as the Krebs cycle or the TCA cycle (tricarboxylic acid cycle)—is a series of chemical reactions to release stored energy through the oxidation of acetyl-CoA derived from carbohydrates, fats, and protein ...
. These similarities to other organisms probably reflect both early origins in the history of life and their high level of efficiency.
Some Euryarchaeota are methanogens (archaea that produce methane as a result of metabolism) living in anaerobic environment
Hypoxia refers to low oxygen conditions. Normally, 20.9% of the gas in the atmosphere is oxygen. The partial pressure of oxygen in the atmosphere is 20.9% of the total barometric pressure. In water, oxygen levels are much lower, approximately 7 p ...
s, such as swamps. This form of metabolism evolved early, and it is even possible that the first free-living organism was a methanogen. A common reaction involves the use of carbon dioxide
Carbon dioxide ( chemical formula ) is a chemical compound made up of molecules that each have one carbon atom covalently double bonded to two oxygen atoms. It is found in the gas state at room temperature. In the air, carbon dioxide is trans ...
as an electron acceptor to oxidize hydrogen
Hydrogen is the chemical element with the symbol H and atomic number 1. Hydrogen is the lightest element. At standard conditions hydrogen is a gas of diatomic molecules having the formula . It is colorless, odorless, tasteless, non-toxic ...
. Methanogenesis involves a range of coenzyme
A cofactor is a non-protein chemical compound or metallic ion that is required for an enzyme's role as a catalyst (a catalyst is a substance that increases the rate of a chemical reaction). Cofactors can be considered "helper molecules" that ass ...
s that are unique to these archaea, such as coenzyme M
Coenzyme M is a coenzyme required for methyl-transfer reactions in the metabolism of archaeal methanogens, and in the metabolism of other substrates in bacteria. It is also a necessary cofactor in the metabolic pathway of alkene-oxidizing bacteri ...
and methanofuran
Methanofurans are a family of chemical compounds found in methanogenic archaea. These species feature a 2-aminomethylfuran linked to phenoxy group. At least three different end groups are recognized: R = tricarboxyheptanoyl (methanofuran), gluta ...
. Other organic compounds such as alcohols, acetic acid
Acetic acid , systematically named ethanoic acid , is an acidic, colourless liquid and organic compound with the chemical formula (also written as , , or ). Vinegar is at least 4% acetic acid by volume, making acetic acid the main component ...
or formic acid are used as alternative electron acceptors by methanogens. These reactions are common in gut-dwelling archaea. Acetic acid
Acetic acid , systematically named ethanoic acid , is an acidic, colourless liquid and organic compound with the chemical formula (also written as , , or ). Vinegar is at least 4% acetic acid by volume, making acetic acid the main component ...
is also broken down into methane and carbon dioxide directly, by ''acetotrophic'' archaea. These acetotrophs are archaea in the order Methanosarcinales
In taxonomy, the Methanosarcinales are an order of the Methanomicrobia.
Large amounts of methane are produced in marine sediments but are then consumed before contacting aerobic waters or the atmosphere. Although no organism that can consume ...
, and are a major part of the communities of microorganisms that produce biogas
Biogas is a mixture of gases, primarily consisting of methane, carbon dioxide and hydrogen sulphide, produced from raw materials such as agricultural waste, manure, municipal waste, plant material, sewage, green waste and food waste. It is a ...
.
 Other archaea use in the atmosphere as a source of carbon, in a process called carbon fixation (they are
Other archaea use in the atmosphere as a source of carbon, in a process called carbon fixation (they are autotroph
An autotroph or primary producer is an organism that produces complex organic compounds (such as carbohydrates, fats, and proteins) using carbon from simple substances such as carbon dioxide,Morris, J. et al. (2019). "Biology: How Life Wo ...
s). This process involves either a highly modified form of the Calvin cycle or another metabolic pathway called the 3-hydroxypropionate/ 4-hydroxybutyrate cycle. The Thermoproteota also use the reverse Krebs cycle
The reverse Krebs cycle (also known as the reverse tricarboxylic acid cycle, the reverse TCA cycle, or the reverse citric acid cycle, or the reductive tricarboxylic acid cycle, or the reductive TCA cycle)
is a sequence of chemical reactions that ...
while the "Euryarchaeota" also use the reductive acetyl-CoA pathway
Reduction, reduced, or reduce may refer to:
Science and technology Chemistry
* Reduction (chemistry), part of a reduction-oxidation (redox) reaction in which atoms have their oxidation state changed.
** Organic redox reaction, a redox reacti ...
. Carbon fixation is powered by inorganic energy sources. No known archaea carry out photosynthesis
Photosynthesis is a process used by plants and other organisms to convert light energy into chemical energy that, through cellular respiration, can later be released to fuel the organism's activities. Some of this chemical energy is stored i ...
( Halobacterium is the only known phototroph archeon but it uses an alternative process to photosynthesis). Archaeal energy sources are extremely diverse, and range from the oxidation of ammonia
Ammonia is an inorganic compound of nitrogen and hydrogen with the formula . A stable binary hydride, and the simplest pnictogen hydride, ammonia is a colourless gas with a distinct pungent smell. Biologically, it is a common nitrogenous wa ...
by the Nitrosopumilales
The Nitrosopumilales are an order of the Archaea class Nitrososphaeria.
Phylogeny
Phylogeny of Nitrososphaerales
Taxonomy
The currently accepted taxonomy is based on the List of Prokaryotic names with Standing in Nomenclature (LPSN) and Nati ...
to the oxidation of hydrogen sulfide or elemental sulfur
Sulfur (or sulphur in British English) is a chemical element with the symbol S and atomic number 16. It is abundant, multivalent and nonmetallic. Under normal conditions, sulfur atoms form cyclic octatomic molecules with a chemical formula ...
by species of '' Sulfolobus'', using either oxygen or metal ions as electron acceptors.
Phototroph
Phototrophs () are organisms that carry out photon capture to produce complex organic compounds (e.g. carbohydrates) and acquire energy. They use the energy from light to carry out various cellular metabolic processes. It is a common misconcep ...
ic archaea use light to produce chemical energy in the form of ATP. In the Halobacteria
Haloarchaea (halophilic archaea, halophilic archaebacteria, halobacteria) are a class of the Euryarchaeota, found in water saturated or nearly saturated with salt. Halobacteria are now recognized as archaea rather than bacteria and are one of ...
, light-activated ion pumps like bacteriorhodopsin and halorhodopsin Halorhodopsin is a light-gated ion pump, specific for chloride ions, found in archaea, known as halobacteria. It is a seven-transmembrane retinylidene protein from microbial rhodopsin family. It is similar in tertiary structure (but not primary se ...
generate ion gradients by pumping ions out of and into the cell across the plasma membrane. The energy stored in these electrochemical gradient
An electrochemical gradient is a gradient of electrochemical potential, usually for an ion that can move across a membrane. The gradient consists of two parts, the chemical gradient, or difference in solute concentration across a membrane, and ...
s is then converted into ATP by ATP synthase. This process is a form of photophosphorylation In the process of photosynthesis, the phosphorylation of ADP to form ATP using the energy of sunlight is called photophosphorylation. Cyclic photophosphorylation occurs in both aerobic and anaerobic conditions, driven by the main primary source of ...
. The ability of these light-driven pumps to move ions across membranes depends on light-driven changes in the structure of a retinol
Retinol, also called vitamin A1, is a fat-soluble vitamin in the vitamin A family found in food and used as a dietary supplement. As a supplement it is used to treat and prevent vitamin A deficiency, especially that which results in xeroph ...
cofactor buried in the center of the protein.
Genetics
Archaea usually have a singlecircular chromosome
A circular chromosome is a chromosome in bacteria, archaea, Mitochondrial DNA#Genome structure and diversity, mitochondria, and Chloroplast DNA#Molecular structure, chloroplasts, in the form of a molecule of circular DNA, unlike the linear chromo ...
, with as many as 5,751,492 base pairs in ''Methanosarcina acetivorans
''Methanosarcina acetivorans'' is a versatile methane producing microbe which is found in such diverse environments as oil wells, trash dumps, deep-sea hydrothermal vents, and oxygen-depleted sediments beneath kelp beds. Only ''M. acetivorans ...
'', the largest known archaeal genome. The tiny 490,885 base-pair genome of ''Nanoarchaeum equitans
''Nanoarchaeum equitans'' is a species of marine archaea that was discovered in 2002 in a hydrothermal vent off the coast of Iceland on the Kolbeinsey Ridge by Karl Stetter. It has been proposed as the first species in a new phylum. Strains of ...
'' is one-tenth of this size and the smallest archaeal genome known; it is estimated to contain only 537 protein-encoding genes. Smaller independent pieces of DNA, called '' plasmids'', are also found in archaea. Plasmids may be transferred between cells by physical contact, in a process that may be similar to bacterial conjugation.
 Archaea are genetically distinct from bacteria and eukaryotes, with up to 15% of the proteins encoded by any one archaeal genome being unique to the domain, although most of these unique genes have no known function. Of the remainder of the unique proteins that have an identified function, most belong to the Euryarchaeota and are involved in methanogenesis. The proteins that archaea, bacteria and eukaryotes share form a common core of cell function, relating mostly to
Archaea are genetically distinct from bacteria and eukaryotes, with up to 15% of the proteins encoded by any one archaeal genome being unique to the domain, although most of these unique genes have no known function. Of the remainder of the unique proteins that have an identified function, most belong to the Euryarchaeota and are involved in methanogenesis. The proteins that archaea, bacteria and eukaryotes share form a common core of cell function, relating mostly to transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
, translation
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transla ...
, and nucleotide metabolism. Other characteristic archaeal features are the organization of genes of related function – such as enzymes that catalyze steps in the same metabolic pathway
In biochemistry, a metabolic pathway is a linked series of chemical reactions occurring within a cell. The reactants, products, and intermediates of an enzymatic reaction are known as metabolites, which are modified by a sequence of chemical reac ...
into novel operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
s, and large differences in tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ...
genes and their aminoacyl tRNA synthetase
An aminoacyl-tRNA synthetase (aaRS or ARS), also called tRNA-ligase, is an enzyme that attaches the appropriate amino acid onto its corresponding tRNA. It does so by catalyzing the transesterification of a specific cognate amino acid or its pre ...
s.
Transcription in archaea more closely resembles eukaryotic than bacterial transcription, with the archaeal RNA polymerase being very close to its equivalent in eukaryotes, while archaeal translation shows signs of both bacterial and eukaryotic equivalents. Although archaea have only one type of RNA polymerase, its structure and function in transcription seems to be close to that of the eukaryotic RNA polymerase II
RNA polymerase II (RNAP II and Pol II) is a multiprotein complex that transcribes DNA into precursors of messenger RNA (mRNA) and most small nuclear RNA (snRNA) and microRNA. It is one of the three RNAP enzymes found in the nucleus of eukaryo ...
, with similar protein assemblies (the general transcription factor
General transcription factors (GTFs), also known as basal transcriptional factors, are a class of protein transcription factors that bind to specific sites ( promoter) on DNA to activate transcription of genetic information from DNA to messenger ...
s) directing the binding of the RNA polymerase to a gene's promoter, but other archaeal transcription factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. The fu ...
s are closer to those found in bacteria. Post-transcriptional modification
Transcriptional modification or co-transcriptional modification is a set of biological processes common to most eukaryotic cells by which an RNA primary transcript is chemically altered following transcription from a gene to produce a mature, fu ...
is simpler than in eukaryotes, since most archaeal genes lack introns, although there are many introns in their transfer RNA and ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosom ...
genes, and introns may occur in a few protein-encoding genes.
Gene transfer and genetic exchange
''Haloferax volcanii
''Haloferax volcanii'' is a species of organism in the genus ''Haloferax'' in the Archaea.
Description and significance
Microbiologist Benjamin Elazari Volcani first discovered ''Haloferax volcanii'', a self-named extremophile, in the 1930s. '' ...
'', an extreme halophilic archaeon, forms cytoplasmic bridges between cells that appear to be used for transfer of DNA from one cell to another in either direction.
When the hyperthermophilic archaea ''Sulfolobus solfataricus
''Saccharolobus solfataricus'' is a species of thermophilic archaeon. It was transferred from the genus ''Sulfolobus'' to the new genus ''Saccharolobus'' with the description of Saccharolobus caldissimus in 2018.
It was first isolated and disco ...
'' and ''Sulfolobus acidocaldarius
''Sulfolobus acidocaldarius'' is a thermoacidophilic archaeon that belongs to the phylum Thermoproteota. ''S. acidocaldarius'' was the first ''Sulfolobus'' species to be described, in 1972 by Thomas D. Brock and collaborators. This species was ...
'' are exposed to DNA-damaging UV irradiation or to the agents bleomycin
-13- (1''H''-imidazol-5-yl)methyl9-hydroxy-5- 1''R'')-1-hydroxyethyl8,10-dimethyl-4,7,12,15-tetraoxo-3,6,11,14-tetraazapentadec-1-yl}-2,4'-bi-1,3-thiazol-4-yl)carbonyl]amino}propyl)(dimethyl)sulfonium
, chemical_formula =
, C=55 , H=84 , N=1 ...
or mitomycin C
Mitomycin C is a mitomycin that is used as a chemotherapeutic agent by virtue of its antitumour activity.
Medical uses
It is given intravenously to treat upper gastro-intestinal cancers (e.g. esophageal carcinoma), anal cancers, and breast ...
, species-specific cellular aggregation is induced. Aggregation in ''S. solfataricus'' could not be induced by other physical stressors, such as pH or temperature shift, suggesting that aggregation is induced specifically by DNA damage
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA d ...
. Ajon et al. showed that UV-induced cellular aggregation mediates chromosomal marker exchange with high frequency in ''S. acidocaldarius''. Recombination rates exceeded those of uninduced cultures by up to three orders of magnitude. Frols et al. and Ajon et al. hypothesized that cellular aggregation enhances species-specific DNA transfer between ''Sulfolobus'' cells in order to provide increased repair of damaged DNA by means of homologous recombination
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids (usually DNA as in cellular organisms but may ...
. This response may be a primitive form of sexual interaction similar to the more well-studied bacterial transformation systems that are also associated with species-specific DNA transfer between cells leading to homologous recombinational repair of DNA damage.
Archaeal viruses
Archaea are the target of a number ofvirus
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsk ...
es in a diverse virosphere distinct from bacterial and eukaryotic viruses. They have been organized into 15-18 DNA-based families so far, but multiple species remain un-isolated and await classification. These families can be informally divided into two groups: archaea-specific and cosmopolitan. Archaeal-specific viruses target only archaean species and currently include 12 families. Numerous unique, previously unidentified viral structures have been observed in this group, including: bottle-shaped, spindle-shaped, coil-shaped, and droplet-shaped viruses. While the reproductive cycles and genomic mechanisms of archaea-specific species may be similar to other viruses, they bear unique characteristics that were specifically developed due to the morphology of host cells they infect. Their virus release mechanisms differ from that of other phages. Bacteriophages generally undergo either lytic
The lytic cycle ( ) is one of the two cycles of viral reproduction (referring to bacterial viruses or bacteriophages), the other being the lysogenic cycle. The lytic cycle results in the destruction of the infected cell and its membrane. Bacter ...
pathways, lysogenic
Lysogeny, or the lysogenic cycle, is one of two cycles of viral reproduction (the lytic cycle being the other). Lysogeny is characterized by integration of the bacteriophage nucleic acid into the host bacterium's genome or formation of a circu ...
pathways, or (rarely) a mix of the two. Most archaea-specific viral strains maintain a stable, somewhat lysogenic, relationship with their hosts – appearing as a chronic infection. This involves the gradual, and continuous, production and release of virions
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsky's ...
without killing the host cell. Prangishyili (2013) noted that it has been hypothesized that tailed archaeal phages originated from bacteriophages capable of infecting haloarchaea
Haloarchaea (halophilic archaea, halophilic archaebacteria, halobacteria) are a class of the Euryarchaeota, found in water saturated or nearly saturated with salt. Halobacteria are now recognized as archaea rather than bacteria and are one of t ...
l species. If the hypothesis is correct, it can be concluded that other double-stranded DNA viruses
A DNA virus is a virus that has a genome made of deoxyribonucleic acid (DNA) that is replicated by a DNA polymerase. They can be divided between those that have two strands of DNA in their genome, called double-stranded DNA (dsDNA) viruses, and ...
that make up the rest of the archaea-specific group are their own unique group in the global viral community. Krupovic et al. (2018) states that the high levels of horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). H ...
, rapid mutation rates in viral genomes, and lack of universal gene sequences have led researchers to perceive the evolutionary pathway of archaeal viruses as a network. The lack of similarities among phylogenetic markers in this network and the global virosphere, as well as external linkages to non-viral elements, may suggest that some species of archaea specific viruses evolved from non-viral mobile genetic elements
Mobile genetic elements (MGEs) sometimes called selfish genetic elements are a type of genetic material that can move around within a genome, or that can be transferred from one species or replicon to another. MGEs are found in all organisms. In ...
(MGE).
These viruses have been studied in most detail in thermophilics, particularly the orders Sulfolobales and Thermoproteales
In taxonomy
Taxonomy is the practice and science of categorization or classification.
A taxonomy (or taxonomical classification) is a scheme of classification, especially a hierarchical classification, in which things are organized into g ...
. Two groups of single-stranded DNA viruses
A DNA virus is a virus that has a genome made of deoxyribonucleic acid (DNA) that is replicated by a DNA polymerase. They can be divided between those that have two strands of DNA in their genome, called double-stranded DNA (dsDNA) viruses, and ...
that infect archaea have been recently isolated. One group is exemplified by the '' Halorubrum pleomorphic virus 1'' (''Pleolipoviridae
''Pleolipoviridae'' is a family of DNA virus
A DNA virus is a virus that has a genome made of deoxyribonucleic acid (DNA) that is replicated by a DNA polymerase. They can be divided between those that have two strands of DNA in their genome, ...
'') infecting halophilic archaea, and the other one by the '' Aeropyrum coil-shaped virus'' (''Spiraviridae
''Spiraviridae'' is a family of viruses that replicate in hyperthermophilic archaea of the genus ''Aeropyrum'', specifically '' Aeropyrum pernix''. The family contains one genus, ''Alphaspiravirus'', which contains one species, ''Aeropyrum coil-s ...
'') infecting a hyperthermophilic (optimal growth at 90–95 °C) host. Notably, the latter virus has the largest currently reported ssDNA genome. Defenses against these viruses may involve RNA interference
RNA interference (RNAi) is a biological process in which RNA molecules are involved in sequence-specific suppression of gene expression by double-stranded RNA, through translational or transcriptional repression. Historically, RNAi was known by ...
from repetitive DNA
Repeated sequences (also known as repetitive elements, repeating units or repeats) are short or long patterns of nucleic acids (DNA or RNA) that occur in multiple copies throughout the genome. In many organisms, a significant fraction of the geno ...
sequences that are related to the genes of the viruses.
Reproduction
Archaea reproduce asexually by binary or multiple fission, fragmentation, orbudding
Budding or blastogenesis is a type of asexual reproduction in which a new organism develops from an outgrowth or bud due to cell division at one particular site. For example, the small bulb-like projection coming out from the yeast cell is kno ...
; mitosis and meiosis
Meiosis (; , since it is a reductional division) is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately r ...
do not occur, so if a species of archaea exists in more than one form, all have the same genetic material. Cell division
Cell division is the process by which a parent cell divides into two daughter cells. Cell division usually occurs as part of a larger cell cycle in which the cell grows and replicates its chromosome(s) before dividing. In eukaryotes, there ar ...
is controlled in a cell cycle
The cell cycle, or cell-division cycle, is the series of events that take place in a cell that cause it to divide into two daughter cells. These events include the duplication of its DNA (DNA replication) and some of its organelles, and sub ...
; after the cell's chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
is replicated and the two daughter chromosomes separate, the cell divides. In the genus '' Sulfolobus'', the cycle has characteristics that are similar to both bacterial and eukaryotic systems. The chromosomes replicate from multiple starting points ( origins of replication) using DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create ...
s that resemble the equivalent eukaryotic enzymes.
In Euryarchaeota the cell division protein FtsZ
FtsZ is a protein encoded by the ''ftsZ'' gene that assembles into a ring at the future site of bacterial cell division (also called the Z ring). FtsZ is a prokaryotic homologue of the eukaryotic protein tubulin. The initials FtsZ mean "Filam ...
, which forms a contracting ring around the cell, and the components of the septum that is constructed across the center of the cell, are similar to their bacterial equivalents. In cren- and thaumarchaea, the cell division machinery Cdv fulfills a similar role. This machinery is related to the eukaryotic ESCRT-III machinery which, while best known for its role in cell sorting, also has been seen to fulfill a role in separation between divided cell, suggesting an ancestral role in cell division.
Both bacteria and eukaryotes, but not archaea, make spores. Some species of Haloarchaea
Haloarchaea (halophilic archaea, halophilic archaebacteria, halobacteria) are a class of the Euryarchaeota, found in water saturated or nearly saturated with salt. Halobacteria are now recognized as archaea rather than bacteria and are one of t ...
undergo phenotypic switching Phenotypic switching is switching between multiple cellular morphologies. David R. Soll described two such systems: the first high frequency switching system between several morphological stages and a second high frequency switching system between ...
and grow as several different cell types, including thick-walled structures that are resistant to osmotic shock
Osmotic shock or osmotic stress is physiologic dysfunction caused by a sudden change in the solute concentration around a cell, which causes a rapid change in the movement of water across its cell membrane. Under hypertonic conditions - conditions ...
and allow the archaea to survive in water at low salt concentrations, but these are not reproductive structures and may instead help them reach new habitats.
Behavior
Communication
Quorum sensing
In biology, quorum sensing or quorum signalling (QS) is the ability to detect and respond to cell population density by gene regulation. As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at ...
was originally thought to not exist in Archaea, but recent studies have shown evidence of some species being able to perform cross-talk through quorum sensing. Other studies have shown syntrophic interactions between archaea and bacteria during biofilm growth. Although research is limited in archaeal quorum sensing, some studies have uncovered LuxR proteins in archaeal species, displaying similarities with bacteria LuxR, and ultimately allowing for the detection of small molecules that are used in high density communication. Similarly to bacteria, Archaea LuxR solos have shown to bind to AHLs (lactones) and non-AHLs ligans, which is a large part in performing intraspecies, interspecies, and interkingdom communication through quorum sensing.
Ecology
Habitats
 Archaea exist in a broad range of
Archaea exist in a broad range of habitat
In ecology, the term habitat summarises the array of resources, physical and biotic factors that are present in an area, such as to support the survival and reproduction of a particular species. A species habitat can be seen as the physical ...
s, and are now recognized as a major part of global ecosystem
An ecosystem (or ecological system) consists of all the organisms and the physical environment with which they interact. These biotic and abiotic components are linked together through nutrient cycles and energy flows. Energy enters the syste ...
s, and may represent about 20% of microbial cells in the oceans. However, the first-discovered archaeans were extremophile
An extremophile (from Latin ' meaning "extreme" and Greek ' () meaning "love") is an organism that is able to live (or in some cases thrive) in extreme environments, i.e. environments that make survival challenging such as due to extreme temper ...
s. Indeed, some archaea survive high temperatures, often above , as found in geysers, black smoker
A hydrothermal vent is a fissure on the seabed from which geothermally heated water discharges. They are commonly found near volcanically active places, areas where tectonic plates are moving apart at mid-ocean ridges, ocean basins, and hotspot ...
s, and oil wells. Other common habitats include very cold habitats and highly saline, acidic, or alkaline water, but archaea include mesophiles that grow in mild conditions, in swamps and marsh
A marsh is a wetland that is dominated by herbaceous rather than woody plant species.Keddy, P.A. 2010. Wetland Ecology: Principles and Conservation (2nd edition). Cambridge University Press, Cambridge, UK. 497 p Marshes can often be found a ...
land, sewage, the ocean
The ocean (also the sea or the world ocean) is the body of salt water that covers approximately 70.8% of the surface of Earth and contains 97% of Earth's water. An ocean can also refer to any of the large bodies of water into which the wo ...
s, the intestinal tract
The gastrointestinal tract (GI tract, digestive tract, alimentary canal) is the tract or passageway of the digestive system that leads from the mouth to the anus. The GI tract contains all the major organs of the digestive system, in humans ...
of animals, and soil
Soil, also commonly referred to as earth or dirt
Dirt is an unclean matter, especially when in contact with a person's clothes, skin, or possessions. In such cases, they are said to become dirty.
Common types of dirt include:
* Debri ...
s. Similar to PGPR, Archaea are now considered as a source of plant growth promotion as well.
Extremophile archaea are members of four main physiological groups. These are the halophile
The halophiles, named after the Greek word for "salt-loving", are extremophiles that thrive in high salt concentrations. While most halophiles are classified into the domain Archaea, there are also bacterial halophiles and some eukaryotic species, ...
s, thermophile
A thermophile is an organism—a type of extremophile—that thrives at relatively high temperatures, between . Many thermophiles are archaea, though they can be bacteria or fungi. Thermophilic eubacteria are suggested to have been among the earl ...
s, alkaliphile Alkaliphiles are a class of extremophilic microbes capable of survival in alkaline ( pH roughly 8.5–11) environments, growing optimally around a pH of 10. These bacteria can be further categorized as obligate alkaliphiles (those that require hig ...
s, and acidophiles
Acidophiles or acidophilic organisms are those that thrive under highly acidic conditions (usually at pH 5.0 or below). These organisms can be found in different branches of the tree of life, including Archaea, Bacteria,Becker, A.Types of Bacte ...
. These groups are not comprehensive or phylum-specific, nor are they mutually exclusive, since some archaea belong to several groups. Nonetheless, they are a useful starting point for classification.
Halophiles, including the genus '' Halobacterium'', live in extremely saline environments such as salt lakes and outnumber their bacterial counterparts at salinities greater than 20–25%. Thermophiles grow best at temperatures above , in places such as hot springs; ''hyperthermophilic'' archaea grow optimally at temperatures greater than . The archaeal ''Methanopyrus kandleri
In taxonomy, ''Methanopyrus'' is a genus of the Methanopyraceae.
''Methanopyrus'' is a genus of methanogen, with a single described species, ''M. kandleri''. It is a rod-shaped hyperthermophile, discovered on the wall of a black smoker from the ...
'' Strain 116 can even reproduce at , the highest recorded temperature of any organism.
Other archaea exist in very acidic or alkaline conditions. For example, one of the most extreme archaean acidophiles is '' Picrophilus torridus'', which grows at pH 0, which is equivalent to thriving in 1.2 molar sulfuric acid.
This resistance to extreme environments has made archaea the focus of speculation about the possible properties of extraterrestrial life. Some extremophile habitats are not dissimilar to those on Mars
Mars is the fourth planet from the Sun and the second-smallest planet in the Solar System, only being larger than Mercury. In the English language, Mars is named for the Roman god of war. Mars is a terrestrial planet with a thin at ...
, leading to the suggestion that viable microbes could be transferred between planets in meteorites.
Recently, several studies have shown that archaea exist not only in mesophilic and thermophilic environments but are also present, sometimes in high numbers, at low temperatures as well. For example, archaea are common in cold oceanic environments such as polar seas. Even more significant are the large numbers of archaea found throughout the world's oceans in non-extreme habitats among the plankton
Plankton are the diverse collection of organisms found in water (or air) that are unable to propel themselves against a current (or wind). The individual organisms constituting plankton are called plankters. In the ocean, they provide a crucia ...
community (as part of the picoplankton
Picoplankton is the fraction of plankton composed by cells between 0.2 and 2 μm that can be either prokaryotic and eukaryotic phototrophs and heterotrophs:
* photosynthetic
* heterotrophic
They are prevalent amongst microbial plankton communit ...
). Although these archaea can be present in extremely high numbers (up to 40% of the microbial biomass), almost none of these species have been isolated and studied in pure culture
A microbiological culture, or microbial culture, is a method of multiplying microbial organisms by letting them reproduce in predetermined culture medium under controlled laboratory conditions. Microbial cultures are foundational and basic diagn ...
. Consequently, our understanding of the role of archaea in ocean ecology is rudimentary, so their full influence on global biogeochemical
Biogeochemistry is the scientific discipline that involves the study of the chemical, physical, geological, and biological processes and reactions that govern the composition of the natural environment (including the biosphere, the cryosphere, th ...
cycles remains largely unexplored. Some marine Thermoproteota are capable of nitrification
''Nitrification'' is the biological oxidation of ammonia to nitrite followed by the oxidation of the nitrite to nitrate occurring through separate organisms or direct ammonia oxidation to nitrate in comammox bacteria. The transformation of am ...
, suggesting these organisms may affect the oceanic nitrogen cycle, although these oceanic Thermoproteota may also use other sources of energy.
Vast numbers of archaea are also found in the sediment
Sediment is a naturally occurring material that is broken down by processes of weathering and erosion, and is subsequently transported by the action of wind, water, or ice or by the force of gravity acting on the particles. For example, sa ...
s that cover the sea floor, with these organisms making up the majority of living cells at depths over 1 meter below the ocean bottom. It has been demonstrated that in all oceanic surface sediments (from 1000- to 10,000-m water depth), the impact of viral infection is higher on archaea than on bacteria and virus-induced lysis of archaea accounts for up to one-third of the total microbial biomass killed, resulting in the release of ~0.3 to 0.5 gigatons of carbon per year globally.
Role in chemical cycling
Archaea recycle elements such ascarbon
Carbon () is a chemical element with the symbol C and atomic number 6. It is nonmetallic and tetravalent—its atom making four electrons available to form covalent chemical bonds. It belongs to group 14 of the periodic table. Carbon mak ...
, nitrogen
Nitrogen is the chemical element with the symbol N and atomic number 7. Nitrogen is a nonmetal and the lightest member of group 15 of the periodic table, often called the pnictogens. It is a common element in the universe, estimated at se ...
, and sulfur
Sulfur (or sulphur in British English) is a chemical element with the symbol S and atomic number 16. It is abundant, multivalent and nonmetallic. Under normal conditions, sulfur atoms form cyclic octatomic molecules with a chemical formula ...
through their various habitats. Archaea carry out many steps in the nitrogen cycle. This includes both reactions that remove nitrogen from ecosystems (such as nitrate-based respiration and denitrification
Denitrification is a microbially facilitated process where nitrate (NO3−) is reduced and ultimately produces molecular nitrogen (N2) through a series of intermediate gaseous nitrogen oxide products. Facultative anaerobic bacteria perform denitr ...
) as well as processes that introduce nitrogen (such as nitrate assimilation and nitrogen fixation
Nitrogen fixation is a chemical process by which molecular nitrogen (), with a strong triple covalent bond, in the air is converted into ammonia () or related nitrogenous compounds, typically in soil or aquatic systems but also in industry. Atmo ...
).
Researchers recently discovered archaeal involvement in ammonia
Ammonia is an inorganic compound of nitrogen and hydrogen with the formula . A stable binary hydride, and the simplest pnictogen hydride, ammonia is a colourless gas with a distinct pungent smell. Biologically, it is a common nitrogenous wa ...
oxidation reactions. These reactions are particularly important in the oceans. The archaea also appear crucial for ammonia oxidation in soils. They produce nitrite, which other microbes then oxidize to nitrate. Plants and other organisms consume the latter.
In the sulfur cycle, archaea that grow by oxidizing sulfur
Sulfur (or sulphur in British English) is a chemical element with the symbol S and atomic number 16. It is abundant, multivalent and nonmetallic. Under normal conditions, sulfur atoms form cyclic octatomic molecules with a chemical formula ...
compounds release this element from rocks, making it available to other organisms, but the archaea that do this, such as ''Sulfolobus'', produce sulfuric acid as a waste product, and the growth of these organisms in abandoned mines can contribute to acid mine drainage
Acid mine drainage, acid and metalliferous drainage (AMD), or acid rock drainage (ARD) is the outflow of acidic water from metal mines or coal mines.
Acid rock drainage occurs naturally within some environments as part of the rock weathering ...
and other environmental damage.
In the carbon cycle
The carbon cycle is the biogeochemical cycle by which carbon is exchanged among the biosphere, pedosphere, geosphere, hydrosphere, and atmosphere of the Earth. Carbon is the main component of biological compounds as well as a major componen ...
, methanogen archaea remove hydrogen and play an important role in the decay of organic matter by the populations of microorganisms that act as decomposer
Decomposers are organisms that break down dead or decaying organisms; they carry out decomposition, a process possible by only certain kingdoms, such as fungi. Like herbivores and predators, decomposers are heterotrophic, meaning that they use o ...
s in anaerobic ecosystems, such as sediments, marshes, and sewage-treatment works.
Interactions with other organisms
 The well-characterized interactions between archaea and other organisms are either mutual or
The well-characterized interactions between archaea and other organisms are either mutual or commensal
Commensalism is a long-term biological interaction (symbiosis) in which members of one species gain benefits while those of the other species neither benefit nor are harmed. This is in contrast with mutualism, in which both organisms benefit fro ...
. There are no clear examples of known archaeal pathogen
In biology, a pathogen ( el, πάθος, "suffering", "passion" and , "producer of") in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ ...
s or parasite
Parasitism is a close relationship between species, where one organism, the parasite, lives on or inside another organism, the host, causing it some harm, and is adapted structurally to this way of life. The entomologist E. O. Wilson has ...
s, but some species of methanogens have been suggested to be involved in infections in the mouth, and ''Nanoarchaeum equitans
''Nanoarchaeum equitans'' is a species of marine archaea that was discovered in 2002 in a hydrothermal vent off the coast of Iceland on the Kolbeinsey Ridge by Karl Stetter. It has been proposed as the first species in a new phylum. Strains of ...
'' may be a parasite of another species of archaea, since it only survives and reproduces within the cells of the Crenarchaeon '' Ignicoccus hospitalis'', and appears to offer no benefit to its host
A host is a person responsible for guests at an event or for providing hospitality during it.
Host may also refer to:
Places
* Host, Pennsylvania, a village in Berks County
People
*Jim Host (born 1937), American businessman
* Michel Host ...
.
Mutualism
Mutualism is an interaction between individuals of different species that results in positive (beneficial) effects on per capita reproduction and/or survival of the interacting populations. One well-understood example of mutualism is the interaction between protozoa and methanogenic archaea in the digestive tracts of animals that digestcellulose
Cellulose is an organic compound with the formula , a polysaccharide consisting of a linear chain of several hundred to many thousands of β(1→4) linked D-glucose units. Cellulose is an important structural component of the primary cell w ...
, such as ruminant
Ruminants (suborder Ruminantia) are hoofed herbivorous grazing or browsing mammals that are able to acquire nutrients from plant-based food by fermenting it in a specialized stomach prior to digestion, principally through microbial actions. The ...
s and termite
Termites are small insects that live in colonies and have distinct castes (eusocial) and feed on wood or other dead plant matter. Termites comprise the infraorder Isoptera, or alternatively the epifamily Termitoidae, within the order Blatto ...
s. In these anaerobic environments, protozoa break down plant cellulose to obtain energy. This process releases hydrogen as a waste product, but high levels of hydrogen reduce energy production. When methanogens convert hydrogen to methane, protozoa benefit from more energy.
In anaerobic protozoa, such as ''Plagiopyla frontata'', archaea reside inside the protozoa and consume hydrogen produced in their hydrogenosome
A hydrogenosome is a membrane-enclosed organelle found in some anaerobic ciliates, flagellates, and fungi. Hydrogenosomes are highly variable organelles that have presumably evolved from protomitochondria to produce molecular hydrogen and ATP i ...
s. Archaea also associate with larger organisms. For example, the marine archaean ''Cenarchaeum symbiosum
''Cenarchaeum symbiosum'' is a species of Archaea in the genus ''Cenarchaeum'', in the phylum Nitrososphaerota in the domain Archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell ...
'' lives within (is an endosymbiont
An ''endosymbiont'' or ''endobiont'' is any organism that lives within the body or cells of another organism most often, though not always, in a mutualistic relationship.
(The term endosymbiosis is from the Greek: ἔνδον ''endon'' "within ...
of) the sponge
Sponges, the members of the phylum Porifera (; meaning 'pore bearer'), are a basal animal clade as a sister of the diploblasts. They are multicellular organisms that have bodies full of pores and channels allowing water to circulate throug ...
'' Axinella mexicana''.
Commensalism
Archaea can also be commensals, benefiting from an association without helping or harming the other organism. For example, the methanogen ''Methanobrevibacter smithii
''Methanobrevibacter smithii'' is the predominant archaeon in the microbiota of the human gut. ''M. smithii'' has a coccobacillus shape. It plays an important role in the efficient digestion of polysaccharides (complex sugars) by consuming the ...
'' is by far the most common archaean in the human flora, making up about one in ten of all the prokaryotes in the human gut. In termites and in humans, these methanogens may in fact be mutualists, interacting with other microbes in the gut to aid digestion. Archaean communities also associate with a range of other organisms, such as on the surface of coral
Corals are marine invertebrates within the class Anthozoa of the phylum Cnidaria. They typically form compact colonies of many identical individual polyps. Coral species include the important reef builders that inhabit tropical oceans and ...
s, and in the region of soil that surrounds plant roots (the rhizosphere
The rhizosphere is the narrow region of soil or substrate that is directly influenced by root secretions and associated soil microorganisms known as the root microbiome. Soil pores in the rhizosphere can contain many bacteria and other microo ...
).
Parasitism
Although Archaea do not have a historical reputation of being pathogens, Archaea are often found with similar genomes to more common pathogen like ''E. coli,'' showing metabolic links and evolutionary history with today's pathogens. Archaea also has inconsistent detection within clinical studies because of the lack of categorization of Archaea into more specific species.Significance in technology and industry
Extremophile
An extremophile (from Latin ' meaning "extreme" and Greek ' () meaning "love") is an organism that is able to live (or in some cases thrive) in extreme environments, i.e. environments that make survival challenging such as due to extreme temper ...
archaea, particularly those resistant either to heat or to extremes of acidity and alkalinity, are a source of enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. A ...
s that function under these harsh conditions. These enzymes have found many uses. For example, thermostable DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create ...
s, such as the Pfu DNA polymerase
''Pfu'' DNA polymerase is an enzyme found in the hyperthermophilic archaeon ''Pyrococcus furiosus'', where it functions to copy the organism's DNA during cell division. In the laboratory setting, ''Pfu'' is used to amplify DNA in the polymeras ...
from ''Pyrococcus furiosus
''Pyrococcus furiosus'' is a heterotrophic, strictly anaerobic, extremophilic, model species of archaea. It is classified as a hyperthermophile because it thrives best under extremely high temperatures, and is notable for having an optimum gr ...
'', revolutionized molecular biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physi ...
by allowing the polymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) ...
to be used in research as a simple and rapid technique for cloning
Cloning is the process of producing individual organisms with identical or virtually identical DNA, either by natural or artificial means. In nature, some organisms produce clones through asexual reproduction. In the field of biotechnology, c ...
DNA. In industry, amylase
An amylase () is an enzyme that catalyses the hydrolysis of starch (Latin ') into sugars. Amylase is present in the saliva of humans and some other mammals, where it begins the chemical process of digestion. Foods that contain large amounts of ...
s, galactosidases and pullulanase
Pullulanase (, ''limit dextrinase'', ''amylopectin 6-glucanohydrolase'', ''bacterial debranching enzyme'', ''debranching enzyme'', ''alpha-dextrin endo-1,6-alpha-glucosidase'', ''R-enzyme'', ''pullulan alpha-1,6-glucanohydrolase'') is a specific ki ...
s in other species of '' Pyrococcus'' that function at over allow food processing at high temperatures, such as the production of low lactose milk and whey
Whey is the liquid remaining after milk has been curdled and strained. It is a byproduct of the manufacturing of cheese or casein and has several commercial uses. Sweet whey is a byproduct resulting from the manufacture of rennet types of har ...
. Enzymes from these thermophilic archaea also tend to be very stable in organic solvents, allowing their use in environmentally friendly processes in green chemistry
Green chemistry, also called sustainable chemistry, is an area of chemistry and chemical engineering focused on the design of products and processes that minimize or eliminate the use and generation of hazardous substances. While environmental che ...
that synthesize organic compounds. This stability makes them easier to use in structural biology
Structural biology is a field that is many centuries old which, and as defined by the Journal of Structural Biology, deals with structural analysis of living material (formed, composed of, and/or maintained and refined by living cells) at every le ...
. Consequently, the counterparts of bacterial or eukaryotic enzymes from extremophile archaea are often used in structural studies.
In contrast with the range of applications of archaean enzymes, the use of the organisms themselves in biotechnology is less developed. Methanogenic archaea are a vital part of sewage treatment
Sewage treatment (or domestic wastewater treatment, municipal wastewater treatment) is a type of wastewater treatment which aims to remove contaminants from sewage to produce an effluent that is suitable for discharge to the surrounding e ...
, since they are part of the community of microorganisms that carry out anaerobic digestion
Anaerobic digestion is a sequence of processes by which microorganisms break down biodegradable material in the absence of oxygen. The process is used for industrial or domestic purposes to Waste management, manage waste or to produce fuels. Mu ...
and produce biogas
Biogas is a mixture of gases, primarily consisting of methane, carbon dioxide and hydrogen sulphide, produced from raw materials such as agricultural waste, manure, municipal waste, plant material, sewage, green waste and food waste. It is a ...
. In mineral processing
In the field of extractive metallurgy, mineral processing, also known as ore dressing, is the process of separating commercially valuable minerals from their ores.
History
Before the advent of heavy machinery the raw ore was broken up using ...
, acidophilic archaea display promise for the extraction of metals from ore
Ore is natural rock or sediment that contains one or more valuable minerals, typically containing metals, that can be mined, treated and sold at a profit.Encyclopædia Britannica. "Ore". Encyclopædia Britannica Online. Retrieved 7 Apr ...
s, including gold
Gold is a chemical element with the symbol Au (from la, aurum) and atomic number 79. This makes it one of the higher atomic number elements that occur naturally. It is a bright, slightly orange-yellow, dense, soft, malleable, and ductile me ...
, cobalt
Cobalt is a chemical element with the symbol Co and atomic number 27. As with nickel, cobalt is found in the Earth's crust only in a chemically combined form, save for small deposits found in alloys of natural meteoric iron. The free element, p ...
and copper
Copper is a chemical element with the symbol Cu (from la, cuprum) and atomic number 29. It is a soft, malleable, and ductile metal with very high thermal and electrical conductivity. A freshly exposed surface of pure copper has a pinkis ...
.
Archaea host a new class of potentially useful antibiotics. A few of these archaeocins have been characterized, but hundreds more are believed to exist, especially within ''Haloarchaea
Haloarchaea (halophilic archaea, halophilic archaebacteria, halobacteria) are a class of the Euryarchaeota, found in water saturated or nearly saturated with salt. Halobacteria are now recognized as archaea rather than bacteria and are one of t ...
'' and '' Sulfolobus''. These compounds differ in structure from bacterial antibiotics, so they may have novel modes of action. In addition, they may allow the creation of new selectable marker A selectable marker is a gene introduced into a cell, especially a bacterium or to cells in culture, that confers a trait suitable for artificial selection. They are a type of reporter gene used in laboratory microbiology, molecular biology, a ...
s for use in archaeal molecular biology.
See also
*Aerobic methane production
Aerobic methane production is a potential biological pathway for atmospheric methane (CH4) production under oxygenated conditions. The existence of this pathway was first theorized in 2006. While significant evidence suggests the existence of th ...
* Earliest known life forms
The earliest known life forms on Earth are believed to be fossilized microorganisms found in hydrothermal vent precipitates, considered to be about 3.42 billion years old. The earliest time for the origin of life on Earth is at least 3.77 bill ...
* List of Archaea genera
* List of sequenced archaeal genomes
This list of sequenced archaeal genomes contains all the archaea known to have publicly available complete genome sequences that have been assembled, annotated and deposited in public databases. ''Methanococcus jannaschii'' was the first archaeon ...
* Nuclear localization sequence
* '' The Surprising Archaea'' (book)
* Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya
* Unique properties of hyperthermophilic archaea
This article discusses the Unique properties of hyperthermophilic archaea. Hyperthermophiles are organisms that can live at temperatures ranging between 70 and 125 °C.Vieille, C. & Zeikus, G. J. "Hyperthermophilic Enzymes: Sources, Uses, and ...
* Branching order of bacterial phyla (Genome Taxonomy Database, 2018)
There are several models of the branching order of bacterial phyla, one of these is the Genome Taxonomy Database (GTDB).
The GTDB is an initiative to establish a standardised microbial taxonomy based on genome phylogeny, primarily funded by an Aus ...
References
Further reading
* * * * * * * *External links
General
Introduction to the Archaea, ecology, systematics and morphology
Oceans of Archaea
nbsp;– E.F. DeLong, ''ASM News'', 2003
Classification
NCBI taxonomy page on Archaea
nbsp;– list of Prokaryotic names with Standing in Nomenclature
Shotgun sequencing finds nanoorganisms
nbsp;– discovery of the ARMAN group of archaea
Genomics
Browse any completed archaeal genome at UCSC
Comparative Analysis of Archaeal Genomes
(at DOE's IMG system) {{Authority control Extremophiles Domains (biology) Systems of bacterial taxonomy