|
Q-FISH
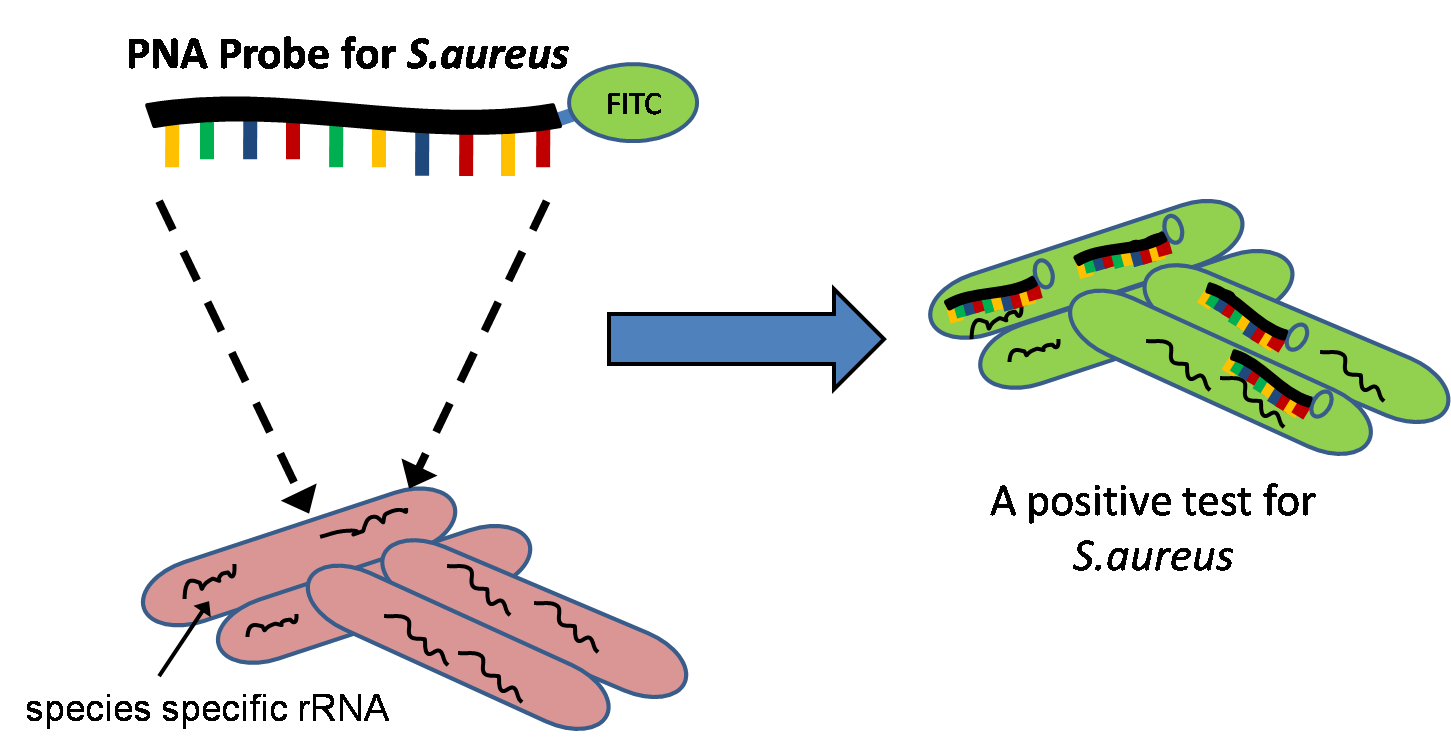
Quantitative Fluorescent ''in situ'' hybridization (Q-FISH) is a cytogenetic technique based on the traditional FISH methodology. In Q-FISH, the technique uses labelled ( Cy3 or FITC) synthetic DNA mimics called peptide nucleic acid (PNA) oligonucleotides to quantify target sequences in chromosomal DNA using fluorescent microscopy and analysis software. Q-FISH is most commonly used to study telomere length, which in vertebrates are repetitive hexameric sequences (TTAGGG) located at the distal end of chromosomes. Telomeres are necessary at chromosome ends to prevent DNA-damage responses as well as genome instability. To this day, the Q-FISH method continues to be utilized in the field of telomere research. PNAs and FISH Due to the fact that PNA backbones contain no charged phosphate groups, binding between PNA and DNA is stronger than that of DNA/DNA or DNA/RNA duplexes. Q-FISH utilizes this unique characteristic of PNAs where at low ionic strengths, PNAs can anneal to complementa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Q-FISH Image From A Human Fibrosarcoma Cell
Quantitative Fluorescent ''in situ'' hybridization (Q-FISH) is a cytogenetic technique based on the traditional FISH methodology. In Q-FISH, the technique uses labelled ( Cy3 or FITC) synthetic DNA mimics called peptide nucleic acid (PNA) oligonucleotides to quantify target sequences in chromosomal DNA using fluorescent microscopy and analysis software. Q-FISH is most commonly used to study telomere length, which in vertebrates are repetitive hexameric sequences (TTAGGG) located at the distal end of chromosomes. Telomeres are necessary at chromosome ends to prevent DNA-damage responses as well as genome instability. To this day, the Q-FISH method continues to be utilized in the field of telomere research. PNAs and FISH Due to the fact that PNA backbones contain no charged phosphate groups, binding between PNA and DNA is stronger than that of DNA/DNA or DNA/RNA duplexes. Q-FISH utilizes this unique characteristic of PNAs where at low ionic strengths, PNAs can anneal to complementa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
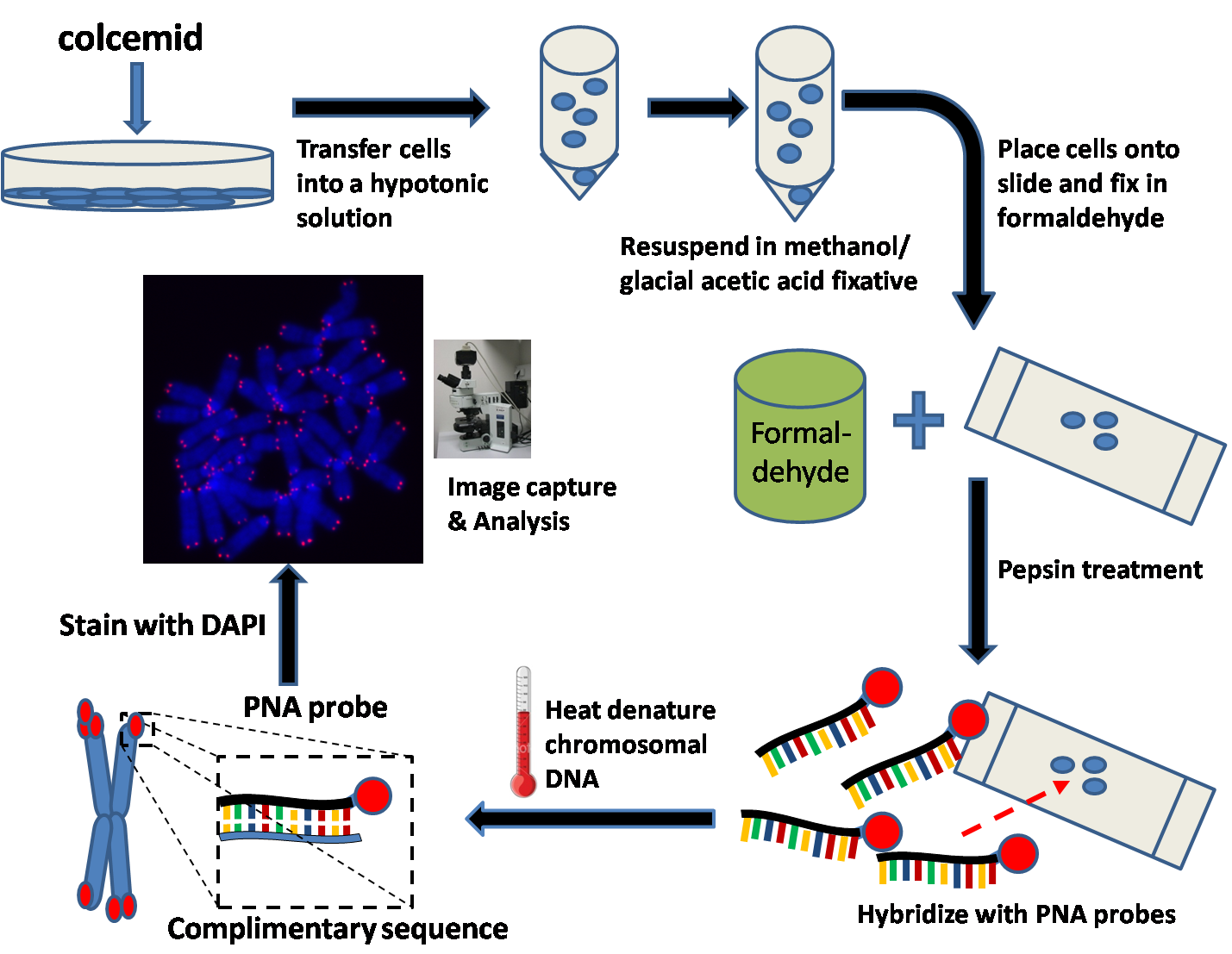
Q-FISH Workflow
Quantitative Fluorescent ''in situ'' hybridization (Q-FISH) is a cytogenetic technique based on the traditional FISH methodology. In Q-FISH, the technique uses labelled ( Cy3 or FITC) synthetic DNA mimics called peptide nucleic acid (PNA) oligonucleotides to quantify target sequences in chromosomal DNA using fluorescent microscopy and analysis software. Q-FISH is most commonly used to study telomere length, which in vertebrates are repetitive hexameric sequences (TTAGGG) located at the distal end of chromosomes. Telomeres are necessary at chromosome ends to prevent DNA-damage responses as well as genome instability. To this day, the Q-FISH method continues to be utilized in the field of telomere research. PNAs and FISH Due to the fact that PNA backbones contain no charged phosphate groups, binding between PNA and DNA is stronger than that of DNA/DNA or DNA/RNA duplexes. Q-FISH utilizes this unique characteristic of PNAs where at low ionic strengths, PNAs can anneal to complementa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Flow-FISH
Flow-FISH (fluorescent in-situ hybridization) is a cytogenetic technique to quantify the copy number of RNA or specific repetitive elements in genomic DNA of whole cell populations via the combination of flow cytometry with cytogenetic fluorescent in situ hybridization staining protocols.Baerlocher GM, Vulto I, de Jong G, Lansdorp PM. Flow cytometry and FISH to measure the average length of telomeres (flow FISH). Nat Protoc 2006; 1:2365–2376. Flow-FISH is most commonly used to quantify the length of telomeres, which are stretches of repetitious DNA (hexameric TTAGGG repeats) at the distal ends of chromosomes in human white blood cells, and a semi-automated method for doing so was published in Nature Protocols. Telomere length in white blood cells has been a subject of interest because telomere length in these cell types (and also of other somatic tissues) declines gradually over the human lifespan, resulting in cell senescence, apoptosis, or transformation. This decline ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Fluorescent In Situ Hybridization
Fluorescence ''in situ'' hybridization (FISH) is a molecular cytogenetic technique that uses fluorescent probes that bind to only particular parts of a nucleic acid sequence with a high degree of sequence complementarity. It was developed by biomedical researchers in the early 1980s to detect and localize the presence or absence of specific DNA sequences on chromosomes. Fluorescence microscopy can be used to find out where the fluorescent probe is bound to the chromosomes. FISH is often used for finding specific features in DNA for use in genetic counseling, medicine, and species identification. FISH can also be used to detect and localize specific RNA targets (mRNA, lncRNA and miRNA) in cells, circulating tumor cells, and tissue samples. In this context, it can help define the spatial-temporal patterns of gene expression within cells and tissues. Probes – RNA and DNA In biology, a probe is a single strand of DNA or RNA that is complementary to a nucleotide sequence ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cyanine
Cyanines, also referred to as tetramethylindo(di)-carbocyanines are a synthetic dye family belonging to the polymethine group. Although the name derives etymologically from terms for shades of blue, the cyanine family covers the electromagnetic spectrum from near IR to UV. Chemically, cyanines are a conjugated system between two nitrogen atoms; in each resonance structure, exactly one nitrogen atom is oxidized to an iminium. Typically, they form part of a nitrogenous heterocyclic system. The main application for cyanine dyes is in biological labeling. Nevertheless, there is a wide literature on both their synthesis and uses, and cyanines are common in some CD and DVD media. Structure Cyanines have been classified in many ways: * ''Streptocyanines'' or ''open chain cyanines'': : R2N+=CH H=CH'n''-NR2 (I) * ''Hemicyanines'': : Aryl=N+=CH H=CH'n''-NR2 (II) * ''Closed chain cyanines'': :Aryl=N+=CH H=CH'n''-N=Aryl (III) Additionally, these classes are recognized: * ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tonicity
In chemical biology, tonicity is a measure of the effective osmotic pressure gradient; the water potential of two solutions separated by a partially-permeable cell membrane. Tonicity depends on the relative concentration of selective membrane-impermeable solutes across a cell membrane which determine the direction and extent of osmotic flux. It is commonly used when describing the swelling-versus-shrinking response of cells immersed in an external solution. Unlike osmotic pressure, tonicity is influenced only by solutes that cannot cross the membrane, as only these exert an effective osmotic pressure. Solutes able to freely cross the membrane do not affect tonicity because they will always equilibrate with equal concentrations on both sides of the membrane without net solvent movement. It is also a factor affecting imbibition. There are three classifications of tonicity that one solution can have relative to another: ''hypertonic'', ''hypotonic'', and ''isotonic''.A hypotonic s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Plasmids
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; however, plasmids are sometimes present in archaea and eukaryotic organisms. In nature, plasmids often carry genes that benefit the survival of the organism and confer selective advantage such as antibiotic resistance. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain only additional genes that may be useful in certain situations or conditions. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms. In the laboratory, plasmids may be introduced into a cell via transformation. Synthetic plasmids are available for procurement over the intern ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Locus (genetics)
In genetics, a locus (plural loci) is a specific, fixed position on a chromosome where a particular gene or genetic marker is located. Each chromosome carries many genes, with each gene occupying a different position or locus; in humans, the total number of protein-coding genes in a complete haploid set of 23 chromosomes is estimated at 19,000–20,000. Genes may possess multiple variants known as alleles, and an allele may also be said to reside at a particular locus. Diploid and polyploid cells whose chromosomes have the same allele at a given locus are called homozygous with respect to that locus, while those that have different alleles at a given locus are called heterozygous. The ordered list of loci known for a particular genome is called a gene map. Gene mapping is the process of determining the specific locus or loci responsible for producing a particular phenotype or biological trait. Association mapping, also known as "linkage disequilibrium mapping", is a method of ma ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Karyotype
A karyotype is the general appearance of the complete set of metaphase chromosomes in the cells of a species or in an individual organism, mainly including their sizes, numbers, and shapes. Karyotyping is the process by which a karyotype is discerned by determining the chromosome complement of an individual, including the number of chromosomes and any abnormalities. A karyogram or idiogram is a graphical depiction of a karyotype, wherein chromosomes are organized in pairs, ordered by size and position of centromere for chromosomes of the same size. Karyotyping generally combines light microscopy and photography, and results in a photomicrographic (or simply micrographic) karyogram. In contrast, a schematic karyogram is a designed graphic representation of a karyotype. In schematic karyograms, just one of the sister chromatids of each chromosome is generally shown for brevity, and in reality they are generally so close together that they look as one on photomicrographs as well ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Photobleaching
In optics, photobleaching (sometimes termed fading) is the photochemical alteration of a dye or a fluorophore molecule such that it is permanently unable to fluoresce. This is caused by cleaving of covalent bonds or non-specific reactions between the fluorophore and surrounding molecules. Such irreversible modifications in covalent bonds are caused by transition from a singlet state to the triplet state of the fluorophores. The number of excitation cycles to achieve full bleaching varies. In microscopy, photobleaching may complicate the observation of fluorescent molecules, since they will eventually be destroyed by the light exposure necessary to stimulate them into fluorescing. This is especially problematic in time-lapse microscopy. However, photobleaching may also be used prior to applying the (primarily antibody-linked) fluorescent molecules, in an attempt to quench autofluorescence. This can help improve the signal-to-noise ratio. Photobleaching may also be exploited to study ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DAPI
DAPI (pronounced 'DAPPY', /ˈdæpiː/), or 4′,6-diamidino-2-phenylindole, is a fluorescent stain that binds strongly to adenine–thymine-rich regions in DNA. It is used extensively in fluorescence microscopy. As DAPI can pass through an intact cell membrane, it can be used to stain both live and fixed cells, though it passes through the membrane less efficiently in live cells and therefore provides a marker for membrane viability. History DAPI was first synthesised in 1971 in the laboratory of Otto Dann as part of a search for drugs to treat trypanosomiasis. Although it was unsuccessful as a drug, further investigation indicated it bound strongly to DNA and became more fluorescent when bound. This led to its use in identifying mitochondrial DNA in ultracentrifugation in 1975, the first recorded use of DAPI as a fluorescent DNA stain. Strong fluorescence when bound to DNA led to the rapid adoption of DAPI for fluorescent staining of DNA for fluorescence microscopy. Its use f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Denaturation (biochemistry)
In biochemistry, denaturation is a process in which proteins or nucleic acids lose the quaternary structure, tertiary structure, and secondary structure which is present in their native state, by application of some external stress or compound such as a strong acid or base, a concentrated inorganic salt, an organic solvent (e.g., alcohol or chloroform), agitation and radiation or heat. If proteins in a living cell are denatured, this results in disruption of cell activity and possibly cell death. Protein denaturation is also a consequence of cell death. Denatured proteins can exhibit a wide range of characteristics, from conformational change and loss of solubility to aggregation due to the exposure of hydrophobic groups. The loss of solubility as a result of denaturation is called ''coagulation''. Denatured proteins lose their 3D structure and therefore cannot function. Protein folding is key to whether a globular or membrane protein can do its job correctly; it must be ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |