Chromatin on:
[Wikipedia]
[Google]
[Amazon]
 Chromatin is a complex of DNA and
Chromatin is a complex of DNA and

 Chromatin undergoes various structural changes during a
Chromatin undergoes various structural changes during a
 In nature, DNA can form three structures, A-, B-, and Z-DNA. A- and B-DNA are very similar, forming right-handed helices, whereas Z-DNA is a left-handed helix with a zig-zag phosphate backbone. Z-DNA is thought to play a specific role in chromatin structure and transcription because of the properties of the junction between B- and Z-DNA.
At the junction of B- and Z-DNA, one pair of bases is flipped out from normal bonding. These play a dual role of a site of recognition by many proteins and as a sink for torsional stress from
In nature, DNA can form three structures, A-, B-, and Z-DNA. A- and B-DNA are very similar, forming right-handed helices, whereas Z-DNA is a left-handed helix with a zig-zag phosphate backbone. Z-DNA is thought to play a specific role in chromatin structure and transcription because of the properties of the junction between B- and Z-DNA.
At the junction of B- and Z-DNA, one pair of bases is flipped out from normal bonding. These play a dual role of a site of recognition by many proteins and as a sink for torsional stress from
 The basic repeat element of chromatin is the nucleosome, interconnected by sections of
The basic repeat element of chromatin is the nucleosome, interconnected by sections of
 With addition of H1, the beads-on-a-string structure in turn coils into a 30 nm diameter helical structure known as the 30 nm fibre or filament. The precise structure of the chromatin fiber in the cell is not known in detail.
This level of chromatin structure is thought to be the form of
With addition of H1, the beads-on-a-string structure in turn coils into a 30 nm diameter helical structure known as the 30 nm fibre or filament. The precise structure of the chromatin fiber in the cell is not known in detail.
This level of chromatin structure is thought to be the form of  The existing models commonly accept that the nucleosomes lie perpendicular to the axis of the fibre, with linker histones arranged internally.
A stable 30 nm fibre relies on the regular positioning of nucleosomes along DNA. Linker DNA is relatively resistant to bending and rotation. This makes the length of linker DNA critical to the stability of the fibre, requiring nucleosomes to be separated by lengths that permit rotation and folding into the required orientation without excessive stress to the DNA.
In this view, different lengths of the linker DNA should produce different folding topologies of the chromatin fiber. Recent theoretical work, based on electron-microscopy images
of reconstituted fibers supports this view.
The existing models commonly accept that the nucleosomes lie perpendicular to the axis of the fibre, with linker histones arranged internally.
A stable 30 nm fibre relies on the regular positioning of nucleosomes along DNA. Linker DNA is relatively resistant to bending and rotation. This makes the length of linker DNA critical to the stability of the fibre, requiring nucleosomes to be separated by lengths that permit rotation and folding into the required orientation without excessive stress to the DNA.
In this view, different lengths of the linker DNA should produce different folding topologies of the chromatin fiber. Recent theoretical work, based on electron-microscopy images
of reconstituted fibers supports this view.

 # Interphase: The structure of chromatin during
# Interphase: The structure of chromatin during
 # ChIP-seq (Chromatin immunoprecipitation sequencing) is recognized as the vastly utilized chromatin identification method it has been using the antibodies that actively selected, identify and combine with proteins including "histones, histone restructuring, transaction factors and cofactors". This has been providing data about the state of chromatin and the transaction of a gene by trimming "oligonucleotides" that are unbound. Chromatin immunoprecipitation sequencing aimed against different histone modifications, can be used to identify chromatin states throughout the genome. Different modifications have been linked to various states of chromatin.
#
# ChIP-seq (Chromatin immunoprecipitation sequencing) is recognized as the vastly utilized chromatin identification method it has been using the antibodies that actively selected, identify and combine with proteins including "histones, histone restructuring, transaction factors and cofactors". This has been providing data about the state of chromatin and the transaction of a gene by trimming "oligonucleotides" that are unbound. Chromatin immunoprecipitation sequencing aimed against different histone modifications, can be used to identify chromatin states throughout the genome. Different modifications have been linked to various states of chromatin.
#
Chromosomes and Chromatin.
* * Cremer, T. 1985. Von der Zellenlehre zur Chromosomentheorie: Naturwissenschaftliche Erkenntnis und Theorienwechsel in der frühen Zell- und Vererbungsforschung, Veröffentlichungen aus der Forschungsstelle für Theoretische Pathologie der Heidelberger Akademie der Wissenschaften. Springer-Vlg., Berlin, Heidelberg. * Elgin, S. C. R. (ed.). 1995. Chromatin Structure and Gene Expression, vol. 9. IRL Press, Oxford, New York, Tokyo. * * * * * * Pollard, T., and W. Earnshaw. 2002. Cell Biology. Saunders. * Saumweber, H. 1987. Arrangement of Chromosomes in Interphase Cell Nuclei, p. 223-234. In W. Hennig (ed.), Structure and Function of Eucaryotic Chromosomes, vol. 14. Springer-Verlag, Berlin, Heidelberg. * * Van Holde KE. 1989. Chromatin. New York:
Chromatin, Histones & Cathepsin
PMAP
''Nature'' journal: recent chromatin publications and news
Protocol for ''in vitro'' Chromatin Assembly
ENCODE threads Explorer
Chromatin patterns at transcription factor binding sites.
 Chromatin is a complex of DNA and
Chromatin is a complex of DNA and protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
found in eukaryotic
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bacte ...
cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important roles in reinforcing the DNA during cell division
Cell division is the process by which a parent cell divides into two daughter cells. Cell division usually occurs as part of a larger cell cycle in which the cell grows and replicates its chromosome(s) before dividing. In eukaryotes, there ...
, preventing DNA damage
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA d ...
, and regulating gene expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. T ...
and DNA replication
In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part for biological inheritan ...
. During mitosis
In cell biology, mitosis () is a part of the cell cycle in which replicated chromosomes are separated into two new nuclei. Cell division by mitosis gives rise to genetically identical cells in which the total number of chromosomes is maintai ...
and meiosis
Meiosis (; , since it is a reductional division) is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately ...
, chromatin facilitates proper segregation of the chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins ar ...
s in anaphase
Anaphase () is the stage of mitosis after the process of metaphase, when replicated chromosomes are split and the newly-copied chromosomes (daughter chromatids) are moved to opposite poles of the cell. Chromosomes also reach their overall maxim ...
; the characteristic shapes of chromosomes visible during this stage are the result of DNA being coiled into highly condensed chromatin.
The primary protein components of chromatin are histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn a ...
s. An octamer
In chemistry and biochemistry, an oligomer () is a molecule that consists of a few repeating units which could be derived, actually or conceptually, from smaller molecules, monomers.Quote: ''Oligomer molecule: A molecule of intermediate relati ...
of two sets of four histone cores (Histone H2A
Histone H2A is one of the five main histone proteins involved in the structure of chromatin in eukaryotic cells.
The other histone proteins are: H1, H2B, H3 and H4.
Background
Histones are proteins that package DNA into nucleosomes. Hist ...
, Histone H2B
Histone H2B is one of the 5 main histone proteins involved in the structure of chromatin in eukaryotic cells. Featuring a main globular domain and long N-terminal and C-terminal tails, H2B is involved with the structure of the nucleosomes.
S ...
, Histone H3
Histone H3 is one of the five main histones involved in the structure of chromatin in eukaryotic cells. Featuring a main globular domain and a long N-terminal tail, H3 is involved with the structure of the nucleosomes of the 'beads on a st ...
, and Histone H4
Histone H4 is one of the five main histone proteins involved in the structure of chromatin in eukaryotic cells. Featuring a main globular domain and a long N-terminal tail, H4 is involved with the structure of the nucleosome of the 'beads on ...
) bind to DNA and function as "anchors" around which the strands are wound.Maeshima, K., Ide, S., & Babokhov, M. (2019). Dynamic chromatin organization without the 30-nm fiber. ''Current opinion in cell biology, 58,'' 95–104. https://doi.org/10.1016/j.ceb.2019.02.003 In general, there are three levels of chromatin organization:
# DNA wraps around histone proteins, forming nucleosome
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundame ...
s and the so-called beads on a string
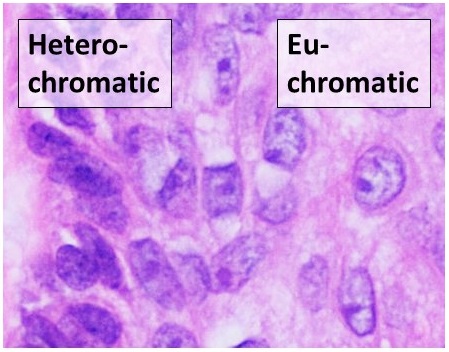
Euchromatin (also called "open chromatin") is a lightly packed form of chromatin ( DNA, RNA, and protein) that is enriched in genes, and is often (but not always) under active transcription. Euchromatin stands in contrast to heterochromatin, whi ...
structure (euchromatin
Euchromatin (also called "open chromatin") is a lightly packed form of chromatin ( DNA, RNA, and protein) that is enriched in genes, and is often (but not always) under active transcription. Euchromatin stands in contrast to heterochromatin, whi ...
).
# Multiple histones wrap into a 30-nanometer
330px, Different lengths as in respect to the molecular scale.
The nanometre (international spelling as used by the International Bureau of Weights and Measures; SI symbol: nm) or nanometer (American and British English spelling differences#-re, ...
fiber consisting of nucleosome arrays in their most compact form (heterochromatin
Heterochromatin is a tightly packed form of DNA or '' condensed DNA'', which comes in multiple varieties. These varieties lie on a continue between the two extremes of constitutive heterochromatin and facultative heterochromatin. Both play a rol ...
).
# Higher-level DNA supercoiling of the 30-nm fiber produces the metaphase
Metaphase ( and ) is a stage of mitosis in the eukaryotic cell cycle in which chromosomes are at their second-most condensed and coiled stage (they are at their most condensed in anaphase). These chromosomes, carrying genetic information, a ...
chromosome (during mitosis and meiosis).
Many organisms, however, do not follow this organization scheme. For example, spermatozoa
A spermatozoon (; also spelled spermatozoön; ; ) is a motile sperm cell, or moving form of the haploid cell that is the male gamete. A spermatozoon joins an ovum to form a zygote. (A zygote is a single cell, with a complete set of chromos ...
and avian
Avian may refer to:
* Birds or Aves, winged animals
*Avian (given name) (russian: Авиа́н, link=no), a male forename
Aviation
*Avro Avian, a series of light aircraft made by Avro in the 1920s and 1930s
*Avian Limited, a hang glider manufactur ...
red blood cell
Red blood cells (RBCs), also referred to as red cells, red blood corpuscles (in humans or other animals not having nucleus in red blood cells), haematids, erythroid cells or erythrocytes (from Greek ''erythros'' for "red" and ''kytos'' for "hol ...
s have more tightly packed chromatin than most eukaryotic cells, and trypanosomatid
Trypanosomatida is a group of kinetoplastid excavates distinguished by having only a single flagellum. The name is derived from the Greek ''trypano'' (borer) and ''soma'' (body) because of the corkscrew-like motion of some trypanosomatid species ...
protozoa
Protozoa (singular: protozoan or protozoon; alternative plural: protozoans) are a group of single-celled eukaryotes, either free-living or parasitic, that feed on organic matter such as other microorganisms or organic tissues and debris. Histo ...
do not condense
Condensation is the change of the state of matter from the gas phase into the liquid phase, and is the reverse of vaporization. The word most often refers to the water cycle. It can also be defined as the change in the state of water vapor to ...
their chromatin into visible chromosomes at all. Prokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
cells have entirely different structures for organizing their DNA (the prokaryotic chromosome equivalent is called a genophore
The nucleoid (meaning ''nucleus-like'') is an irregularly shaped region within the prokaryotic cell that contains all or most of the genetic material. The chromosome of a prokaryote is circular, and its length is very large compared to the cell d ...
and is localized within the nucleoid
The nucleoid (meaning ''nucleus-like'') is an irregularly shaped region within the prokaryotic cell that contains all or most of the genetic material. The chromosome of a prokaryote is circular, and its length is very large compared to the cell ...
region).
The overall structure of the chromatin network further depends on the stage of the cell cycle
The cell cycle, or cell-division cycle, is the series of events that take place in a cell that cause it to divide into two daughter cells. These events include the duplication of its DNA (DNA replication) and some of its organelles, and sub ...
. During interphase
Interphase is the portion of the cell cycle that is not accompanied by visible changes under the microscope, and includes the G1, S and G2 phases. During interphase, the cell grows (G1), replicates its DNA (S) and prepares for mitosis (G2). A c ...
, the chromatin is structurally loose to allow access to RNA and DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to crea ...
s that transcribe and replicate the DNA. The local structure of chromatin during interphase depends on the specific gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
s present in the DNA. Regions of DNA containing genes which are actively transcribed ("turned on") are less tightly compacted and closely associated with RNA polymerases in a structure known as euchromatin
Euchromatin (also called "open chromatin") is a lightly packed form of chromatin ( DNA, RNA, and protein) that is enriched in genes, and is often (but not always) under active transcription. Euchromatin stands in contrast to heterochromatin, whi ...
, while regions containing inactive genes ("turned off") are generally more condensed and associated with structural proteins in heterochromatin
Heterochromatin is a tightly packed form of DNA or '' condensed DNA'', which comes in multiple varieties. These varieties lie on a continue between the two extremes of constitutive heterochromatin and facultative heterochromatin. Both play a rol ...
. Epigenetic
In biology, epigenetics is the study of stable phenotypic changes (known as ''marks'') that do not involve alterations in the DNA sequence. The Greek prefix '' epi-'' ( "over, outside of, around") in ''epigenetics'' implies features that are ...
modification of the structural proteins in chromatin via methylation
In the chemical sciences, methylation denotes the addition of a methyl group on a substrate, or the substitution of an atom (or group) by a methyl group. Methylation is a form of alkylation, with a methyl group replacing a hydrogen atom. These ...
and acetylation
:
In organic chemistry, acetylation is an organic esterification reaction with acetic acid. It introduces an acetyl group into a chemical compound. Such compounds are termed ''acetate esters'' or simply '' acetates''. Deacetylation is the oppos ...
also alters local chromatin structure and therefore gene expression. There is limited understanding of chromatin structure and it is active area of research in molecular biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and phys ...
.
Dynamic chromatin structure and hierarchy
cell cycle
The cell cycle, or cell-division cycle, is the series of events that take place in a cell that cause it to divide into two daughter cells. These events include the duplication of its DNA (DNA replication) and some of its organelles, and sub ...
. Histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn a ...
proteins are the basic packers and arrangers of chromatin and can be modified by various post-translational modifications to alter chromatin packing ( histone modification). Most modifications occur on histone tails. The positively charged histone cores only partially counteract the negative charge of the DNA phosphate backbone resulting in a negative net charge of the overall structure. An imbalance of charge within the polymer causes electrostatic
Electrostatics is a branch of physics that studies electric charges at rest ( static electricity).
Since classical times, it has been known that some materials, such as amber, attract lightweight particles after rubbing. The Greek word for ...
repulsion between neighboring chromatin regions that promote interactions with positively charged proteins, molecules, and cations. As these modifications occur, the electrostatic environment surrounding the chromatin will flux and the level of chromatin compaction will alter. The consequences in terms of chromatin accessibility and compaction depend both on the modified amino acid and the type of modification. For example, histone acetylation results in loosening and increased accessibility of chromatin for replication and transcription. Lysine trimethylation can either lead to increased transcriptional activity (trimethylation of histone H3
Histone H3 is one of the five main histones involved in the structure of chromatin in eukaryotic cells. Featuring a main globular domain and a long N-terminal tail, H3 is involved with the structure of the nucleosomes of the 'beads on a st ...
lysine 4) or transcriptional repression and chromatin compaction (trimethylation of histone H3 lysine 9 or 27). Several studies suggested that different modifications could occur simultaneously. For example, it was proposed that a bivalent Bivalent may refer to:
* Bivalent (chemistry), a molecule formed from two or more atoms bound together
*Bivalent (engine), an engine that can operate on two different types of fuel
*Bivalent (genetics), a pair of homologous chromosomes
*Bivalent log ...
structure (with trimethylation of both lysine 4 and 27 on histone H3) is involved in early mammalian development. Another study tested the role of H4K16ac
H4K16ac is an epigenetic modification to the DNA packaging protein Histone H4. It is a mark that indicates the acetylation at the 16th lysine residue of the histone H4 protein.
H4K16ac is unusual in that it has both transcriptional activation ...
on chromatin structure and found that homogeneous
Homogeneity and heterogeneity are concepts often used in the sciences and statistics relating to the uniformity of a substance or organism. A material or image that is homogeneous is uniform in composition or character (i.e. color, shape, siz ...
acetylation inhibited 30 nm chromatin formation and blocked adenosine triphosphate
Adenosine triphosphate (ATP) is an organic compound that provides energy to drive many processes in living cells, such as muscle contraction, nerve impulse propagation, condensate dissolution, and chemical synthesis. Found in all known forms ...
remodeling. This singular modification changed the dynamics of the chromatin which shows that acetylation of H4 at K16 is vital for proper intra- and inter- functionality of chromatin structure.
Polycomb-group proteins play a role in regulating genes through modulation of chromatin structure.
For additional information, see Chromatin variant, Histone modifications in chromatin regulation and RNA polymerase control by chromatin structure
RNA polymerase II (RNAP II and Pol II) is a multiprotein complex that transcribes DNA into precursors of messenger RNA (mRNA) and most small nuclear RNA (snRNA) and microRNA. It is one of the three RNAP enzymes found in the nucleus of eukaryotic ...
.
DNA structure
 In nature, DNA can form three structures, A-, B-, and Z-DNA. A- and B-DNA are very similar, forming right-handed helices, whereas Z-DNA is a left-handed helix with a zig-zag phosphate backbone. Z-DNA is thought to play a specific role in chromatin structure and transcription because of the properties of the junction between B- and Z-DNA.
At the junction of B- and Z-DNA, one pair of bases is flipped out from normal bonding. These play a dual role of a site of recognition by many proteins and as a sink for torsional stress from
In nature, DNA can form three structures, A-, B-, and Z-DNA. A- and B-DNA are very similar, forming right-handed helices, whereas Z-DNA is a left-handed helix with a zig-zag phosphate backbone. Z-DNA is thought to play a specific role in chromatin structure and transcription because of the properties of the junction between B- and Z-DNA.
At the junction of B- and Z-DNA, one pair of bases is flipped out from normal bonding. These play a dual role of a site of recognition by many proteins and as a sink for torsional stress from RNA polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template.
Using the enzyme helicase, RNAP locally opens th ...
or nucleosome binding.DNA bases are stored as a code structure with four chemical bases such as ''“Adenine (A), Guanine (G), Cytosine (C), and Thymine (T)”''. The order and sequences of these chemical structures of DNA are reflected as information available for the creation and control of human organisms. ''“A with T and C with G”'' pairing up to build the DNA base pair. ''Sugar and phosphate'' molecules are also paired with these bases, making DNA nucleotides arrange 2 long spiral strands unitedly called ''“double helix”''. In eukaryotes, DNA consists of a cell nucleus and the DNA is providing strength and direction to the mechanism of heredity. Moreover, between the nitrogenous bonds of the 2 DNA, homogenous bonds are forming.
Nucleosomes and beads-on-a-string
 The basic repeat element of chromatin is the nucleosome, interconnected by sections of
The basic repeat element of chromatin is the nucleosome, interconnected by sections of linker DNA
In molecular biology, linker DNA is double-stranded DNA (38-53 base pairs long) in between two nucleosome cores that, in association with histone H1, holds the cores together. Linker DNA is seen as the string in the "beads and string model", whi ...
, a far shorter arrangement than pure DNA in solution.
In addition to core histones, a linker histone H1
Histone H1 is one of the five main histone protein families which are components of chromatin in eukaryotic cells. Though highly conserved, it is nevertheless the most variable histone in sequence across species.
Structure
Metazoan H1 prote ...
exists that contacts the exit/entry of the DNA strand on the nucleosome. The nucleosome core particle, together with histone H1, is known as a chromatosome. Nucleosomes, with about 20 to 60 base pairs of linker DNA, can form, under non-physiological conditions, an approximately 10 nm beads on a string
Euchromatin (also called "open chromatin") is a lightly packed form of chromatin ( DNA, RNA, and protein) that is enriched in genes, and is often (but not always) under active transcription. Euchromatin stands in contrast to heterochromatin, whi ...
fibre.
The nucleosomes bind DNA non-specifically, as required by their function in general DNA packaging. There are, however, large DNA sequence preferences that govern nucleosome positioning. This is due primarily to the varying physical properties of different DNA sequences: For instance, adenine
Adenine () ( symbol A or Ade) is a nucleobase (a purine derivative). It is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The three others are guanine, cytosine and thymine. Its deriv ...
(A), and thymine
Thymine () ( symbol T or Thy) is one of the four nucleobases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidin ...
(T) is more favorably compressed into the inner minor grooves. This means nucleosomes can bind preferentially at one position approximately every 10 base pairs (the helical repeat of DNA)- where the DNA is rotated to maximise the number of A and T bases that will lie in the inner minor groove. (See nucleic acid structure
Nucleic acid structure refers to the structure of nucleic acids such as DNA and RNA. Chemically speaking, DNA and RNA are very similar. Nucleic acid structure is often divided into four different levels: primary, secondary, tertiary, and quater ...
.)
30-nanometer chromatin fiber
 With addition of H1, the beads-on-a-string structure in turn coils into a 30 nm diameter helical structure known as the 30 nm fibre or filament. The precise structure of the chromatin fiber in the cell is not known in detail.
This level of chromatin structure is thought to be the form of
With addition of H1, the beads-on-a-string structure in turn coils into a 30 nm diameter helical structure known as the 30 nm fibre or filament. The precise structure of the chromatin fiber in the cell is not known in detail.
This level of chromatin structure is thought to be the form of heterochromatin
Heterochromatin is a tightly packed form of DNA or '' condensed DNA'', which comes in multiple varieties. These varieties lie on a continue between the two extremes of constitutive heterochromatin and facultative heterochromatin. Both play a rol ...
, which contains mostly transcriptionally silent genes. Electron microscopy studies have demonstrated that the 30 nm fiber is highly dynamic such that it unfolds into a 10 nm fiber beads-on-a-string structure when transversed by an RNA polymerase engaged in transcription.
 The existing models commonly accept that the nucleosomes lie perpendicular to the axis of the fibre, with linker histones arranged internally.
A stable 30 nm fibre relies on the regular positioning of nucleosomes along DNA. Linker DNA is relatively resistant to bending and rotation. This makes the length of linker DNA critical to the stability of the fibre, requiring nucleosomes to be separated by lengths that permit rotation and folding into the required orientation without excessive stress to the DNA.
In this view, different lengths of the linker DNA should produce different folding topologies of the chromatin fiber. Recent theoretical work, based on electron-microscopy images
of reconstituted fibers supports this view.
The existing models commonly accept that the nucleosomes lie perpendicular to the axis of the fibre, with linker histones arranged internally.
A stable 30 nm fibre relies on the regular positioning of nucleosomes along DNA. Linker DNA is relatively resistant to bending and rotation. This makes the length of linker DNA critical to the stability of the fibre, requiring nucleosomes to be separated by lengths that permit rotation and folding into the required orientation without excessive stress to the DNA.
In this view, different lengths of the linker DNA should produce different folding topologies of the chromatin fiber. Recent theoretical work, based on electron-microscopy images
of reconstituted fibers supports this view.
Spatial organization of chromatin in the cell nucleus
The spatial arrangement of the chromatin within the nucleus is not random - specific regions of the chromatin can be found in certain territories. Territories are, for example, the lamina-associated domains (LADs), and thetopologically associating domain
A topologically associating domain (TAD) is a self-interacting genomic region, meaning that DNA sequences within a TAD physically interact with each other more frequently than with sequences outside the TAD. The median size of a TAD in mouse cells ...
s (TADs), which are bound together by protein complexes. Currently, polymer models such as the Strings & Binders Switch (SBS) model and the Dynamic Loop (DL) model are used to describe the folding of chromatin within the nucleus. The arrangement of chromatin within the nucleus may also play a role in nuclear stress and restoring nuclear membrane deformation by mechanical stress. When chromatin is condensed, the nucleus becomes more rigid. When chromatin is decondensed, the nucleus becomes more elastic with less force
In physics, a force is an influence that can change the motion of an object. A force can cause an object with mass to change its velocity (e.g. moving from a state of rest), i.e., to accelerate. Force can also be described intuitively as a ...
exerted on the inner nuclear membrane. This observation sheds light on other possible cellular functions of chromatin organization outside of genomic regulation.
Cell-cycle dependent structural organization

interphase
Interphase is the portion of the cell cycle that is not accompanied by visible changes under the microscope, and includes the G1, S and G2 phases. During interphase, the cell grows (G1), replicates its DNA (S) and prepares for mitosis (G2). A c ...
of mitosis
In cell biology, mitosis () is a part of the cell cycle in which replicated chromosomes are separated into two new nuclei. Cell division by mitosis gives rise to genetically identical cells in which the total number of chromosomes is maintai ...
is optimized to allow simple access of transcription and DNA repair
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA d ...
factors to the DNA while compacting the DNA into the nucleus
Nucleus ( : nuclei) is a Latin word for the seed inside a fruit. It most often refers to:
* Atomic nucleus, the very dense central region of an atom
*Cell nucleus, a central organelle of a eukaryotic cell, containing most of the cell's DNA
Nucl ...
. The structure varies depending on the access required to the DNA. Genes
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a ba ...
that require regular access by RNA polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template.
Using the enzyme helicase, RNAP locally opens th ...
require the looser structure provided by euchromatin.
# Metaphase: The metaphase
Metaphase ( and ) is a stage of mitosis in the eukaryotic cell cycle in which chromosomes are at their second-most condensed and coiled stage (they are at their most condensed in anaphase). These chromosomes, carrying genetic information, a ...
structure of chromatin differs vastly to that of interphase
Interphase is the portion of the cell cycle that is not accompanied by visible changes under the microscope, and includes the G1, S and G2 phases. During interphase, the cell grows (G1), replicates its DNA (S) and prepares for mitosis (G2). A c ...
. It is optimised for physical strength and manageability, forming the classic chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins ar ...
structure seen in karyotype
A karyotype is the general appearance of the complete set of metaphase chromosomes in the cells of a species or in an individual organism, mainly including their sizes, numbers, and shapes. Karyotyping is the process by which a karyotype is disce ...
s. The structure of the condensed chromatin is thought to be loops of 30 nm fibre to a central scaffold of proteins. It is, however, not well-characterised. Chromosome scaffolds play an important role to hold the chromatin into compact chromosomes. Loops of 30 nm structure further condense with scaffold, into higher order structures. Chromosome scaffolds are made of proteins including condensin
Condensins are large protein complexes that play a central role in chromosome assembly and segregation during mitosis and meiosis (Figure 1). Their subunits were originally identified as major components of mitotic chromosomes assembled in ''Xenop ...
, type IIA topoisomerase and kinesin family member 4 (KIF4). The physical strength of chromatin is vital for this stage of division to prevent shear damage to the DNA as the daughter chromosomes are separated. To maximise strength the composition of the chromatin changes as it approaches the centromere, primarily through alternative histone H1 analogues. During mitosis, although most of the chromatin is tightly compacted, there are small regions that are not as tightly compacted. These regions often correspond to promoter regions of genes that were active in that cell type prior to chromatin formation. The lack of compaction of these regions is called bookmarking
Bookmarking (also "gene bookmarking" or "mitotic bookmarking") refers to a potential mechanism of transmission of gene expression programs through cell division.
During mitosis, gene transcription is silenced and most transcription factors are re ...
, which is an epigenetic
In biology, epigenetics is the study of stable phenotypic changes (known as ''marks'') that do not involve alterations in the DNA sequence. The Greek prefix '' epi-'' ( "over, outside of, around") in ''epigenetics'' implies features that are ...
mechanism believed to be important for transmitting to daughter cells the "memory" of which genes were active prior to entry into mitosis. This bookmarking
Bookmarking (also "gene bookmarking" or "mitotic bookmarking") refers to a potential mechanism of transmission of gene expression programs through cell division.
During mitosis, gene transcription is silenced and most transcription factors are re ...
mechanism is needed to help transmit this memory because transcription ceases during mitosis
In cell biology, mitosis () is a part of the cell cycle in which replicated chromosomes are separated into two new nuclei. Cell division by mitosis gives rise to genetically identical cells in which the total number of chromosomes is maintai ...
.
Chromatin and bursts of transcription
Chromatin and its interaction with enzymes has been researched, and a conclusion being made is that it is relevant and an important factor in gene expression. Vincent G. Allfrey, a professor at Rockefeller University, stated that RNA synthesis is related to histone acetylation. The lysine amino acid attached to the end of the histones is positively charged. The acetylation of these tails would make the chromatin ends neutral, allowing for DNA access. When the chromatin decondenses, the DNA is open to entry of molecular machinery. Fluctuations between open and closed chromatin may contribute to the discontinuity of transcription, ortranscriptional bursting Transcriptional bursting, also known as transcriptional pulsing, is a fundamental property of genes in which transcription from DNA to RNA can occur in "bursts" or "pulses", which has been observed in diverse organisms, from bacteria to mammals.
...
. Other factors are probably involved, such as the association and dissociation of transcription factor complexes with chromatin. Specifically, RNA polymerase and transcriptional proteins have been shown to congregate into droplets via phase separation, and recent studies have suggested that 10 nm chromatin demonstrates liquid-like behavior increasing the targetability of genomic DNA. The interactions between linker histones and disordered tail regions act as an electrostatic glue organizing large-scale chromatin into a dynamic, liquid-like domain. Decreased chromatin compaction comes with increased chromatin mobility and easier transcriptional access to DNA. The phenomenon, as opposed to simple probabilistic models of transcription, can account for the high variability in gene expression occurring between cells in isogenic populations.
Alternative chromatin organizations
During metazoanspermiogenesis
Spermiogenesis is the final stage of spermatogenesis, during which the spermatids develop into mature spermatozoa. At the beginning of the stage, the spermatid is a more or less circular cell containing a nucleus, Golgi apparatus, centriole an ...
, the spermatid's chromatin is remodeled into a more spaced-packaged, widened, almost crystal-like structure. This process is associated with the cessation of transcription and involves nuclear
Nuclear may refer to:
Physics
Relating to the nucleus of the atom:
*Nuclear engineering
*Nuclear physics
*Nuclear power
*Nuclear reactor
*Nuclear weapon
*Nuclear medicine
*Radiation therapy
*Nuclear warfare
Mathematics
*Nuclear space
*Nuclear ...
protein exchange. The histones are mostly displaced, and replaced by protamine
Protamines are small, arginine-rich, nuclear proteins that replace histones late in the haploid phase of spermatogenesis and are believed essential for sperm head condensation and DNA stabilization. They may allow for denser packaging of DNA in t ...
s (small, arginine
Arginine is the amino acid with the formula (H2N)(HN)CN(H)(CH2)3CH(NH2)CO2H. The molecule features a guanidino group appended to a standard amino acid framework. At physiological pH, the carboxylic acid is deprotonated (−CO2−) and both the am ...
-rich proteins). It is proposed that in yeast, regions devoid of histones become very fragile after transcription; HMO1, an HMG-box protein, helps in stabilizing nucleosomes-free chromatin.
Chromatin and DNA repair
A variety of internal and external agents can cause DNA damage in cells. Many factors influence how the repair route is selected, including the cell cycle phase and chromatin segment where the break occurred. In terms of initiating 5’ end DNA repair, the p53 binding protein 1(53BP1) and BRCA 1 are important protein components that influence Double-strand break repair pathway selection. The 53BP1 complex attaches to chromatin near DNA break and activities downstream factors such as Rap1-Interacting Factor 1 (RIF1) and the Shieldin, which protects DNA ends against nucleolytic destruction. DNA damage process occurs within the condition of chromatin, and the constantly changing chromatin environment has a large effect on it. Accessing and repairing the damaged cell of DNA, the genome condenses into chromatin and repairing it through modifying the histone residues. Through altering the chromatin structure, histones residues are adding chemical groups namely phosphate, acetyl and one or more than one methyl group and these are controlling the expressions of gene building by proteins to acquire DNA. Moreover, resynthesis of the delighted zone, DNA will be repaired by processing and restructuring the damaged bases. In order to maintain genomic integrity, “homologous recombination and classical non-homologous end joining process” has been followed by DNA to be repaired. The packaging of eukaryotic DNA into chromatin presents a barrier to all DNA-based processes that require recruitment of enzymes to their sites of action. To allow the critical cellular process of DNA repair, the chromatin must be remodeled. In eukaryotes,ATP-dependent chromatin remodeling
Chromatin remodeling is the dynamic modification of chromatin architecture to allow access of condensed genomic DNA to the regulatory Transcription (genetics), transcription machinery proteins, and thereby control gene expression. Such remodeling i ...
complexes and histone-modifying enzymes are two predominant factors employed to accomplish this remodeling process.
Chromatin relaxation occurs rapidly at the site of a DNA damage. This process is initiated by PARP1
Poly DP-ribosepolymerase 1 (PARP-1) also known as NAD+ ADP-ribosyltransferase 1 or poly DP-ribosesynthase 1 is an enzyme that in humans is encoded by the ''PARP1'' gene. It is the most abundant of the PARP family of enzymes, accounting for 90% o ...
protein that starts to appear at DNA damage in less than a second, with half maximum accumulation within 1.6 seconds after the damage occurs. Next the chromatin remodeler Alc1
Chromodomain-helicase-DNA-binding protein 1-like (ALC1) is an enzyme that in humans is encoded by the ''CHD1L'' gene. It has been implicated in chromatin remodeling and DNA relaxation process required for DNA replication, repair and transcriptio ...
quickly attaches to the product of PARP1, and completes arrival at the DNA damage within 10 seconds of the damage. About half of the maximum chromatin relaxation, presumably due to action of Alc1, occurs by 10 seconds. This then allows recruitment of the DNA repair enzyme MRE11, to initiate DNA repair, within 13 seconds.
γH2AX, the phosphorylated form of H2AX
H2A histone family member X (usually abbreviated as H2AX) is a type of histone protein from the H2A family encoded by the ''H2AFX'' gene. An important phosphorylated form is γH2AX (S139), which forms when double-strand breaks appear.
In humans ...
is also involved in the early steps leading to chromatin decondensation after DNA damage occurrence. The histone variant H2AX constitutes about 10% of the H2A histones in human chromatin. γH2AX (H2AX phosphorylated on serine 139) can be detected as soon as 20 seconds after irradiation of cells (with DNA double-strand break formation), and half maximum accumulation of γH2AX occurs in one minute. The extent of chromatin with phosphorylated γH2AX is about two million base pairs at the site of a DNA double-strand break. γH2AX does not, itself, cause chromatin decondensation, but within 30 seconds of irradiation, RNF8
E3 ubiquitin-protein ligase RNF8 is an enzyme that in humans is encoded by the ''RNF8'' gene. RNF8 has activity both in immune system functions and in DNA repair.
Function
The protein encoded by this gene contains a RING finger motif and an F ...
protein can be detected in association with γH2AX. RNF8 mediates extensive chromatin decondensation, through its subsequent interaction with CHD4, a component of the nucleosome remodeling and deacetylase complex NuRD
In the field of molecular biology, the Mi-2/NuRD (Nucleosome Remodeling Deacetylase) complex, is a group of associated proteins with both ATP-dependent chromatin remodeling and histone deacetylase activities. , Mi-2/NuRD was the only known protei ...
.
After undergoing relaxation subsequent to DNA damage, followed by DNA repair, chromatin recovers to a compaction state close to its pre-damage level after about 20 min.
Methods to investigate chromatin

 # ChIP-seq (Chromatin immunoprecipitation sequencing) is recognized as the vastly utilized chromatin identification method it has been using the antibodies that actively selected, identify and combine with proteins including "histones, histone restructuring, transaction factors and cofactors". This has been providing data about the state of chromatin and the transaction of a gene by trimming "oligonucleotides" that are unbound. Chromatin immunoprecipitation sequencing aimed against different histone modifications, can be used to identify chromatin states throughout the genome. Different modifications have been linked to various states of chromatin.
#
# ChIP-seq (Chromatin immunoprecipitation sequencing) is recognized as the vastly utilized chromatin identification method it has been using the antibodies that actively selected, identify and combine with proteins including "histones, histone restructuring, transaction factors and cofactors". This has been providing data about the state of chromatin and the transaction of a gene by trimming "oligonucleotides" that are unbound. Chromatin immunoprecipitation sequencing aimed against different histone modifications, can be used to identify chromatin states throughout the genome. Different modifications have been linked to various states of chromatin.
# DNase-seq DNase-seq ( DNase I hypersensitive sites sequencing) is a method in molecular biology used to identify the location of regulatory regions, based on the genome-wide sequencing of regions sensitive to cleavage by DNase I. FAIRE-Seq is a successor of ...
(DNase I hypersensitive sites Sequencing) uses the sensitivity of accessible regions in the genome to the DNase I enzyme to map open or accessible regions in the genome.
# FAIRE-seq (Formaldehyde-Assisted Isolation of Regulatory Elements sequencing) uses the chemical properties of protein-bound DNA in a two-phase separation method to extract nucleosome depleted regions from the genome.
# ATAC-seq (Assay for Transposable Accessible Chromatin sequencing) uses the Tn5 transposase to integrate (synthetic) transposons into accessible regions of the genome consequentially highlighting the localisation of nucleosomes and transcription factors across the genome.
# DNA footprinting is a method aimed at identifying protein-bound DNA. It uses labeling and fragmentation coupled to gel electrophoresis to identify areas of the genome that have been bound by proteins.
# MNase-seq (Micrococcal Nuclease sequencing) uses the micrococcal nuclease enzyme to identify nucleosome positioning throughout the genome.
# Chromosome conformation capture
Chromosome conformation capture techniques (often abbreviated to 3C technologies or 3C-based methods) are a set of molecular biology methods used to analyze the spatial organization of chromatin in a cell. These methods quantify the number of int ...
determines the spatial organization of chromatin in the nucleus, by inferring genomic locations that physically interact.
# MACC profiling (Micrococcal nuclease ACCessibility profiling) uses titration series of chromatin digests with micrococcal nuclease to identify chromatin accessibility as well as to map nucleosomes and non-histone DNA-binding proteins in both open and closed regions of the genome.
Chromatin and knots
It has been a puzzle how decondensed interphase chromosomes remain essentially unknotted. The natural expectation is that in the presence of type II DNA topoisomerases that permit passages of double-stranded DNA regions through each other, all chromosomes should reach the state of topological equilibrium. The topological equilibrium in highly crowded interphase chromosomes forming chromosome territories would result in formation of highly knotted chromatin fibres. However, Chromosome Conformation Capture (3C) methods revealed that the decay of contacts with the genomic distance in interphase chromosomes is practically the same as in the crumpled globule state that is formed when long polymers condense without formation of any knots. To remove knots from highly crowded chromatin, one would need an active process that should not only provide the energy to move the system from the state of topological equilibrium but also guide topoisomerase-mediated passages in such a way that knots would be efficiently unknotted instead of making the knots even more complex. It has been shown that the process of chromatin-loop extrusion is ideally suited to actively unknot chromatin fibres in interphase chromosomes.Chromatin: alternative definitions
The term, introduced byWalther Flemming
Walther Flemming (21 April 1843 – 4 August 1905) was a German biologist and a founder of cytogenetics.
He was born in Sachsenberg (now part of Schwerin) as the fifth child and only son of the psychiatrist Carl Friedrich Flemming (1799–18 ...
, has multiple meanings:
# Simple and concise definition: Chromatin is a macromolecular complex of a DNA macromolecule and protein macromolecules (and RNA). The proteins package and arrange the DNA and control its functions within the cell nucleus.
# A biochemists' operational definition: Chromatin is the DNA/protein/RNA complex extracted from eukaryotic lysed interphase nuclei. Just which of the multitudinous substances present in a nucleus will constitute a part of the extracted material partly depends on the technique each researcher uses. Furthermore, the composition and properties of chromatin vary from one cell type to another, during the development of a specific cell type, and at different stages in the cell cycle.
# The ''DNA + histone = chromatin'' definition: The DNA double helix in the cell nucleus is packaged by special proteins termed histones. The formed protein/DNA complex is called chromatin. The basic structural unit of chromatin is the nucleosome.
The first definition allows for "chromatins" to be defined in other domains of life like bacteria and archaea, using any DNA-binding proteins that condenses the molecule. These proteins are usually referred to nucleoid-associated proteins (NAPs); examples include AsnC/LrpC with HU. In addition, some archaea do produce nucleosomes from proteins homologous to eukaryotic histones.
Nobel Prizes
The following scientists were recognized for their contributions to chromatin research withNobel Prize
The Nobel Prizes ( ; sv, Nobelpriset ; no, Nobelprisen ) are five separate prizes that, according to Alfred Nobel's will of 1895, are awarded to "those who, during the preceding year, have conferred the greatest benefit to humankind." Alfr ...
s:
See also
* Active chromatin sequence *Chromatid
A chromatid (Greek ''khrōmat-'' 'color' + ''-id'') is one half of a duplicated chromosome. Before replication, one chromosome is composed of one DNA molecule. In replication, the DNA molecule is copied, and the two molecules are known as chr ...
* DAnCER
Dance is a performing art form consisting of sequences of movement, either improvised or purposefully selected. This movement has aesthetic and often symbolic value. Dance can be categorized and described by its choreography, by its repertoire ...
database (2010)
* Epigenetics
In biology, epigenetics is the study of stable phenotypic changes (known as ''marks'') that do not involve alterations in the DNA sequence. The Greek prefix '' epi-'' ( "over, outside of, around") in ''epigenetics'' implies features that are ...
* Histone-modifying enzymes
* Position-effect variegation Position-effect variegation (PEV) is a variegation caused by the silencing of a gene in some cells through its abnormal juxtaposition with heterochromatin via rearrangement or transposition. It is also associated with changes in chromatin conforma ...
* Transcriptional bursting Transcriptional bursting, also known as transcriptional pulsing, is a fundamental property of genes in which transcription from DNA to RNA can occur in "bursts" or "pulses", which has been observed in diverse organisms, from bacteria to mammals.
...
Notes
References
Additional sources
* Cooper, Geoffrey M. 2000. The Cell, 2nd edition, A Molecular Approach. Chapter 4.Chromosomes and Chromatin.
* * Cremer, T. 1985. Von der Zellenlehre zur Chromosomentheorie: Naturwissenschaftliche Erkenntnis und Theorienwechsel in der frühen Zell- und Vererbungsforschung, Veröffentlichungen aus der Forschungsstelle für Theoretische Pathologie der Heidelberger Akademie der Wissenschaften. Springer-Vlg., Berlin, Heidelberg. * Elgin, S. C. R. (ed.). 1995. Chromatin Structure and Gene Expression, vol. 9. IRL Press, Oxford, New York, Tokyo. * * * * * * Pollard, T., and W. Earnshaw. 2002. Cell Biology. Saunders. * Saumweber, H. 1987. Arrangement of Chromosomes in Interphase Cell Nuclei, p. 223-234. In W. Hennig (ed.), Structure and Function of Eucaryotic Chromosomes, vol. 14. Springer-Verlag, Berlin, Heidelberg. * * Van Holde KE. 1989. Chromatin. New York:
Springer-Verlag
Springer Science+Business Media, commonly known as Springer, is a German multinational publishing company of books, e-books and peer-reviewed journals in science, humanities, technical and medical (STM) publishing.
Originally founded in 1842 ...
. .
* Van Holde, K., J. Zlatanova, G. Arents, and E. Moudrianakis. 1995. Elements of chromatin structure: histones, nucleosomes, and fibres, p. 1-26. In S. C. R. Elgin (ed.), Chromatin structure and gene expression. IRL Press at Oxford University Press, Oxford.
External links
Chromatin, Histones & Cathepsin
PMAP
The Proteolysis Map
The Proteolysis MAP (PMAP) is an integrated web resource focused on proteases.
Rationale
PMAP is to aid the protease researchers in reasoning about proteolytic networks and metabolic pathways.
History and funding
PMAP was originally created ...
-animation
''Nature'' journal: recent chromatin publications and news
Protocol for ''in vitro'' Chromatin Assembly
ENCODE threads Explorer
Chromatin patterns at transcription factor binding sites.
Nature (journal)
''Nature'' is a British weekly scientific journal founded and based in London, England. As a multidisciplinary publication, ''Nature'' features peer-reviewed research from a variety of academic disciplines, mainly in science and technology. ...
{{portal bar, Biology, Science
Molecular genetics
Nuclear substructures