Pseudomonas aeruginosa on:
[Wikipedia]
[Google]
[Amazon]
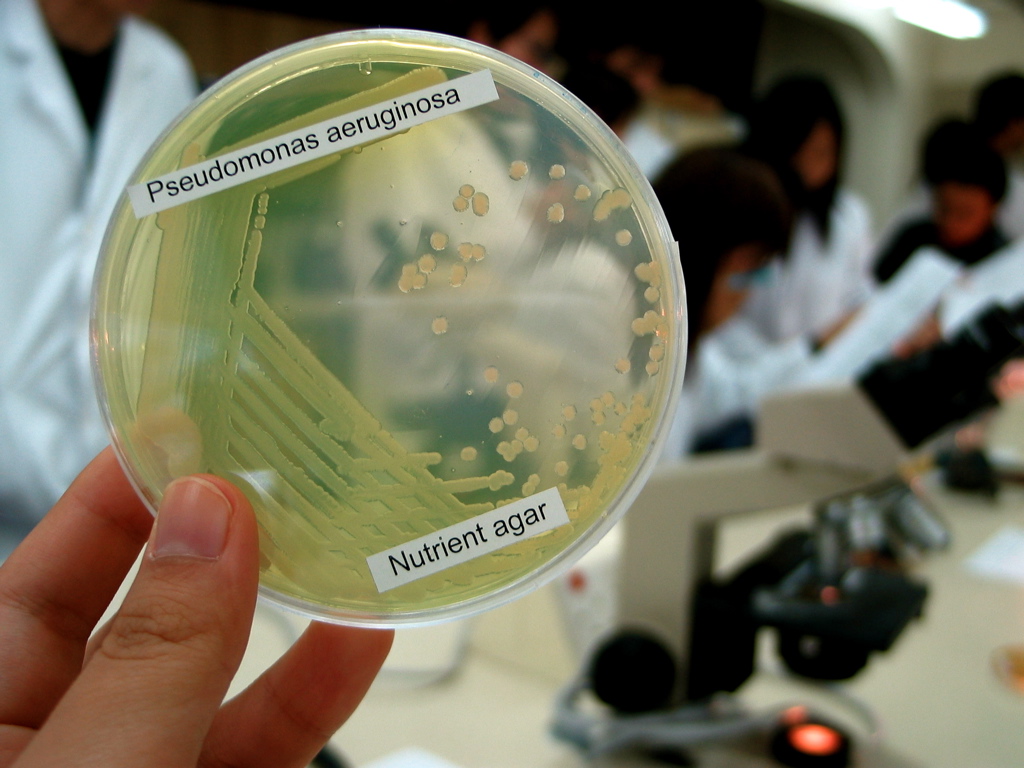 ''Pseudomonas aeruginosa'' is a common encapsulated,
''Pseudomonas aeruginosa'' is a common encapsulated,

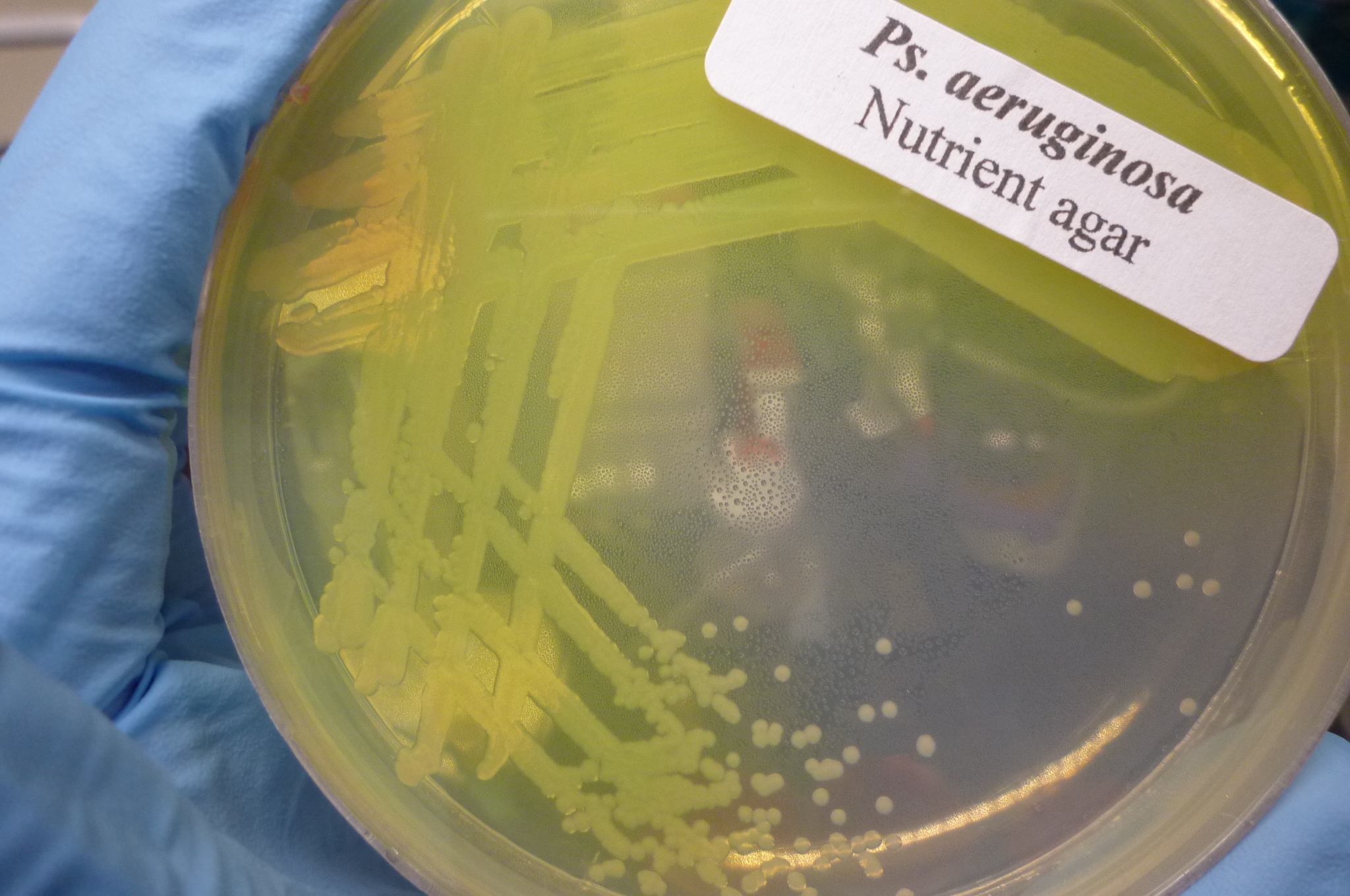 The word ''Pseudomonas'' means "false unit", from the Greek ''pseudēs'' (
The word ''Pseudomonas'' means "false unit", from the Greek ''pseudēs'' (
 An
An
 Depending on the nature of infection, an appropriate specimen is collected and sent to a
Depending on the nature of infection, an appropriate specimen is collected and sent to a
 Clinical identification of ''P. aeruginosa'' may include identifying the production of both pyocyanin and fluorescein, as well as its ability to grow at 42 °C. ''P. aeruginosa'' is capable of growth in
Clinical identification of ''P. aeruginosa'' may include identifying the production of both pyocyanin and fluorescein, as well as its ability to grow at 42 °C. ''P. aeruginosa'' is capable of growth in
 Due to widespread resistance to many common first-line antibiotics,
Due to widespread resistance to many common first-line antibiotics,  cephalosporins (
cephalosporins (
Type strain of ''Pseudomonas aeruginosa'' at Bac''Dive'' - the Bacterial Diversity Metadatabase
* Johanna M. Sweere ''et al.'' (2019)
Bacteriophage trigger antiviral immunity and prevent clearance of bacterial infection
Science 29 Mar 2019: Vol. 363, Issue 6434, eaat9691. doi:10.1126/science.aat9691 about Pseudomonas aeruginosa filamentous phages (Pf-phages), ''
UM researchers publish new discoveries on bacterial viruses
On: EurekAlert! 1 Apr 2019. Source: University of Montana {{Authority control Antibiotic-resistant bacteria Bacteria described in 1872 Gram-negative bacteria Pseudomonadales
 ''Pseudomonas aeruginosa'' is a common encapsulated,
''Pseudomonas aeruginosa'' is a common encapsulated, gram-negative
Gram-negative bacteria are bacteria that do not retain the crystal violet stain used in the Gram staining method of bacterial differentiation. They are characterized by their cell envelopes, which are composed of a thin peptidoglycan cell wall ...
, aerobic
Aerobic means "requiring air," in which "air" usually means oxygen.
Aerobic may also refer to
* Aerobic exercise, prolonged exercise of moderate intensity
* Aerobics, a form of aerobic exercise
* Aerobic respiration, the aerobic process of cel ...
–facultatively anaerobic
A facultative anaerobic organism is an organism that makes ATP by aerobic respiration if oxygen is present, but is capable of switching to fermentation if oxygen is absent.
Some examples of facultatively anaerobic bacteria are ''Staphylococcus' ...
, rod-shaped
A bacillus (), also called a bacilliform bacterium or often just a rod (when the context makes the sense clear), is a rod-shaped bacterium or archaeon. Bacilli are found in many different taxonomic groups of bacteria. However, the name '' Baci ...
bacterium
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
that can cause disease
A disease is a particular abnormal condition that negatively affects the structure or function of all or part of an organism, and that is not immediately due to any external injury. Diseases are often known to be medical conditions that a ...
in plants and animals, including humans. A species
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriate s ...
of considerable medical importance, ''P. aeruginosa'' is a multidrug resistant pathogen
In biology, a pathogen ( el, πάθος, "suffering", "passion" and , "producer of") in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ ...
recognized for its ubiquity, its intrinsically advanced antibiotic resistance
Antimicrobial resistance (AMR) occurs when microbes evolve mechanisms that protect them from the effects of antimicrobials. All classes of microbes can evolve resistance. Fungi evolve antifungal resistance. Viruses evolve antiviral resistance. ...
mechanisms, and its association with serious illnesses – hospital-acquired infections
A hospital-acquired infection, also known as a nosocomial infection (from the Greek , meaning "hospital"), is an infection that is acquired in a hospital or other health care facility. To emphasize both hospital and nonhospital settings, it is ...
such as ventilator-associated pneumonia
Ventilator-associated pneumonia (VAP) is a type of lung infection that occurs in people who are on mechanical ventilation breathing machines in hospitals. As such, VAP typically affects critically ill persons that are in an intensive care unit (I ...
and various sepsis
Sepsis, formerly known as septicemia (septicaemia in British English) or blood poisoning, is a life-threatening condition that arises when the body's response to infection causes injury to its own tissues and organs. This initial stage is follo ...
syndromes.
The organism is considered opportunistic
Opportunism is the practice of taking advantage of circumstances – with little regard for principles or with what the consequences are for others. Opportunist actions are expedient actions guided primarily by self-interested motives. The term ...
insofar as serious infection often occurs during existing disease
A disease is a particular abnormal condition that negatively affects the structure or function of all or part of an organism, and that is not immediately due to any external injury. Diseases are often known to be medical conditions that a ...
s or conditions – most notably cystic fibrosis
Cystic fibrosis (CF) is a rare genetic disorder that affects mostly the lungs, but also the pancreas, liver, kidneys, and intestine. Long-term issues include difficulty breathing and coughing up mucus as a result of frequent lung infections. O ...
and traumatic burns. It generally affects the immunocompromised
Immunodeficiency, also known as immunocompromisation, is a state in which the immune system's ability to fight infectious diseases and cancer is compromised or entirely absent. Most cases are acquired ("secondary") due to extrinsic factors that a ...
but can also infect the immunocompetent
In immunology, immunocompetence is the ability of the body to produce a normal immune response following exposure to an antigen. Immunocompetence is the opposite of immunodeficiency (also known as ''immuno-incompetence'' or being ''immuno-compro ...
as in hot tub folliculitis
Hot tub folliculitis (''pseudomonal'' folliculitis) is a common type of folliculitis, a condition which causes inflammation of hair follicles.
This condition is caused by an infection of hair follicles by a non-pathogenic strain of the bacterium ...
. Treatment of ''P. aeruginosa'' infections can be difficult due to its natural resistance to antibiotics. When more advanced antibiotic drug regimens are needed adverse effects
An adverse effect is an undesired harmful effect resulting from a medication or other intervention, such as surgery. An adverse effect may be termed a "side effect", when judged to be secondary to a main or therapeutic effect. The term complica ...
may result.
It is citrate
Citric acid is an organic compound with the chemical formula HOC(CO2H)(CH2CO2H)2. It is a colorless weak organic acid. It occurs naturally in citrus fruits. In biochemistry, it is an intermediate in the citric acid cycle, which occurs in the ...
, catalase, and oxidase positive
The oxidase test is used to determine if an organism possesses the cytochrome c oxidase enzyme. The test is used as an aid for the differentiation of ''Neisseria'', ''Moraxella'', '' Campylobacter'' and ''Pasteurella'' species (oxidase positive). I ...
. It is found in soil, water, skin flora, and most man-made environments throughout the world. It thrives not only in normal atmospheres, but also in low-oxygen atmospheres, thus has colonized many natural and artificial environments. It uses a wide range of organic material for food; in animals, its versatility enables the organism to infect damaged tissues or those with reduced immunity. The symptoms of such infections are generalized inflammation
Inflammation (from la, wikt:en:inflammatio#Latin, inflammatio) is part of the complex biological response of body tissues to harmful stimuli, such as pathogens, damaged cells, or Irritation, irritants, and is a protective response involving im ...
and sepsis
Sepsis, formerly known as septicemia (septicaemia in British English) or blood poisoning, is a life-threatening condition that arises when the body's response to infection causes injury to its own tissues and organs. This initial stage is follo ...
. If such colonizations occur in critical body organs, such as the lung
The lungs are the primary organs of the respiratory system in humans and most other animals, including some snails and a small number of fish. In mammals and most other vertebrates, two lungs are located near the backbone on either side of t ...
s, the urinary tract
The urinary system, also known as the urinary tract or renal system, consists of the kidneys, ureters, bladder, and the urethra. The purpose of the urinary system is to eliminate waste from the body, regulate blood volume and blood pressure, c ...
, and kidney
The kidneys are two reddish-brown bean-shaped organs found in vertebrates. They are located on the left and right in the retroperitoneal space, and in adult humans are about in length. They receive blood from the paired renal arteries; blood ...
s, the results can be fatal. Because it thrives on moist surfaces, this bacterium is also found on and in medical equipment, including catheter
In medicine, a catheter (/ˈkæθətər/) is a thin tubing (material), tube made from medical grade materials serving a broad range of functions. Catheters are medical devices that can be inserted in the body to treat diseases or perform a surgi ...
s, causing cross-infection
An infection is the invasion of tissues by pathogens, their multiplication, and the reaction of host tissues to the infectious agent and the toxins they produce. An infectious disease, also known as a transmissible disease or communicable dise ...
s in hospital
A hospital is a health care institution providing patient treatment with specialized health science and auxiliary healthcare staff and medical equipment. The best-known type of hospital is the general hospital, which typically has an emerge ...
s and clinic
A clinic (or outpatient clinic or ambulatory care clinic) is a health facility that is primarily focused on the care of outpatients. Clinics can be privately operated or publicly managed and funded. They typically cover the primary care needs ...
s. It is also able to decompose hydrocarbons and has been used to break down tarballs and oil from oil spills. ''P. aeruginosa'' is not extremely virulent
Virulence is a pathogen's or microorganism's ability to cause damage to a host.
In most, especially in animal systems, virulence refers to the degree of damage caused by a microbe to its host. The pathogenicity of an organism—its ability to ...
in comparison with other major pathogenic bacterial species – for example the Gram-positive
In bacteriology, gram-positive bacteria are bacteria that give a positive result in the Gram stain test, which is traditionally used to quickly classify bacteria into two broad categories according to their type of cell wall.
Gram-positive bact ...
''Staphylococcus aureus
''Staphylococcus aureus'' is a Gram-positive spherically shaped bacterium, a member of the Bacillota, and is a usual member of the microbiota of the body, frequently found in the upper respiratory tract and on the skin. It is often positive ...
'' and ''Streptococcus pyogenes
''Streptococcus pyogenes'' is a species of Gram-positive, aerotolerant bacteria in the genus ''Streptococcus''. These bacteria are extracellular, and made up of non-motile and non-sporing cocci (round cells) that tend to link in chains. They are ...
'' – though ''P. aeruginosa'' is capable of extensive colonization, and can aggregate into enduring biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
s.
Nomenclature

Greek
Greek may refer to:
Greece
Anything of, from, or related to Greece, a country in Southern Europe:
*Greeks, an ethnic group.
*Greek language, a branch of the Indo-European language family.
**Proto-Greek language, the assumed last common ancestor ...
: ψευδής, false) and ( la, monas, from Greek
Greek may refer to:
Greece
Anything of, from, or related to Greece, a country in Southern Europe:
*Greeks, an ethnic group.
*Greek language, a branch of the Indo-European language family.
**Proto-Greek language, the assumed last common ancestor ...
: μονάς, a single unit). The stem word ''mon'' was used early in the history of microbiology to refer to germs, e.g., kingdom Monera
Monera (/məˈnɪərə/) (Greek - μονήρης (monḗrēs), "single", "solitary") is a biological kingdom that is made up of prokaryotes. As such, it is composed of single-celled organisms that lack a nucleus.
The taxon Monera was first p ...
.
The species name ''aeruginosa'' is a Latin word meaning verdigris
Verdigris is the common name for blue-green, copper-based pigments that form a patina on copper, bronze, and brass. The technical literature is ambiguous as to its chemical composition. Some sources refer to "neutral verdigris" as copper(II) ...
("copper rust"), referring to the blue-green color of laboratory cultures of the species. This blue-green pigment is a combination of two metabolites of ''P. aeruginosa'', pyocyanin
Pyocyanin (PCN−) is one of the many toxic compounds produced and secreted by the Gram negative bacterium ''Pseudomonas aeruginosa''. Pyocyanin is a blue secondary metabolite, turning red below pH 4.9, with the ability to oxidise and reduce other ...
(blue) and pyoverdine
Pyoverdines (alternatively, and less commonly, spelled as pyoverdins) are fluorescent siderophores produced by certain pseudomonads. Pyoverdines are important virulence factors, and are required for pathogenesis in many biological models of inf ...
(green), which impart the blue-green characteristic color of cultures. Another assertion from 1956 is that ''aeruginosa'' may be derived from the Greek prefix ''ae-'' meaning "old or aged", and the suffix ''ruginosa'' means wrinkled or bumpy.
The names pyocyanin and pyoverdine are from the Greek, with ''pyo-'', meaning "pus", ''cyanin'', meaning "blue", and ''verdine'', meaning "green". Hence, the term "pyocyanic bacteria" refers specifically to the "blue pus" characteristic of a ''P. aeruginosa'' infection. Pyoverdine in the absence of pyocyanin is a fluorescent-yellow color.

Biology
Genome
Thegenome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
of ''Pseudomonas aeruginosa'' consists of a relatively large circular chromosome (5.5–6.8 Mb) that carries between 5,500 and 6,000 open reading frame
In molecular biology, open reading frames (ORFs) are defined as spans of DNA sequence between the start and stop codons. Usually, this is considered within a studied region of a prokaryotic DNA sequence, where only one of the six possible readin ...
s, and sometimes plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; how ...
s of various sizes depending on the strain. Comparison of 389 genomes from different ''P. aeruginosa'' strains showed that just 17.5% is shared. This part of the genome is the ''P. aeruginosa'' core genome.
A comparative genomic study (in 2020) analyzed 494 complete genomes from the ''Pseudomonas'' genus, of which 189 were ''P. aeruginosa'' strains. The study observed that their protein count and GC content
In molecular biology and genetics, GC-content (or guanine-cytosine content) is the percentage of nitrogenous bases in a DNA or RNA molecule that are either guanine (G) or cytosine (C). This measure indicates the proportion of G and C bases out ...
ranged between 5500 and 7352 (average: 6192) and between 65.6 and 66.9% (average: 66.1%), respectively. This comparative analysis further identified 1811 aeruginosa-core proteins, which accounts for more than 30% of the proteome. The higher percentage of aeruginosa-core proteins in this latter analysis could partly be attributed to the use of complete genomes. Although ''P. aeruginosa'' is a very well-defined monophyletic species, phylogenomically and in terms of ANIm values, it is surprisingly diverse in terms of protein content, thus revealing a very dynamic accessory proteome, in accordance with several analyses. It appears that, on average, industrial strains have the largest genomes, followed by environmental strains, and then clinical isolates. The same comparative study (494 ''Pseudomonas'' strains, of which 189 are ''P. aeruginosa'') identified that 41 of the 1811 ''P. aeruginosa'' core proteins were present only in this species and not in any other member of the genus, with 26 (of the 41) being annotated as hypothetical. Furthermore, another 19 orthologous protein groups are present in at least 188/189 ''P. aeruginosa'' strains and absent in all the other strains of the genus.
Population structure
The population of ''P. aeruginosa'' can be classified in three main lineages, genetically characterised by the model strains PAO1, PA14, and the more divergent PA7. While ''P. aeruginosa'' is generally thought of as an opportunistic pathogen, several widespread clones appear to have become more specialised pathogens, particularly in cystic fibrosis patients, including the Liverpool epidemic strain (LES) which is found mainly in the UK, DK2 in Denmark, and AUST-02 in Australia (also previously known as AES-2 and P2). There is also a clone that is frequently found infecting the reproductive tracts of horses.Metabolism
''P. aeruginosa'' is afacultative anaerobe
A facultative anaerobic organism is an organism that makes ATP by aerobic respiration if oxygen is present, but is capable of switching to fermentation if oxygen is absent.
Some examples of facultatively anaerobic bacteria are '' Staphylococc ...
, as it is well adapted to proliferate in conditions of partial or total oxygen depletion. This organism can achieve anaerobic
Anaerobic means "living, active, occurring, or existing in the absence of free oxygen", as opposed to aerobic which means "living, active, or occurring only in the presence of oxygen." Anaerobic may also refer to:
* Anaerobic adhesive, a bonding a ...
growth with nitrate
Nitrate is a polyatomic ion
A polyatomic ion, also known as a molecular ion, is a covalent bonded set of two or more atoms, or of a metal complex, that can be considered to behave as a single unit and that has a net charge that is not zer ...
or nitrite as a terminal electron acceptor
An electron acceptor is a chemical entity that accepts electrons transferred to it from another compound. It is an oxidizing agent that, by virtue of its accepting electrons, is itself reduced in the process. Electron acceptors are sometimes mista ...
. When oxygen, nitrate, and nitrite are absent, it is able to ferment arginine and pyruvate by substrate-level phosphorylation. Adaptation to microaerobic or anaerobic environments is essential for certain lifestyles of ''P. aeruginosa'', for example, during lung infection in cystic fibrosis
Cystic fibrosis (CF) is a rare genetic disorder that affects mostly the lungs, but also the pancreas, liver, kidneys, and intestine. Long-term issues include difficulty breathing and coughing up mucus as a result of frequent lung infections. O ...
and primary ciliary dyskinesia
Primary ciliary dyskinesia (PCD) is a rare, autosomal recessive genetic ciliopathy, that causes defects in the action of cilia lining the upper and lower respiratory tract, sinuses, Eustachian tube, middle ear, Fallopian tube, and flagella of ...
, where thick layers of lung mucus
Mucus ( ) is a slippery aqueous secretion produced by, and covering, mucous membranes. It is typically produced from cells found in mucous glands, although it may also originate from mixed glands, which contain both serous and mucous cells. It ...
and bacterially-produced alginate
Alginic acid, also called algin, is a naturally occurring, edible polysaccharide found in brown algae. It is hydrophilic and forms a viscous gum when hydrated. With metals such as sodium and calcium, its salts are known as alginates. Its colour ...
surrounding mucoid bacterial cells can limit the diffusion of oxygen. ''P. aeruginosa'' growth within the human body can be asymptomatic until the bacteria form a biofilm, which overwhelms the immune system. These biofilms are found in the lungs of people with cystic fibrosis and primary ciliary dyskinesia, and can prove fatal.
Cellular cooperation
''P. aeruginosa'' relies oniron
Iron () is a chemical element with symbol Fe (from la, ferrum) and atomic number 26. It is a metal that belongs to the first transition series and group 8 of the periodic table. It is, by mass, the most common element on Earth, right in f ...
as a nutrient source to grow. However, iron is not easily accessible because it is not commonly found in the environment. Iron is usually found in a largely insoluble ferric form. Furthermore, excessively high levels of iron can be toxic to ''P. aeruginosa''. To overcome this and regulate proper intake of iron, ''P. aeruginosa'' uses siderophore
Siderophores (Greek: "iron carrier") are small, high-affinity iron- chelating compounds that are secreted by microorganisms such as bacteria and fungi. They help the organism accumulate iron. Although a widening range of siderophore functions is n ...
s, which are secreted molecules that bind and transport iron. These iron-siderophore complexes, however, are not specific. The bacterium that produced the siderophores does not necessarily receive the direct benefit of iron intake. Rather, all members of the cellular population are equally likely to access the iron-siderophore complexes. Members of the cellular population that can efficiently produce these siderophores are commonly referred to as cooperators; members that produce little to no siderophores are often referred to as cheaters. Research has shown when cooperators and cheaters are grown together, cooperators have a decrease in fitness, while cheaters have an increase in fitness. The magnitude of change in fitness increases with increasing iron limitation. With an increase in fitness, the cheaters can outcompete the cooperators; this leads to an overall decrease in fitness of the group, due to lack of sufficient siderophore production. These observations suggest that having a mix of cooperators and cheaters can reduce the virulent nature of ''P. aeruginosa''.
Enzymes
LigDs form a subfamily of theDNA ligase
DNA ligase is a specific type of enzyme, a ligase, () that facilitates the joining of DNA strands together by catalyzing the formation of a phosphodiester bond. It plays a role in repairing single-strand breaks in duplex DNA in living organ ...
s. These all have a LigDom/ligase domain, but many bacterial LigDs also have separate polymerase domains/PolDoms and nuclease domains/NucDoms. In ''P. aeruginosa''s case the nuclease domains are N-terminus
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the ami ...
, and the polymerase domains are C-terminus
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is ...
, extensions of the single central ligase domain.
Pathogenesis
opportunistic
Opportunism is the practice of taking advantage of circumstances – with little regard for principles or with what the consequences are for others. Opportunist actions are expedient actions guided primarily by self-interested motives. The term ...
, nosocomial
A hospital-acquired infection, also known as a nosocomial infection (from the Greek , meaning "hospital"), is an infection that is acquired in a hospital or other health care facility. To emphasize both hospital and nonhospital settings, it is ...
pathogen of immunocompromised
Immunodeficiency, also known as immunocompromisation, is a state in which the immune system's ability to fight infectious diseases and cancer is compromised or entirely absent. Most cases are acquired ("secondary") due to extrinsic factors that a ...
individuals, ''P. aeruginosa'' typically infects the airway, urinary tract
The urinary system, also known as the urinary tract or renal system, consists of the kidneys, ureters, bladder, and the urethra. The purpose of the urinary system is to eliminate waste from the body, regulate blood volume and blood pressure, c ...
, burns Burns may refer to:
* Burn, an injury (plural)
People:
* Burns (surname), includes list of people and characters
Business:
* Burns London, a British guitar maker
Places:
;In the United States
* Burns, Colorado, unincorporated community in Eagle ...
, and wounds, and also causes other blood infections.
It is the most common cause of infections of burn injuries and of the outer ear
The outer ear, external ear, or auris externa is the external part of the ear, which consists of the auricle (also pinna) and the ear canal. It gathers sound energy and focuses it on the eardrum (tympanic membrane).
Structure
Auricle
Th ...
(otitis externa
Otitis externa, also called swimmer's ear, is inflammation of the ear canal. It often presents with ear pain, swelling of the ear canal, and occasionally decreased hearing. Typically there is pain with movement of the outer ear. A high fever is ...
), and is the most frequent colonizer of medical devices (e.g., catheter
In medicine, a catheter (/ˈkæθətər/) is a thin tubing (material), tube made from medical grade materials serving a broad range of functions. Catheters are medical devices that can be inserted in the body to treat diseases or perform a surgi ...
s). ''Pseudomonas'' can be spread by equipment that gets contaminated and is not properly cleaned or on the hands of healthcare workers. ''Pseudomonas'' can, in rare circumstances, cause community-acquired pneumonia
Community-acquired pneumonia (CAP) refers to pneumonia (any of several lung diseases) contracted by a person outside of the healthcare system. In contrast, hospital-acquired pneumonia (HAP) is seen in patients who have recently visited a hospital ...
s, as well as ventilator
A ventilator is a piece of medical technology that provides mechanical ventilation by moving breathable air into and out of the lungs, to deliver breaths to a patient who is physically unable to breathe, or breathing insufficiently. Ventilators ...
-associated pneumonias, being one of the most common agents isolated in several studies. Pyocyanin
Pyocyanin (PCN−) is one of the many toxic compounds produced and secreted by the Gram negative bacterium ''Pseudomonas aeruginosa''. Pyocyanin is a blue secondary metabolite, turning red below pH 4.9, with the ability to oxidise and reduce other ...
is a virulence factor
Virulence factors (preferably known as pathogenicity factors or effectors in plant science) are cellular structures, molecules and regulatory systems that enable microbial pathogens (bacteria, viruses, fungi, and protozoa) to achieve the following ...
of the bacteria and has been known to cause death in ''C. elegans
''Caenorhabditis elegans'' () is a free-living transparent nematode about 1 mm in length that lives in temperate soil environments. It is the type species of its genus. The name is a blend of the Greek ''caeno-'' (recent), ''rhabditis'' (r ...
'' by oxidative stress
Oxidative stress reflects an imbalance between the systemic manifestation of reactive oxygen species and a biological system's ability to readily Detoxification, detoxify the reactive intermediates or to repair the resulting damage. Disturbances ...
. However, salicylic acid can inhibit pyocyanin production. One in ten hospital-acquired infections is from ''Pseudomonas''. Cystic fibrosis
Cystic fibrosis (CF) is a rare genetic disorder that affects mostly the lungs, but also the pancreas, liver, kidneys, and intestine. Long-term issues include difficulty breathing and coughing up mucus as a result of frequent lung infections. O ...
patients are also predisposed to ''P. aeruginosa'' infection of the lungs due to a functional loss in chloride ion movement across cell membranes as a result of a mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mi ...
. ''P. aeruginosa'' may also be a common cause of "hot-tub rash" (dermatitis
Dermatitis is inflammation of the skin, typically characterized by itchiness, redness and a rash. In cases of short duration, there may be small blisters, while in long-term cases the skin may become thickened. The area of skin involved can ...
), caused by lack of proper, periodic attention to water quality. Since these bacteria thrive in moist environments, such as hot tubs and swimming pools, they can cause skin rash or swimmer's ear. ''Pseudomonas'' is also a common cause of postoperative infection in radial keratotomy surgery patients. The organism is also associated with the skin lesion ecthyma gangrenosum
Ecthyma gangrenosum is a type of skin lesion characterized by vesicles or blisters which rapidly evolve into pustules and necrotic ulcers with undermined tender erythematous border. "Ecthyma" means a pus forming infection of the skin with an ulcer ...
. ''P. aeruginosa'' is frequently associated with osteomyelitis
Osteomyelitis (OM) is an infection of bone. Symptoms may include pain in a specific bone with overlying redness, fever, and weakness. The long bones of the arms and legs are most commonly involved in children e.g. the femur and humerus, while the ...
involving puncture wounds of the foot, believed to result from direct inoculation with ''P. aeruginosa'' via the foam padding found in tennis shoes, with diabetic patients at a higher risk.
A comparative genomic analysis of 494 compete ''Pseudomonas'' genomes, including 189 complete ''P. aeruginosa'' genomes, identified several proteins that are shared by the vast majority of ''P. aeruginosa'' strains, but are not observed in other analyzed ''Pseudomonas'' genomes. These aeruginosa-specific core proteins, such as ''CntL, CntM, PlcB, Acp1, MucE, SrfA, Tse1, Tsi2, Tse3,'' and ''EsrC'' are known to play an important role in this species' pathogenicity.
Toxins
''P. aeruginosa'' uses thevirulence factor
Virulence factors (preferably known as pathogenicity factors or effectors in plant science) are cellular structures, molecules and regulatory systems that enable microbial pathogens (bacteria, viruses, fungi, and protozoa) to achieve the following ...
exotoxin A
The Pseudomonas exotoxin (or exotoxin A) is an exotoxin produced by ''Pseudomonas aeruginosa''. ''Vibrio cholerae'' produces a similar protein called the Cholix toxin ().
It inhibits elongation factor-2. It does so by ADP-ribosylation of EF2 u ...
to inactivate eukaryotic elongation factor 2 via ADP-ribosylation in the host cell, much as the diphtheria toxin
Diphtheria toxin is an exotoxin secreted by '' Corynebacterium diphtheriae'', the pathogenic bacterium that causes diphtheria. The toxin gene is encoded by a prophageA prophage is a virus that has inserted itself into the genome of the host ...
does. Without elongation factor 2, eukaryotic cells
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bact ...
cannot synthesize proteins and necrotise. The release of intracellular contents induces an immunologic response in immunocompetent
In immunology, immunocompetence is the ability of the body to produce a normal immune response following exposure to an antigen. Immunocompetence is the opposite of immunodeficiency (also known as ''immuno-incompetence'' or being ''immuno-compro ...
patients.
In addition ''P. aeruginosa'' uses an exoenzyme, ExoU, which degrades the plasma membrane of eukaryotic cells, leading to lysis
Lysis ( ) is the breaking down of the membrane of a cell, often by viral, enzymic, or osmotic (that is, "lytic" ) mechanisms that compromise its integrity. A fluid containing the contents of lysed cells is called a ''lysate''. In molecular bio ...
. Increasingly, it is becoming recognized that the iron-acquiring siderophore
Siderophores (Greek: "iron carrier") are small, high-affinity iron- chelating compounds that are secreted by microorganisms such as bacteria and fungi. They help the organism accumulate iron. Although a widening range of siderophore functions is n ...
, pyoverdine
Pyoverdines (alternatively, and less commonly, spelled as pyoverdins) are fluorescent siderophores produced by certain pseudomonads. Pyoverdines are important virulence factors, and are required for pathogenesis in many biological models of inf ...
, also functions as a toxin by removing iron
Iron () is a chemical element with symbol Fe (from la, ferrum) and atomic number 26. It is a metal that belongs to the first transition series and group 8 of the periodic table. It is, by mass, the most common element on Earth, right in f ...
from mitochondria
A mitochondrion (; ) is an organelle found in the Cell (biology), cells of most Eukaryotes, such as animals, plants and Fungus, fungi. Mitochondria have a double lipid bilayer, membrane structure and use aerobic respiration to generate adenosi ...
, inflicting damage on this organelle.
Phenazines
Phenazines
Phenazine is an organic compound with the formula (C6H4)2N2. It is a dibenzo annulated pyrazine, and the parent substance of many dyestuffs, such as the toluylene red, indulines, and safranines (and the closely related eurhodines). Phenazine cr ...
are redox-active pigments produced by ''P. aeruginosa''. These pigments are involved in quorum sensing
In biology, quorum sensing or quorum signalling (QS) is the ability to detect and respond to cell population density by gene regulation. As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at ...
, virulence
Virulence is a pathogen's or microorganism's ability to cause damage to a host.
In most, especially in animal systems, virulence refers to the degree of damage caused by a microbe to its host. The pathogenicity of an organism—its ability to ...
, and iron acquisition. ''P. aeruginosa'' produces several pigments all produced by a biosynthetic pathway: phenazine-1-carboxamide (PCA), 1-hydroxyphenazine, 5-methylphenazine-1-carboxylic acid betaine, pyocyanin
Pyocyanin (PCN−) is one of the many toxic compounds produced and secreted by the Gram negative bacterium ''Pseudomonas aeruginosa''. Pyocyanin is a blue secondary metabolite, turning red below pH 4.9, with the ability to oxidise and reduce other ...
and aeruginosin A. Two operons are involved in phenazine biosynthesis: ''phzA1B1C1D1E1F1G1'' and ''phzA2B2C2D2E2F2G2''. The enzymes encoded by these operons convert chorismic acid
Chorismic acid, more commonly known as its anionic form chorismate, is an important biochemical intermediate in plants and microorganisms. It is a precursor for:
* The aromatic amino acids phenylalanine, tryptophan, and tyrosine
* Indole, indole d ...
to PCA. The products of three key genes, ''phzH'', ''phzM'', and ''phzS'' then convert PCA to the other phenazines mentioned above. Though phenazine biosynthesis is well studied, questions remain as to the final structure of the brown phenazine
Phenazine is an organic compound with the formula (C6H4)2N2. It is a dibenzo annulated pyrazine, and the parent substance of many dyestuffs, such as the toluylene red, indulines, and safranines (and the closely related eurhodines). Phenazine ...
pyomelanin.
When pyocyanin biosynthesis is inhibited, a decrease in ''P. aeruginosa'' pathogenicity is observed ''in vitro
''In vitro'' (meaning in glass, or ''in the glass'') studies are performed with microorganisms, cells, or biological molecules outside their normal biological context. Colloquially called "test-tube experiments", these studies in biology an ...
''. This suggests that pyocyanin is mostly responsible for the initial colonization of ''P. aeruginosa'' ''in vivo''.
Triggers
With lowphosphate
In chemistry, a phosphate is an anion, salt, functional group or ester derived from a phosphoric acid. It most commonly means orthophosphate, a derivative of orthophosphoric acid .
The phosphate or orthophosphate ion is derived from phospho ...
levels, ''P. aeruginosa'' has been found to activate from benign symbiont to express lethal toxins inside the intestinal tract and severely damage or kill the host, which can be mitigated by providing excess phosphate instead of antibiotics.
Plants and invertebrates
In higher plants, ''P. aeruginosa'' induces soft rot, for example in ''Arabidopsis thaliana
''Arabidopsis thaliana'', the thale cress, mouse-ear cress or arabidopsis, is a small flowering plant native to Eurasia and Africa. ''A. thaliana'' is considered a weed; it is found along the shoulders of roads and in disturbed land.
A winter a ...
'' (Thale cress) and ''Lactuca sativa
Lettuce (''Lactuca sativa'') is an annual plant of the family Asteraceae. It is most often grown as a leaf vegetable, but sometimes for its stem and seeds. Lettuce is most often used for salads, although it is also seen in other kinds of food, ...
'' (lettuce). It is also pathogenic to invertebrate animals, including the nematode '' Caenorhabditis elegans'', the fruit fly ''Drosophila
''Drosophila'' () is a genus of flies, belonging to the family Drosophilidae, whose members are often called "small fruit flies" or (less frequently) pomace flies, vinegar flies, or wine flies, a reference to the characteristic of many species ...
'', and the moth ''Galleria mellonella
''Galleria mellonella'', the greater wax moth or honeycomb moth, is a moth of the family Pyralidae. ''G. mellonella'' is found throughout the world. It is one of two species of wax moths, with the other being the lesser wax moth. ''G. mellonella' ...
.'' The associations of virulence factors are the same for plant and animal infections. In both insects and plants, ''P. aeruginosa'' virulence
Virulence is a pathogen's or microorganism's ability to cause damage to a host.
In most, especially in animal systems, virulence refers to the degree of damage caused by a microbe to its host. The pathogenicity of an organism—its ability to ...
is highly quorum sensing
In biology, quorum sensing or quorum signalling (QS) is the ability to detect and respond to cell population density by gene regulation. As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at ...
(QS) dependent. Its QS is in turn highly dependent upon such genes as acyl-homoserine-lactone synthase, and lasI.
Quorum sensing
''P. aeruginosa'' is an opportunistic pathogen with the ability to coordinate gene expression in order to compete against other species for nutrients or colonization. Regulation ofgene expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. The ...
can occur through cell-cell communication or quorum sensing
In biology, quorum sensing or quorum signalling (QS) is the ability to detect and respond to cell population density by gene regulation. As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at ...
(QS) via the production of small molecules called autoinducers that are released into the external environment. These signals, when reaching specific concentrations correlated with specific population cell densities, activate their respective regulators thus altering gene expression and coordinating behavior. ''P. aeruginosa'' employs five interconnected QS systems – las, rhl, pqs, iqs, and pch – that each produce unique signaling molecules. The las and rhl systems are responsible for the activation of numerous QS-controlled genes, the pqs system is involved in quinolone signaling, and the iqs system plays an important role in intercellular communication. QS in ''P. aeruginosa'' is organized in a hierarchical manner. At the top of the signaling hierarchy is the las system, since the las regulator initiates the QS regulatory system by activating the transcription of a number of other regulators, such as rhl. So, the las system defines a hierarchical QS cascade from the las to the rhl regulons. Detection of these molecules indicates ''P. aeruginosa'' is growing as biofilm within the lungs of cystic fibrosis patients. The impact of QS and especially las systems on the pathogenicity of ''P. aeruginosa'' is unclear, however. Studies have shown that lasR-deficient mutants are associated with more severe outcomes in cystic fibrosis patients and are found in up to 63% of chronically infected cystic fibrosis patients despite impaired QS activity.
QS is known to control expression of a number of virulence factors
Virulence factors (preferably known as pathogenicity factors or effectors in plant science) are cellular structures, molecules and regulatory systems that enable microbial pathogens (bacteria, viruses, fungi, and protozoa) to achieve the followin ...
in a hierarchical manner, including the pigment pyocyanin. However, although the las system initiates the regulation of gene expression, its absence does not lead to loss of virulence factors. Recently, it has been demonstrated that the rhl system partially controls las-specific factors, such as proteolytic enzymes responsible for elastolytic and staphylolytic activities, but in a delayed manner. So, las is a direct and indirect regulator of QS-controlled genes. Another form of gene regulation
Regulation of gene expression, or gene regulation, includes a wide range of mechanisms that are used by cells to increase or decrease the production of specific gene products (protein or RNA). Sophisticated programs of gene expression are wi ...
that allows the bacteria to rapidly adapt to surrounding changes is through environmental signaling. Recent studies have discovered anaerobiosis can significantly impact the major regulatory circuit of QS. This important link between QS and anaerobiosis has a significant impact on production of virulence factors
Virulence factors (preferably known as pathogenicity factors or effectors in plant science) are cellular structures, molecules and regulatory systems that enable microbial pathogens (bacteria, viruses, fungi, and protozoa) to achieve the followin ...
of this organism. Garlic
Garlic (''Allium sativum'') is a species of bulbous flowering plant in the genus ''Allium''. Its close relatives include the onion, shallot, leek, chive, Allium fistulosum, Welsh onion and Allium chinense, Chinese onion. It is native to South A ...
experimentally blocks quorum sensing in ''P. aeruginosa''.
Biofilms formation and cyclic di-GMP
As in most Gram negative bacteria, ''P. aeruginosa''biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
formation is regulated by one single molecule: cyclic di-GMP
Cyclic di-GMP (also called cyclic diguanylate and c-di-GMP) is a second messenger used in signal transduction in a wide variety of bacteria. Cyclic di-GMP is not known to be used by archaea, and has only been observed in eukaryotes in ''Dictyos ...
. At low cyclic di-GMP concentration, ''P. aeruginosa'' has a free-swimming mode of life. But when cyclic di-GMP levels increase, ''P. aeruginosa'' start to establish sessile communities on surfaces. The intracellular concentration of cyclic di-GMP increases within seconds when ''P. aeruginosa'' touches a surface (''e.g.'': a rock, plastic, host tissues...). This activates the production of adhesive pili, that serve as "anchors" to stabilize the attachment of ''P. aeruginosa'' on the surface. At later stages, bacteria will start attaching irreversibly by producing a strongly adhesive matrix. At the same time, cyclic di-GMP represses the synthesis of the flagellar machinery, preventing ''P. aeruginosa'' from swimming. When suppressed, the biofilms are less adherent and easier to treat.
The biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
matrix of ''P. aeruginosa'' is composed of nucleic acids, amino acids, carbohydrates, and various ions. It mechanically and chemically protects ''P. aeruginosa'' from aggression by the immune system and some toxic compounds. ''P. aeruginosa'' biofilm's matrix is composed of 2 types of sugars (or "exopolysacharides") named PSL and PEL:
* Polysaccharide synthesis locus (PSL) and cyclic di-GMP form a positive feedback loop. PSL stimulates cyclic di-GMP production, while high cyclic di-GMP turns on the operon and increases activity of the operon. This 15-gene operon is responsible for the cell-cell and cell-surface interactions required for cell communication. It is also responsible for the sequestering of the extracellular polymeric substance matrix.
* PEL is a cationic exopolysaccharide that cross-links extracellular DNA in the ''P. aeruginosa'' biofilm matrix.
Upon certain cues or stresses, ''P. aeruginosa'' revert the biofilm program and detach. Recent studies have shown that the dispersed cells from ''P. aeruginosa'' biofilms have lower cyclic di-GMP levels and different physiologies from those of planktonic and biofilm cells. Such dispersed cells are found to be highly virulent against macrophages and ''C. elegans'', but highly sensitive towards iron stress, as compared with planktonic cells.
Biofilms and treatment resistance
Biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
s of ''P. aeruginosa'' can cause chronic opportunistic infection
An opportunistic infection is an infection caused by pathogens (bacteria, fungi, parasites or viruses) that take advantage of an opportunity not normally available. These opportunities can stem from a variety of sources, such as a weakened immune ...
s, which are a serious problem for medical care in industrialized societies, especially for immunocompromised patients and the elderly. They often cannot be treated effectively with traditional antibiotic
An antibiotic is a type of antimicrobial substance active against bacteria. It is the most important type of antibacterial agent for fighting bacterial infections, and antibiotic medications are widely used in the treatment and prevention of ...
therapy. Biofilms seem to protect these bacteria from adverse environmental factors. ''P. aeruginosa'' can cause nosocomial infections and is considered a model organism for the study of antibiotic-resistant bacteria. Researchers consider it important to learn more about the molecular mechanisms that cause the switch from planktonic growth to a biofilm phenotype and about the role of QS in treatment-resistant bacteria such as ''P. aeruginosa''. This should contribute to better clinical management of chronically infected patients, and should lead to the development of new drugs.
Scientists have been examining the possible genetic basis for ''P. aeruginosa'' resistance to antibiotics such as tobramycin
Tobramycin is an aminoglycoside antibiotic derived from '' Streptomyces tenebrarius'' that is used to treat various types of bacterial infections, particularly Gram-negative infections. It is especially effective against species of ''Pseudomonas ...
. One locus
Locus (plural loci) is Latin for "place". It may refer to:
Entertainment
* Locus (comics), a Marvel Comics mutant villainess, a member of the Mutant Liberation Front
* ''Locus'' (magazine), science fiction and fantasy magazine
** ''Locus Award' ...
identified as being an important genetic determinant of the resistance in this species is ''ndvB'', which encodes periplasm
The periplasm is a concentrated gel-like matrix in the space between the inner cytoplasmic membrane and the bacterial outer membrane called the ''periplasmic space'' in gram-negative bacteria. Using cryo-electron microscopy it has been found that ...
ic glucans that may interact with antibiotics and cause them to become sequestered into the periplasm. These results suggest a genetic basis exists behind bacterial antibiotic resistance, rather than the biofilm simply acting as a diffusion barrier to the antibiotic.
Diagnosis
 Depending on the nature of infection, an appropriate specimen is collected and sent to a
Depending on the nature of infection, an appropriate specimen is collected and sent to a bacteriology Bacteriology is the branch and specialty of biology that studies the morphology, ecology, genetics and biochemistry of bacteria as well as many other aspects related to them. This subdivision of microbiology involves the identification, classificat ...
laboratory for identification. As with most bacteriological specimens, a Gram stain
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish b ...
is performed, which may show Gram-negative rods and/or white blood cells
White blood cells, also called leukocytes or leucocytes, are the cells of the immune system that are involved in protecting the body against both infectious disease and foreign invaders. All white blood cells are produced and derived from mult ...
. ''P. aeruginosa'' produces colonies with a characteristic "grape-like" or "fresh-tortilla" odor on bacteriological media. In mixed cultures, it can be isolated as clear colonies on MacConkey agar
MacConkey agar is a selective and differential culture medium for bacteria. It is designed to selectively isolate Gram-negative and enteric (normally found in the intestinal tract) bacteria and differentiate them based on lactose fermentation. ...
(as it does not ferment lactose) which will test positive for oxidase
In biochemistry, an oxidase is an enzyme that catalyzes oxidation-reduction reactions, especially one involving dioxygen (O2) as the electron acceptor. In reactions involving donation of a hydrogen atom, oxygen is reduced to water (H2O) or hydro ...
. Confirmatory tests include production of the blue-green pigment pyocyanin on cetrimide agar and growth at 42 °C. A TSI slant
250px, TSI agar slant results: (from left) preinoculated (as control),
''P. aeruginosa'', ''E. coli'', '' Salmonella Typhimurium'', ''Shigella flexneri'' ">Shigella_flexneri.html" ;"title="Salmonella Typhimurium'', ''Shigella flexneri">Salmonel ...
is often used to distinguish nonfermenting ''Pseudomonas'' species from enteric pathogens in faecal specimens.
When ''P. aeruginosa'' is isolated from a normally sterile site (blood, bone, deep collections), it is generally considered dangerous, and almost always requires treatment. However, ''P. aeruginosa'' is frequently isolated from nonsterile sites (mouth swabs, sputum, etc.), and, under these circumstances, it may represent colonization and not infection. The isolation of ''P. aeruginosa'' from nonsterile specimens should, therefore, be interpreted cautiously, and the advice of a microbiologist or infectious diseases physician/pharmacist should be sought prior to starting treatment. Often, no treatment is needed.
Classification
Morphological, physiological, and biochemical characteristics of ''Pseudomonas aeruginosa'' are shown in the Table below. Note: + = Positive, - =Negative ''P. aeruginosa'' is a Gram-negative,aerobic
Aerobic means "requiring air," in which "air" usually means oxygen.
Aerobic may also refer to
* Aerobic exercise, prolonged exercise of moderate intensity
* Aerobics, a form of aerobic exercise
* Aerobic respiration, the aerobic process of cel ...
(and at times facultatively anaerobic
A facultative anaerobic organism is an organism that makes ATP by aerobic respiration if oxygen is present, but is capable of switching to fermentation if oxygen is absent.
Some examples of facultatively anaerobic bacteria are ''Staphylococcus' ...
), rod-shaped bacterium with unipolar motility. It has been identified as an opportunistic pathogen
An opportunistic infection is an infection caused by pathogens (bacteria, fungi, parasites or viruses) that take advantage of an opportunity not normally available. These opportunities can stem from a variety of sources, such as a weakened immune ...
of both humans and plants. ''P. aeruginosa'' is the type species
In zoological nomenclature, a type species (''species typica'') is the species name with which the name of a genus or subgenus is considered to be permanently taxonomically associated, i.e., the species that contains the biological type specimen ...
of the genus ''Pseudomonas
''Pseudomonas'' is a genus of Gram-negative, Gammaproteobacteria, belonging to the family Pseudomonadaceae and containing 191 described species. The members of the genus demonstrate a great deal of metabolic diversity and consequently are able ...
''.
Identification of ''P. aeruginosa'' can be complicated by the fact individual isolates often lack motility. The colony morphology itself also displays several varieties. The main two types are large, smooth, with a flat edge and elevated center and small, rough, and convex. A third type, mucoid, can also be found. The large colony can typically be found in clinal settings while the small is found in nature. The third, however, is present in biological settings and has been found in respiratory and in the urinary tract. Furthermore, mutations in the gene lasR drastically alter colony morphology and typically lead to failure to hydrolyze gelatin or hemolyze.
In certain conditions, ''P. aeruginosa'' can secrete a variety of pigments, including pyocyanin
Pyocyanin (PCN−) is one of the many toxic compounds produced and secreted by the Gram negative bacterium ''Pseudomonas aeruginosa''. Pyocyanin is a blue secondary metabolite, turning red below pH 4.9, with the ability to oxidise and reduce other ...
(blue), pyoverdine
Pyoverdines (alternatively, and less commonly, spelled as pyoverdins) are fluorescent siderophores produced by certain pseudomonads. Pyoverdines are important virulence factors, and are required for pathogenesis in many biological models of inf ...
(yellow and fluorescent
Fluorescence is the emission of light by a substance that has absorbed light or other electromagnetic radiation. It is a form of luminescence. In most cases, the emitted light has a longer wavelength, and therefore a lower photon energy, ...
), pyorubin (red), and pyomelanin (brown). These can be used to identify the organism.
 Clinical identification of ''P. aeruginosa'' may include identifying the production of both pyocyanin and fluorescein, as well as its ability to grow at 42 °C. ''P. aeruginosa'' is capable of growth in
Clinical identification of ''P. aeruginosa'' may include identifying the production of both pyocyanin and fluorescein, as well as its ability to grow at 42 °C. ''P. aeruginosa'' is capable of growth in diesel
Diesel may refer to:
* Diesel engine, an internal combustion engine where ignition is caused by compression
* Diesel fuel, a liquid fuel used in diesel engines
* Diesel locomotive, a railway locomotive in which the prime mover is a diesel engin ...
and jet fuels, where it is known as a hydrocarbon
In organic chemistry, a hydrocarbon is an organic compound consisting entirely of hydrogen and carbon. Hydrocarbons are examples of group 14 hydrides. Hydrocarbons are generally colourless and hydrophobic, and their odors are usually weak or ...
-using microorganism
A microorganism, or microbe,, ''mikros'', "small") and ''organism'' from the el, ὀργανισμός, ''organismós'', "organism"). It is usually written as a single word but is sometimes hyphenated (''micro-organism''), especially in olde ...
, causing microbial corrosion Microbial corrosion, also called microbiologically influenced corrosion (MIC), microbially induced corrosion (MIC) or biocorrosion, is "corrosion affected by the presence or activity (or both) of microorganisms in biofilms on the surface of the cor ...
. It creates dark, gellish mats sometimes improperly called "algae
Algae (; singular alga ) is an informal term for a large and diverse group of photosynthetic eukaryotic organisms. It is a polyphyletic grouping that includes species from multiple distinct clades. Included organisms range from unicellular mic ...
" because of their appearance.
Treatment
Many ''P. aeruginosa'' isolates are resistant to a large range of antibiotics and may demonstrate additional resistance after unsuccessful treatment. It should usually be possible to guide treatment according to laboratory sensitivities, rather than choosing an antibioticempirically
In philosophy, empiricism is an epistemological theory that holds that knowledge or justification comes only or primarily from sensory experience. It is one of several views within epistemology, along with rationalism and skepticism. Empir ...
. If antibiotics are started empirically, then every effort should be made to obtain cultures (before administering the first dose of antibiotic), and the choice of antibiotic used should be reviewed when the culture results are available.
 Due to widespread resistance to many common first-line antibiotics,
Due to widespread resistance to many common first-line antibiotics, carbapenems
Carbapenems are a class of very effective antibiotic agents most commonly used for the treatment of severe bacterial infections. This class of antibiotics is usually reserved for known or suspected multidrug-resistant (MDR) bacterial infections. ...
, polymyxins
Polymyxins are antibiotics. Polymyxins B and E (also known as colistin) are used in the treatment of Gram-negative bacterial infections. They work mostly by breaking up the bacterial cell membrane. They are part of a broader class of molecules ...
, and more recently tigecycline
Tigecycline, sold under the brand name Tygacil, is an tetracycline antibiotic medication for a number of bacterial infections. It is a glycylcycline administered intravenously. It was developed in response to the growing rate of antibiotic resist ...
were considered to be the drugs of choice; however, resistance to these drugs has also been reported. Despite this, they are still being used in areas where resistance has not yet been reported. Use of β-lactamase inhibitors such as sulbactam has been advised in combination with antibiotics to enhance antimicrobial action even in the presence of a certain level of resistance. Combination therapy after rigorous antimicrobial susceptibility testing has been found to be the best course of action in the treatment of multidrug-resistant ''P. aeruginosa''. Some next-generation antibiotics that are reported as being active against ''P. aeruginosa'' include doripenem, ceftobiprole, and ceftaroline. However, these require more clinical trials for standardization. Therefore, research for the discovery of new antibiotics and drugs against P. ''aeruginosa'' is very much needed.
Antibiotics that may have activity against ''P. aeruginosa'' include:
* aminoglycoside
Aminoglycoside is a medicinal and bacteriologic category of traditional Gram-negative antibacterial medications that inhibit protein synthesis and contain as a portion of the molecule an amino-modified glycoside (sugar). The term can also refer ...
s ( gentamicin, amikacin
Amikacin is an antibiotic medication used for a number of bacterial infections. This includes joint infections, intra-abdominal infections, meningitis, pneumonia, sepsis, and urinary tract infections. It is also used for the treatment of mu ...
, tobramycin
Tobramycin is an aminoglycoside antibiotic derived from '' Streptomyces tenebrarius'' that is used to treat various types of bacterial infections, particularly Gram-negative infections. It is especially effective against species of ''Pseudomonas ...
, but ''not'' kanamycin
Kanamycin A, often referred to simply as kanamycin, is an antibiotic used to treat severe bacterial infections and tuberculosis. It is not a first line treatment. It is used by mouth, injection into a vein, or injection into a muscle. Kanamycin ...
)
* quinolones
Quinolone may refer to:
* 2-Quinolone
* 4-Quinolone
4-Quinolone is an organic compound derived from quinoline. It and 2-quinolone are the two most important parent (meaning simplified) quinolones. 4-Quinolone exists in equilibrium with a mino ...
(ciprofloxacin
Ciprofloxacin is a fluoroquinolone antibiotic used to treat a number of bacterial infections. This includes bone and joint infections, intra abdominal infections, certain types of infectious diarrhea, respiratory tract infections, skin inf ...
, levofloxacin
Levofloxacin, sold under the brand name Levaquin among others, is an antibiotic medication. It is used to treat a number of bacterial infections including acute bacterial sinusitis, pneumonia, H. pylori (in combination with other medications), ...
, but not moxifloxacin
Moxifloxacin is an antibiotic, used to treat bacterial infections, including pneumonia, conjunctivitis, endocarditis, tuberculosis, and sinusitis. It can be given by mouth, by injection into a vein, and as an eye drop.
Common side effects in ...
)
*  cephalosporins (
cephalosporins (ceftazidime
Ceftazidime, sold under the brand name Fortaz among others, is a third-generation cephalosporin antibiotic useful for the treatment of a number of bacterial infections. Specifically it is used for joint infections, meningitis, pneumonia, sepsis, ...
, cefepime
Cefepime is a fourth-generation cephalosporin antibiotic. Cefepime has an extended spectrum of activity against Gram-positive and Gram-negative bacteria, with greater activity against both types of organism than third-generation agents. A 2007 ...
, cefoperazone
Cefoperazone is a third-generation cephalosporin antibiotic, marketed by Pfizer under the name Cefobid. It is one of few cephalosporin antibiotics effective in treating ''Pseudomonas'' bacterial infections which are otherwise resistant to these ...
, cefpirome, ceftobiprole
Ceftobiprole (Zevtera/Mabelio) is a fifth-generation cephalosporin for the treatment of hospital-acquired pneumonia (excluding ventilator-associated pneumonia) and community-acquired pneumonia. It is marketed by Basilea Pharmaceutica in the Unit ...
, but not cefuroxime
Cefuroxime, sold under the brand name Zinacef among others, is a second-generation cephalosporin antibiotic used to treat and prevent a number of bacterial infections. These include pneumonia, meningitis, otitis media, sepsis, urinary tract inf ...
, cefotaxime
Cefotaxime is an antibiotic used to treat a number of bacterial infections in human, other animals and plant tissue culture. Specifically in humans it is used to treat joint infections, pelvic inflammatory disease, meningitis, pneumonia, urin ...
, or ceftriaxone
Ceftriaxone, sold under the brand name Rocephin, is a third-generation cephalosporin antibiotic used for the treatment of a number of bacterial infections. These include middle ear infections, endocarditis, meningitis, pneumonia, bone and joint ...
)
* antipseudomonal penicillins: carboxypenicillin
The carboxypenicillins are a group of antibiotics. They belong to the penicillin family and comprise the members carbenicillin and ticarcillin.
Chemical structure
The carboxypenicillins feature the beta-lactam backbone of all penicillins b ...
s (carbenicillin
Carbenicillin is a bactericidal antibiotic belonging to the carboxypenicillin subgroup of the penicillins. It was discovered by scientists at Beecham and marketed as Pyopen. It has Gram-negative coverage which includes ''Pseudomonas aeruginosa'' ...
and ticarcillin
Ticarcillin is a carboxypenicillin. It can be sold and used in combination with clavulanate as ticarcillin/clavulanic acid. Because it is a penicillin, it also falls within the larger class of beta-lactam antibiotics. Its main clinical use is as ...
), and ureidopenicillin
The ureidopenicillins are a group of penicillins which are active against ''Pseudomonas aeruginosa''.
There are three ureidopenicillins in clinical use:
*Azlocillin
*Piperacillin
*Mezlocillin
They are mostly ampicillin derivatives in which the ...
s (mezlocillin
Mezlocillin is a broad-spectrum penicillin antibiotic. It is active against both Gram-negative and some Gram-positive bacteria. Unlike most other extended spectrum penicillins, it is excreted by the liver, therefore it is useful for biliary tract ...
, azlocillin
Azlocillin is an acyl ampicillin antibiotic with an extended spectrum of activity and greater ''in vitro'' potency than the carboxy penicillins.
Azlocillin is similar to mezlocillin and piperacillin. It demonstrates antibacterial activity ...
, and piperacillin
Piperacillin is a broad-spectrum β-lactam antibiotic of the ureidopenicillin class. The chemical structure of piperacillin and other ureidopenicillins incorporates a polar side chain that enhances penetration into Gram-negative bacteria and red ...
). ''P. aeruginosa'' is intrinsically resistant to all other penicillin
Penicillins (P, PCN or PEN) are a group of β-lactam antibiotics originally obtained from ''Penicillium'' moulds, principally '' P. chrysogenum'' and '' P. rubens''. Most penicillins in clinical use are synthesised by P. chrysogenum using ...
s.
* carbapenem
Carbapenems are a class of very effective antibiotic agents most commonly used for the treatment of severe bacterial infections. This class of antibiotics is usually reserved for known or suspected multidrug-resistant (MDR) bacterial infections. ...
s (meropenem
Meropenem, sold under the brand name Merrem among others, is an intravenous β-lactam antibiotic used to treat a variety of bacterial infections. Some of these include meningitis, intra-abdominal infection, pneumonia, sepsis, and anthrax.
...
, imipenem
Imipenem (trade name Primaxin among others) is an intravenous β-lactam antibiotic discovered by Merck scientists Burton Christensen, William Leanza, and Kenneth Wildonger in the mid-1970s. Carbapenems are highly resistant to the β-lactamase enz ...
, doripenem
Doripenem (Doribax, Finibax) is an antibiotic drug in the carbapenem class. It is a beta-lactam antibiotic drug able to kill ''Pseudomonas aeruginosa''.
Doripenem can be used for bacterial infections such as: complex abdominal infections, pneumon ...
, but not ertapenem
Ertapenem, sold under the brand name Invanz, is a carbapenem antibiotic medication used for the treatment of infections of the abdomen, the lungs, the upper part of the female reproductive system, and the diabetic foot.
The most common side eff ...
)
* polymyxin
Polymyxins are antibiotics. Polymyxins B and E (also known as colistin) are used in the treatment of Gram-negative bacterial infections. They work mostly by breaking up the bacterial cell membrane. They are part of a broader class of molecules ...
s (polymyxin B
Polymyxin B, sold under the brand name Poly-Rx among others, is an antibiotic used to treat meningitis, pneumonia, sepsis, and urinary tract infections. While it is useful for many Gram negative infections, it is not useful for Gram positive inf ...
and colistin
Colistin, also known as polymyxin E, is an antibiotic medication used as a last-resort treatment for multidrug-resistant Gram-negative infections including pneumonia. These may involve bacteria such as ''Pseudomonas aeruginosa'', '' Klebsiella ...
)
* monobactam
Monobactams are monocyclic and bacterially-produced β-lactam antibiotics. The β-lactam ring is not fused to another ring, in contrast to most other β-lactams. Monobactams are effective only against aerobic Gram-negative bacteria (e.g., ''Nei ...
s (aztreonam
Aztreonam, sold under the brand name Azactam among others, is an antibiotic used primarily to treat infections caused by gram-negative bacteria such as ''Pseudomonas aeruginosa''. This may include bone infections, endometritis, intra abdominal ...
)
As fluoroquinolones are one of the few antibiotic classes widely effective against ''P. aeruginosa'', in some hospitals, their use is severely restricted to avoid the development of resistant strains. On the rare occasions where infection is superficial and limited (for example, ear infections or nail infections), topical
A topical medication is a medication that is applied to a particular place on or in the body. Most often topical medication means application to body surfaces such as the skin or mucous membranes to treat ailments via a large range of classes ...
gentamicin or colistin may be used.
For pseudomonal wound infections, acetic acid with concentrations from 0.5% to 5% can be an effective bacteriostatic agent in eliminating the bacteria from the wound. Usually a sterile gauze soaked with acetic acid is placed on the wound after irrigation with normal saline. Dressing would be done once per day. Pseudomonas is usually eliminated in 90% of the cases after 10 to 14 days of treatment.
Antibiotic resistance
One of the most worrisome characteristics of ''P. aeruginosa'' is its low antibiotic susceptibility, which is attributable to a concerted action of multidrug efflux pumps with chromosomally encoded antibiotic resistance genes (e.g., ''mexAB'', ''mexXY'', etc.) and the low permeability of the bacterial cellular envelopes. In addition to this intrinsic resistance, ''P. aeruginosa'' easily develops acquired resistance either bymutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mi ...
in chromosomally encoded genes or by the horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). H ...
of antibiotic resistance determinants. Development of multidrug resistance
Multiple drug resistance (MDR), multidrug resistance or multiresistance is antimicrobial resistance shown by a species of microorganism to at least one antimicrobial drug in three or more antimicrobial categories. Antimicrobial categories are c ...
by ''P. aeruginosa'' isolates requires several different genetic events, including acquisition of different mutations and/or horizontal transfer of antibiotic resistance genes. Hypermutation favours the selection of mutation-driven antibiotic resistance in ''P. aeruginosa'' strains producing chronic infections, whereas the clustering of several different antibiotic resistance genes in integron Integrons are genetic mechanisms that allow bacteria to adapt and evolve rapidly through the stockpiling and expression of new genes. These genes are embedded in a specific genetic structure called gene cassette (a term that is lately changing to in ...
s favors the concerted acquisition of antibiotic resistance determinants. Some recent studies have shown phenotypic resistance associated to biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
formation or to the emergence of small-colony variants may be important in the response of ''P. aeruginosa'' populations to antibiotic treatment.
Mechanisms underlying antibiotic resistance have been found to include production of antibiotic-degrading or antibiotic-inactivating enzymes, outer membrane proteins to evict the antibiotics, and mutations to change antibiotic targets. Presence of antibiotic-degrading enzymes such as extended-spectrum β-lactamases like PER-1, PER-2, and VEB-1, AmpC cephalosporinases, carbapenemases like serine oxacillinases, metallo-b-lactamases, OXA-type carbapenemases, and aminoglycoside-modifying enzymes, among others, have been reported. ''P. aeruginosa'' can also modify the targets of antibiotic action: for example, methylation of 16S rRNA to prevent aminoglycoside binding and modification of DNA, or topoisomerase to protect it from the action of quinolones. ''P. aeruginosa'' has also been reported to possess multidrug efflux pumps systems that confer resistance against a number of antibiotic classes, and the MexAB-OprM ( Resistance-nodulation-division (RND) family) is considered as the most important''.'' An important factor found to be associated with antibiotic resistance is the decrease in the virulence capabilities of the resistant strain. Such findings have been reported in the case of rifampicin-resistant and colistin-resistant strains, in which decrease in infective ability, quorum sensing, and motility have been documented.
Mutations in DNA gyrase
DNA gyrase, or simply gyrase, is an enzyme within the class of topoisomerase and is a subclass of Type II topoisomerases that reduces topological strain in an ATP dependent manner while double-stranded DNA is being unwound by elongating RNA-poly ...
are commonly associated with antibiotic resistance in ''P. aeruginosa''. These mutations, when combined with others, confer high resistance without hindering survival. Additionally, genes involved in cyclic-di-GMP signaling may contribute to resistance. When ''P. aeruginosa'' is grown under ''in vitro'' conditions designed to mimic a cystic fibrosis patient's lungs, these genes mutate repeatedly.
Two small RNA
Small RNA (sRNA) are polymeric RNA molecules that are less than 200 nucleotides in length, and are usually non-coding. RNA silencing is often a function of these molecules, with the most common and well-studied example being RNA interference (RN ...
s, Sr0161 and ErsA, were shown to interact with mRNA encoding the major porin OprD responsible for the uptake of carbapenem antibiotics
Carbapenems are a class of very effective antibiotic agents most commonly used for the treatment of severe bacterial infections. This class of antibiotics is usually reserved for known or suspected multidrug-resistant (MDR) bacterial infections. ...
into the periplasm
The periplasm is a concentrated gel-like matrix in the space between the inner cytoplasmic membrane and the bacterial outer membrane called the ''periplasmic space'' in gram-negative bacteria. Using cryo-electron microscopy it has been found that ...
. The sRNAs bind to the 5'UTR of ''oprD'', causing increase in bacterial resistance to meropenem
Meropenem, sold under the brand name Merrem among others, is an intravenous β-lactam antibiotic used to treat a variety of bacterial infections. Some of these include meningitis, intra-abdominal infection, pneumonia, sepsis, and anthrax.
...
. Another sRNA, Sr006, may positively regulate (post-transcriptionally) the expression of PagL, an enzyme responsible for deacylation of lipid A. This reduces the pro-inflammatory property of lipid A. Furthermore, similar to a process found in ''Salmonella'', Sr006 regulation of PagL expression may aid in polymyxin B
Polymyxin B, sold under the brand name Poly-Rx among others, is an antibiotic used to treat meningitis, pneumonia, sepsis, and urinary tract infections. While it is useful for many Gram negative infections, it is not useful for Gram positive inf ...
resistance.
Prevention
Probiotic prophylaxis may prevent colonization and delay onset of ''Pseudomonas'' infection in an ICU setting. Immunoprophylaxis against ''Pseudomonas'' is being investigated. The risk of contracting ''P. aeruginosa'' can be reduced by avoiding pools, hot tubs, and other bodies of standing water; regularly disinfecting and/or replacing equipment that regularly encounters moisture (such as contact lens equipment and solutions); and washing one's hands often (which is protective against many other pathogens as well). However, even the best hygiene practices cannot totally protect an individual against ''P. aeruginosa,'' given how common ''P. aeruginosa'' is in the environment.Experimental therapies
Phage therapy
Phage therapy, viral phage therapy, or phagotherapy is the therapeutic use of bacteriophages for the treatment of pathogenic bacterial infections. This therapeutic approach emerged at the beginning of the 20th century but was progressively re ...
against ''P. aeruginosa'' has been investigated as a possible effective treatment, which can be combined with antibiotics, has no contraindications and minimal adverse effects. Phages are produced as sterile liquid, suitable for intake, applications etc.
Phage therapy against ear infections caused by ''P. aeruginosa'' was reported in the journal ''Clinical Otolaryngology'' in August 2009.
Research
In 2013, João Xavier described an experiment in which ''P. aeruginosa'', when subjected to repeated rounds of conditions in which it needed to swarm to acquire food, developed the ability to "hyperswarm" at speeds 25% faster than baseline organisms, by developing multiple flagella, whereas the baseline organism has a single flagellum. This result was notable in the field ofexperimental evolution
Experimental evolution is the use of laboratory experiments or controlled field manipulations to explore evolutionary dynamics. Evolution may be observed in the laboratory as individuals/populations adapt to new environmental conditions by natura ...
in that it was highly repeatable.
''P. aeruginosa'' has been studied for use in bioremediation
Bioremediation broadly refers to any process wherein a biological system (typically bacteria, microalgae, fungi, and plants), living or dead, is employed for removing environmental pollutants from air, water, soil, flue gasses, industrial effluent ...
and use in processing polyethylene
Polyethylene or polythene (abbreviated PE; IUPAC name polyethene or poly(methylene)) is the most commonly produced plastic. It is a polymer, primarily used for packaging ( plastic bags, plastic films, geomembranes and containers including bo ...
in municipal solid waste
Municipal solid waste (MSW), commonly known as trash or garbage in the United States and rubbish in Britain, is a waste type consisting of everyday items that are discarded by the public. "Garbage" can also refer specifically to food waste, ...
.
Distribution
Pest risk analysis
theEast African Community
The East African Community (EAC) is an intergovernmental organisation composed of seven countries in the Great Lakes region of East Africa: the Democratic Republic of the Congo, the United Republic of Tanzania, the Republics of Kenya, Burun ...
considers ''P. aeruginosa'' to be a quarantine concern. The presence of '' Phaseolus vulgaris''-pathogenic strains of ''P. aeruginosa'' in Kenya
)
, national_anthem = "Ee Mungu Nguvu Yetu"()
, image_map =
, map_caption =
, image_map2 =
, capital = Nairobi
, coordinates =
, largest_city = Nairobi
, ...
for the rest of the area. A pest risk analysis by the EAC was based on this bacterium's CABI's Crop Protection Compendium listing, following Kaaya & Darji 1989's initial detection in Kenya.
See also
*Bacteriological water analysis
Bacteriological water analysis is a method of analysing water to estimate the numbers of bacteria present and, if needed, to find out what sort of bacteria they are. It represents one aspect of water quality. It is a microbiological analytical p ...
* Contamination control
Contamination control is the generic term for all activities aiming to control the existence, growth and proliferation of contamination in certain areas. Contamination control may refer to the atmosphere as well as to surfaces, to particulate matte ...
* Nosocomial infection
A hospital-acquired infection, also known as a nosocomial infection (from the Greek , meaning "hospital"), is an infection that is acquired in a hospital or other health care facility. To emphasize both hospital and nonhospital settings, it is so ...
* NrsZ small RNA
NrsZ (nitrogen regulated small RNA) is a bacterial small RNA found in the opportunistic pathogen ''Pseudomonas aeruginosa'' PAO1. Its transcription is induced during nitrogen limitation by the NtrB/C two-component system (an important regulator of ...
* AsponA antisense RNA AsponA is a small asRNA transcribed antisense to the penicillin-binding protein 1A gene called ''ponA.'' It was identified by RNAseq and the expression was validated by 5' and 3' RACE experiments in '' Pseudomanas aeruginosa''. ''AsponA'' express ...
* Repression of heat shock gene expression (ROSE) element
The repression of heat shock gene expression (ROSE) element is an RNA element found in the 5' UTR of some heat shock protein's mRNAs. The ROSE element is an RNA thermometer that negatively regulates heat shock gene expression. The secondary st ...
* Pseudomon-1 RNA motif (ErsA sRNA)
* PrrF RNA
The PrrF RNAs are small non-coding RNAs involved in iron homeostasis and are encoded by all ''Pseudomonas'' species. The PrrF RNAs are analogs of the RyhB RNA, which is encoded by enteric bacteria. Expression of the PrrF RNAs is repressed by the ...
* ''Pseudomonas'' sRNA P16 (RgsA sRNA)
References
External links
Type strain of ''Pseudomonas aeruginosa'' at Bac''Dive'' - the Bacterial Diversity Metadatabase
* Johanna M. Sweere ''et al.'' (2019)
Bacteriophage trigger antiviral immunity and prevent clearance of bacterial infection
Science 29 Mar 2019: Vol. 363, Issue 6434, eaat9691. doi:10.1126/science.aat9691 about Pseudomonas aeruginosa filamentous phages (Pf-phages), ''
Inoviridae
Filamentous bacteriophage is a family of viruses (''Inoviridae'') that infect bacteria. The phages are named for their filamentous shape, a worm-like chain (long, thin and flexible, reminiscent of a length of cooked spaghetti), about 6 nm ...
''. See also:
:UM researchers publish new discoveries on bacterial viruses
On: EurekAlert! 1 Apr 2019. Source: University of Montana {{Authority control Antibiotic-resistant bacteria Bacteria described in 1872 Gram-negative bacteria Pseudomonadales