|
Ancient Pathogen Genomics
Ancient pathogen genomics is a scientific field related to the study of pathogen genomes recovered from ancient human, plant or animal remains. Ancient pathogens are microorganisms, now extinct, that in the past centuries caused several epidemics and deaths worldwide. Their genome, referred to as ancient DNA (aDNA), is isolated from the burial's remains (bones and teeth) of victims of the pandemics caused by these pathogens. The analysis of the genomic features of ancient pathogen genomes allows researchers to understand the evolution of modern microbial strains that can hypothetically generate new pandemics or outbreaks. The analysis of aDNA is carried out by bioinformatic tools and molecular biology techniques to compare ancient pathogens with the modern descendants. The comparison also provides phylogenetic information of these strains. Reconstructing ancient pathogen genomes through NGS technologies Pathogen DNA detection in ancient remains can be achieved with laboratory or ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Scientific Field
The branches of science, also referred to as sciences, scientific fields or scientific disciplines, are commonly divided into three major groups: *Formal sciences: the study of formal systems, such as those under the branches of logic and mathematics, which use an ''a priori'', as opposed to empirical, methodology. *Natural sciences: the study of natural phenomena (including cosmological, geological, physical, chemical, and biological factors of the universe). Natural science can be divided into two main branches: physical science and life science (or biology). *Social sciences: the study of human behavior in its social and cultural aspects. Scientific knowledge must be based on observable phenomena and must be capable of being verified by other researchers working under the same conditions. This verifiability may well vary even ''within'' a scientific discipline. Natural, social, and formal science make up the fundamental sciences, which form the basis of interdisciplinarity- ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Deamination
Deamination is the removal of an amino group from a molecule. Enzymes that catalyse this reaction are called deaminases. In the human body, deamination takes place primarily in the liver, however it can also occur in the kidney. In situations of excess protein intake, deamination is used to break down amino acids for energy. The amino group is removed from the amino acid and converted to ammonia. The rest of the amino acid is made up of mostly carbon and hydrogen, and is recycled or oxidized for energy. Ammonia is toxic to the human system, and enzymes convert it to urea or uric acid by addition of carbon dioxide molecules (which is not considered a deamination process) in the urea cycle, which also takes place in the liver. Urea and uric acid can safely diffuse into the blood and then be excreted in urine. Deamination reactions in DNA Cytosine Spontaneous deamination is the hydrolysis reaction of cytosine into uracil, releasing ammonia in the process. This can occur in vitro thr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Uracil-DNA Glycosylase
Uracil-DNA glycosylase is also known as UNG or UDG. Its most important function is to prevent mutagenesis by eliminating uracil from DNA molecules by cleaving the N-glycosidic bond and initiating the base-excision repair (BER) pathway. Function The human gene encodes one of several uracil-DNA glycosylases. Alternative promoter usage and splicing of this gene leads to two different isoforms: the mitochondrial UNG1 and the nuclear UNG2. One important function of uracil-DNA glycosylases is to prevent mutagenesis by eliminating uracil from DNA molecules by cleaving the N-glycosidic bond and initiating the base-excision repair (BER) pathway. Uracil bases occur from cytosine deamination or misincorporation of dUMP residues. After a mutation occurs, the mutagenic threat of uracil propagates through any subsequent DNA replication steps. Once unzipped, mismatched guanine and uracil pairs are separated, and DNA polymerase inserts complementary bases to form a guanine-cytosine (GC) pair in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
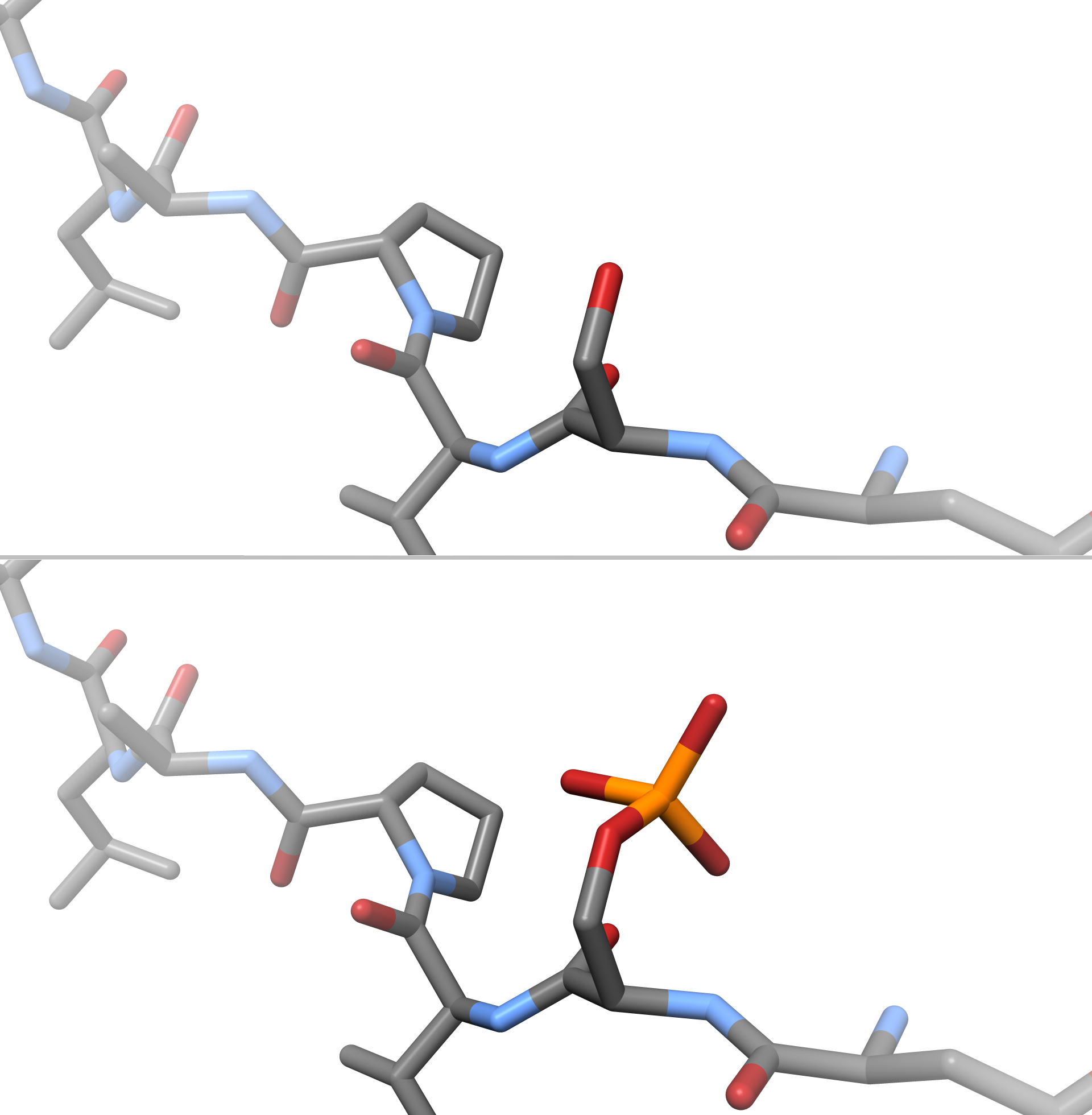
Phosphorylation
In chemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology and could be driven by natural selection. Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. Protein phosphorylation often activates (or deactivates) many enzymes. Glucose Phosphorylation of sugars is often the first stage in their catabolism. Phosphorylation allows cells to accumulate sugars because the phosphate group prevents the molecules from diffusing back across their transporter. Phosphorylation of glucose is a key reaction in sugar metabolism. The chemical equation for the conversion of D-glucose to D-glucose-6-phosphate in the first step of glycolysis is given by :D-glucose + ATP → D-glucose-6-phosphate + ADP : ΔG° = −16.7 kJ/mol (° indicates measurement at standard condition) Hepatic cells are freely permeable to glucose, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Enzyme
A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class of the broader endonuclease group of enzymes. Restriction enzymes are commonly classified into five types, which differ in their structure and whether they cut their DNA substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. To cut DNA, all restriction enzymes make two incisions, once through each sugar-phosphate backbone (i.e. each strand) of the DNA double helix. These enzymes are found in bacteria and archaea and provide a defense mechanism against invading viruses. Inside a prokaryote, the restriction enzymes selectively cut up ''foreign'' DNA in a process called ''restriction digestion''; meanwhile, host DNA is protected by a modification enzyme (a methyltransferase) that modifi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Microarray
A microarray is a multiplex lab-on-a-chip. Its purpose is to simultaneously detect the expression of thousands of genes from a sample (e.g. from a tissue). It is a two-dimensional array on a solid substrate—usually a glass slide or silicon thin-film cell—that assays (tests) large amounts of biological material using high-throughput screening miniaturized, multiplexed and parallel processing and detection methods. The concept and methodology of microarrays was first introduced and illustrated in antibody microarrays (also referred to as antibody matrix) by Tse Wen Chang in 1983 in a scientific publication and a series of patents. The "gene chip" industry started to grow significantly after the 1995 ''Science Magazine'' article by the Ron Davis and Pat Brown labs at Stanford University. With the establishment of companies, such as Affymetrix, Agilent, Applied Microarrays, Arrayjet, Illumina, and others, the technology of DNA microarrays has become the most sophisticated and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Streptavidin
Streptavidin is a 66.0 (tetramer) kDa protein purified from the bacterium '' Streptomyces avidinii''. Streptavidin homo-tetramers have an extraordinarily high affinity for biotin (also known as vitamin B7 or vitamin H). With a dissociation constant (Kd) on the order of ≈10−14 mol/L, the binding of biotin to streptavidin is one of the strongest non-covalent interactions known in nature. Streptavidin is used extensively in molecular biology and bionanotechnology due to the streptavidin-biotin complex's resistance to organic solvents, denaturants (e.g. guanidinium chloride), detergents (e.g. SDS, Triton X-100), proteolytic enzymes, and extremes of temperature and pH. Structure The crystal structure of streptavidin with biotin bound was reported by two groups in 1989. The structure was solved using multi wavelength anomalous diffraction by Hendrickson et al. at Columbia University and using multiple isomorphous replacement by Weber et al. at E. I. DuPont Central Research ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create two identical DNA duplexes from a single original DNA duplex. During this process, DNA polymerase "reads" the existing DNA strands to create two new strands that match the existing ones. These enzymes catalyze the chemical reaction : deoxynucleoside triphosphate + DNAn pyrophosphate + DNAn+1. DNA polymerase adds nucleotides to the three prime (3')-end of a DNA strand, one nucleotide at a time. Every time a cell divides, DNA polymerases are required to duplicate the cell's DNA, so that a copy of the original DNA molecule can be passed to each daughter cell. In this way, genetic information is passed down from generation to generation. Before replication can take place, an enzyme called helicase unwinds the DNA molecule from its tightl ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biotin
Biotin (or vitamin B7) is one of the B vitamins. It is involved in a wide range of metabolic processes, both in humans and in other organisms, primarily related to the utilization of fats, carbohydrates, and amino acids. The name ''biotin'', borrowed from the German , derives from the Ancient Greek word (; 'life') and the suffix "-in" (a suffix used in chemistry usually to indicate 'forming'). Chemical description Biotin is classified as a heterocyclic compound, with a sulfur-containing ring fused ureido and tetrahydrothiophene group. A C5-carboxylic acid side chain is appended to one of the rings. The ureido ring, containing the −N−CO−N− group, serves as the carbon dioxide carrier in carboxylation reactions. Biotin is a coenzyme for five carboxylase enzymes, which are involved in the catabolism of amino acids and fatty acids, synthesis of fatty acids, and gluconeogenesis. Biotinylation of histone proteins in nuclear chromatin plays a role in chromatin stability and g ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nick (DNA)
A nick is a discontinuity in a double stranded DNA molecule where there is no phosphodiester bond between adjacent nucleotides of one strand typically through damage or enzyme action. Nicks allow DNA strands to untwist during replication, and are also thought to play a role in the DNA mismatch repair mechanisms that fix errors on both the leading and lagging daughter strands.Williams JS, Kunkel TA. Ribonucleotides in DNA: Origins, repair and consequences. DNA repair. 2014;19:27-37. doi:10.1016/j.dnarep.2014.03.029 Formation of nicks The diagram shows the effects of nicks on intersecting DNA in a twisted plasmid. Nicking can be used to dissipate the energy held up by intersecting states. The nicks allow the DNA to take on a circular shape.Chao Ji, Lingyun Zhang, and Pengye Wang,Configuration Transitions of Free Circular DNA System Induced by Nicks" Journal of Nanomaterials, vol. 2015, Article ID 546851, 7 pages, 2015. doi:10.1155/2015/546851 Nicked DNA can be the result of D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Library (biology)
In molecular biology, a library is a collection of DNA fragments that is stored and propagated in a population of micro-organisms through the process of molecular cloning. There are different types of DNA libraries, including cDNA libraries (formed from reverse-transcribed RNA), genomic libraries (formed from genomic DNA) and randomized mutant libraries (formed by de novo gene synthesis where alternative nucleotides or codons are incorporated). DNA library technology is a mainstay of current molecular biology, genetic engineering, and protein engineering, and the applications of these libraries depend on the source of the original DNA fragments. There are differences in the cloning vectors and techniques used in library preparation, but in general each DNA fragment is uniquely inserted into a cloning vector and the pool of recombinant DNA molecules is then transferred into a population of bacteria (a Bacterial Artificial Chromosome or BAC library) or yeast such that each or ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |