Q-FISH Workflow on:
[Wikipedia]
[Google]
[Amazon]
Quantitative Fluorescent ''in situ'' hybridization (Q-FISH) is a
Another application of Q-FISH is the detection of telomeric fusions, where the ends of chromosomes have been fused together at the telomere, which are sometimes called interstitial (within the chromosome) telomeric sequences (ITSs). Studying telomeric fusions can sometimes show the course of evolution. For example, one human chromosome has an ITS that is hypothesized to be the equivalent of two chromosomes in chimpanzees that fused together. Observing the regulation of telomere length in different species also reveals important information about karyotype evolution and relevance to human illnesses. In another example, the
cytogenetic
Cytogenetics is essentially a branch of genetics, but is also a part of cell biology/cytology (a subdivision of human anatomy), that is concerned with how the chromosomes relate to cell behaviour, particularly to their behaviour during mitosis an ...
technique based on the traditional FISH
Fish are aquatic, craniate, gill-bearing animals that lack limbs with digits. Included in this definition are the living hagfish, lampreys, and cartilaginous and bony fish as well as various extinct related groups. Approximately 95% of li ...
methodology. In Q-FISH, the technique uses labelled ( Cy3 or FITC) synthetic DNA mimics called peptide nucleic acid
Peptide nucleic acid (PNA) is an artificially synthesized polymer similar to DNA or RNA.
Synthetic peptide nucleic acid oligomers have been used in recent years in molecular biology procedures, diagnostic assays, and antisense therapies. Due to ...
(PNA) oligonucleotides
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids c ...
to quantify target sequences in chromosomal
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
DNA using fluorescent microscopy
A fluorescence microscope is an optical microscope that uses fluorescence instead of, or in addition to, scattering, reflection, and attenuation or absorption, to study the properties of organic or inorganic substances. "Fluorescence microsc ...
and analysis software. Q-FISH is most commonly used to study telomere
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ...
length, which in vertebrates
Vertebrates () comprise all animal taxa within the subphylum Vertebrata () ( chordates with backbones), including all mammals, birds, reptiles, amphibians, and fish. Vertebrates represent the overwhelming majority of the phylum Chordata, ...
are repetitive hexameric sequences (TTAGGG) located at the distal end of chromosomes. Telomeres are necessary at chromosome ends to prevent DNA-damage responses as well as genome instability
Genome instability (also genetic instability or genomic instability) refers to a high frequency of mutations within the genome of a cellular lineage. These mutations can include changes in nucleic acid sequences, chromosomal rearrangements or aneup ...
. To this day, the Q-FISH method continues to be utilized in the field of telomere research.
PNAs and FISH
Due to the fact that PNA backbones contain no charged phosphate groups, binding between PNA and DNA is stronger than that of DNA/DNA or DNA/RNA duplexes. Q-FISH utilizes this unique characteristic of PNAs where at low ionic strengths, PNAs can anneal to complementary single-stranded DNA sequences while single-stranded DNA cannot. By using conditions that only allow labeled (CCCTAA)3 PNA to hybridize to (TTAGGG)n target sequences, Q-FISH is able to quantify the hybridization of PNAs to telomeric sequences.General method/protocol for cultured cells

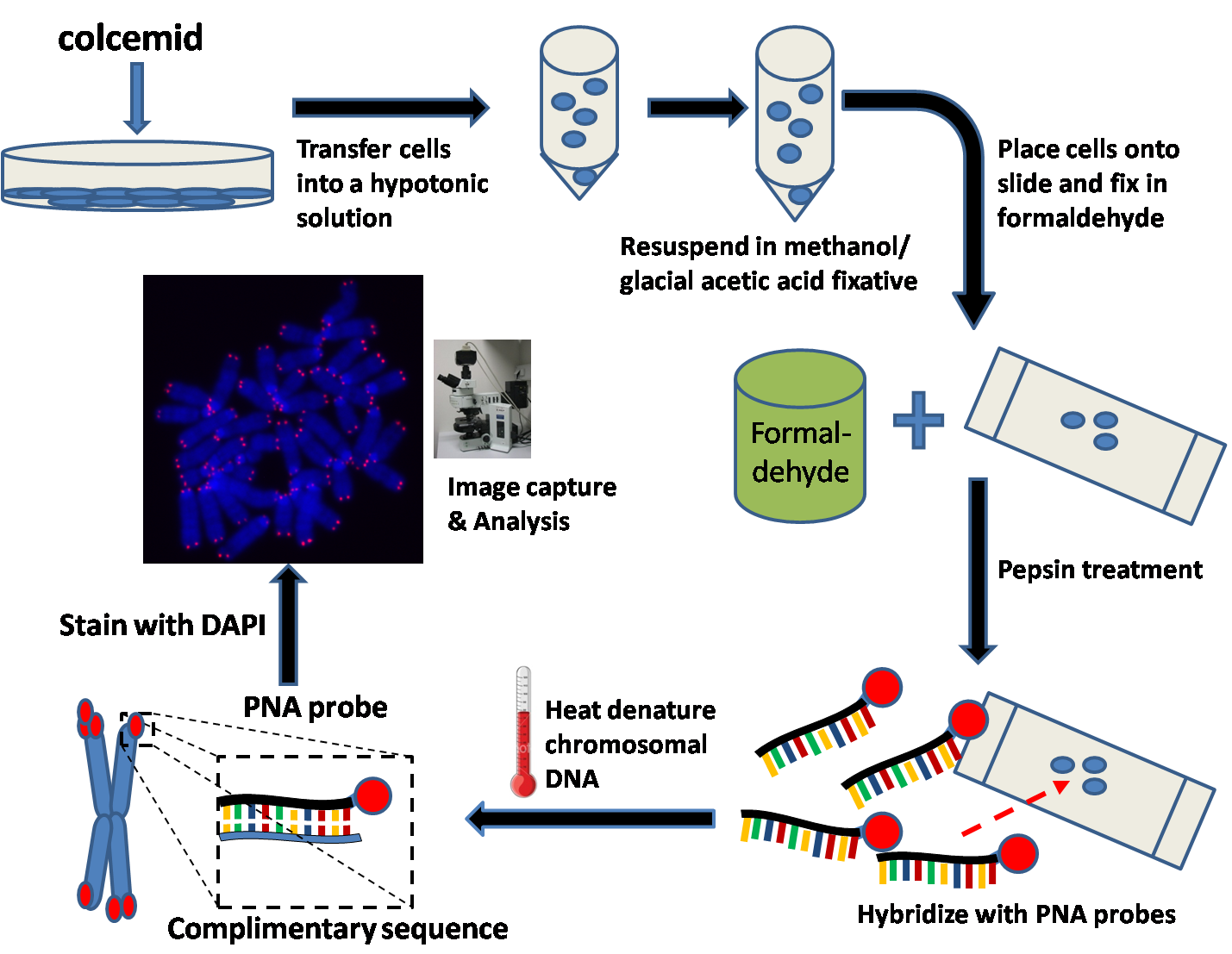
Prepare metaphase-arrested cells
Several hours prior to harvesting thecultured
Culture () is an umbrella term which encompasses the social behavior, institutions, and norms found in human societies, as well as the knowledge, beliefs, arts, laws, customs, capabilities, and habits of the individuals in these groups.Tylor ...
cells, colcemid is added to the culture medium. Colcemid acts to arrest cells in the metaphase
Metaphase ( and ) is a stage of mitosis in the eukaryotic cell cycle in which chromosomes are at their second-most condensed and coiled stage (they are at their most condensed in anaphase). These chromosomes, carrying genetic information, align ...
state by disrupting microtubules
Microtubules are polymers of tubulin that form part of the cytoskeleton and provide structure and shape to eukaryotic cells. Microtubules can be as long as 50 micrometres, as wide as 23 to 27 nm and have an inner diameter between 11 an ...
in mitotic
In cell biology, mitosis () is a part of the cell cycle in which replicated chromosomes are separated into two new nuclei. Cell division by mitosis gives rise to genetically identical cells in which the total number of chromosomes is maintai ...
cells. Cells are then trypsin
Trypsin is an enzyme in the first section of the small intestine that starts the digestion of protein molecules by cutting these long chains of amino acids into smaller pieces. It is a serine protease from the PA clan superfamily, found in the dig ...
ized and resuspended in a hypotonic
In chemical biology, tonicity is a measure of the effective osmotic pressure gradient; the water potential of two solutions separated by a partially-permeable cell membrane. Tonicity depends on the relative concentration of selective membrane-imp ...
buffer. This will swell the collected cells and spread the chromosomes.
Fix cells
The hypotonic solution is then removed by centrifugation and resuspended in a methanol/glacial acetic acid fixative.Prepare slides
Place a few drops of the cell suspension onto a microscope slide and let air dry overnight. The following day, immerse the slide inphosphate buffered saline
Phosphate-buffered saline (PBS) is a buffer solution (pH ~ 7.4) commonly used in biological research. It is a water-based salt solution containing disodium hydrogen phosphate, sodium chloride and, in some formulations, potassium chloride and potass ...
(PBS) for several minutes.
Fix slides in formaldehyde
Transfer the slides into a 4%formaldehyde
Formaldehyde ( , ) (systematic name methanal) is a naturally occurring organic compound with the formula and structure . The pure compound is a pungent, colourless gas that polymerises spontaneously into paraformaldehyde (refer to section F ...
solution and fix for several minutes. Wash slides several times with PBS.
Treat slides with pepsin
Slides are then transferred into apepsin
Pepsin is an endopeptidase that breaks down proteins into smaller peptides. It is produced in the gastric chief cells of the stomach lining and is one of the main digestive enzymes in the digestive systems of humans and many other animals, w ...
solution. Pepsin is a protease
A protease (also called a peptidase, proteinase, or proteolytic enzyme) is an enzyme that catalyzes (increases reaction rate or "speeds up") proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the ...
and acts to digest proteins into peptides.
Hybridize PNA probe (Cy3 or FITC labelled PNAs)
A small volume of the hybridization mixture is placed onto a coverslip and then placed gently onto the microscope slide which contains the fixed cells.Heat denature DNA
The slide is then placed into a preheated oven where the chromosomal DNA in the cell is denatured at 80 °C for several minutes. The slide is then left at room temperature for several hours to allow the PNA to hybridize to complementary DNA.Wash slides to remove unbound PNAs and counterstain DNA (DAPI or PI)
Slides are then carefully washed in various wash solutions to remove unbound PNA. Microscope mounting medium is then placed onto the cells. This medium generally containsDAPI
DAPI (pronounced 'DAPPY', /ˈdæpiː/), or 4′,6-diamidino-2-phenylindole, is a fluorescent stain that binds strongly to adenine–thymine-rich regions in DNA. It is used extensively in fluorescence microscopy. As DAPI can pass through an inta ...
(a DNA counterstain) and an antifade solution to preserve the PNA fluorescence and reduce photobleaching
In optics, photobleaching (sometimes termed fading) is the photochemical alteration of a dye or a fluorophore molecule such that it is permanently unable to fluoresce. This is caused by cleaving of covalent bonds or non-specific reactions between t ...
.
Image capture and analysis
Before experimental samples are imaged, fluorescent reference beads are imaged in order to ensure the proper set-up of the camera and fluorescent microscope. In addition, these reference beads will be imaged prior to every acquisition session. This will ensure that the differences between samples are not due to errors in the lamp or camera.Slijepcevic, Predrag. "Telomere length measurement by Q-FISH." Methods in Cell Science (2001) 23:17-22 A metaphase cell is then manually selected and centered for the camera. Two types of images are taken: pictures of the stained chromosomes in their metaphase state and fluorescent images of the telomeres. The two images can then be superimposed to generate a combined image. This image can then bekaryotype
A karyotype is the general appearance of the complete set of metaphase chromosomes in the cells of a species or in an individual organism, mainly including their sizes, numbers, and shapes. Karyotyping is the process by which a karyotype is disce ...
d or assigned nomenclature. Furthermore, the intra-chromosomal distribution of telomere length in p-arms versus q-arms can be measured.
Data from different experiments may be used to normalize the fluorescence intensity while plasmids
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; how ...
with a known number of telomeric repeats can be used as standards to help relate telomere fluorescence and telomere length. In addition to the fluorescent reference beads, signal strength from sister chromatids should be equal and therefore can be used as another control to gauge the precision of the data. Lastly, it is important the images are not saturated. If the fluorescence intensity reaches saturation, telomere lengths become underestimated. Q-FISH image analysis software is available for free from the Flintbox Network aApplications and significance
Q-FISH has been used extensively to quantitate information regarding telomere length distribution and associating it with various illnesses. In this context, Q-FISH is particularly relevant because it is able to detect and quantify critically short telomeres. It has been shown that it is the frequency of these critically short telomeres, rather than the average telomere length, that is important in telomere dysfunction.Hemann MT., Strong, MA., Hao, LY., Greider, CW. "The shortest telomere, not average telomere length, is critical for cell viability and chromosome stability." ''Cell'' (2001) 107:67-77.Samper, E., Flores, JM., Blasco, MA. "Restoration of telomerase activity rescues chromosomal instability and premature aging in Terc-/- mice with short telomeres." ''EMBO Rep'' (2001) 2:800-807. While Q-FISH supplies accurate information about telomere length, its relevance can be extended by combining Q-FISH with other FISH related techniques, such asflow-FISH
Flow-FISH (fluorescent in-situ hybridization) is a cytogenetic technique to quantify the copy number of RNA or specific repetitive elements in genomic DNA of whole cell populations via the combination of flow cytometry with cytogenetic fluoresc ...
. In flow-FISH, flow cytometry
Flow cytometry (FC) is a technique used to detect and measure physical and chemical characteristics of a population of cells or particles.
In this process, a sample containing cells or particles is suspended in a fluid and injected into the flo ...
is utilized to measure fluorescence intensity (and thus telomere length) in a large population of cells rather than just a handful of cells in Q-FISH. Conversely, unlike Q-FISH, flow-FISH is unable to determine telomere length in a particular chromosome within an individual cell.Baerlocher, GM., Vultro, I., de Jong, G., Lansdorp, PM. "Flow cytometry and FISH to measure the average length of telomeres." ''Nature Protocols'' (2006) 1(5):2365-2376. However, although Q-FISH is generally considered low-throughput and not suitable for population studies, groups have developed high-throughput (HT) Q-FISH protocols that use automated machinery to perform Q-FISH on interphase nuclei in 96well plates.Canela, A., Vera, E., Klatt, P., Blasco, MA. "High-throughput telomere length quantification by FISH and its application to human population studies." ''PNAS'' (2007) 104(13):5300-5305.
Similarly, other methods like multiplex-FISH and cenM-FISH have been developed which can also be used in conjunction with Q-FISH. Multiplex-FISH uses a variety of probes to visualize the 24 chromosomes in different colours and identify intra- or inter-chromosomal rearrangements.Uhrig, S., Schuffenhauer, S., Fauth, C., Wirtz, A., Daumer-Haas, C., Apacik, C., Cohen, M., Müller-Navia, J., Cremer, T., Murken, J., and Speicher, MR. "Multiplex-FISH for Pre- and Post-Natal Diagnostic Applications." American Journal of Human Genetics (1999) 65:448-462. Centromere-specific multi-colour FISH (cenM-FISH) uses the multi-coloured probes from multiplex-FISH as well as centromere
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers a ...
specific labeled probes to identify and distinguish centromere regions. The relation between centromere abnormalities or chromosomal rearrangements and telomere length may have high clinical impact, since all appear important in pre- or post-natal diagnostics and tumor developments.Nietzel, A., Rocchi, M., Starke, H., Heller, A., Fielder, W., Wlodarska, I., Loncarevic, IF., Beensen, V., Claussen, U., and Liehr, T. "A new multi-colour FISH approach for the characterization of marker chromosomes: centromere-specific multicolor-FISH (cenM-FISH)." Human Genetics (2001) 108:199-204. These experiments can provide more enlightenment about the role of telomeres and the importance of telomere length.
Another application of Q-FISH is the detection of telomeric fusions, where the ends of chromosomes have been fused together at the telomere, which are sometimes called interstitial (within the chromosome) telomeric sequences (ITSs). Studying telomeric fusions can sometimes show the course of evolution. For example, one human chromosome has an ITS that is hypothesized to be the equivalent of two chromosomes in chimpanzees that fused together. Observing the regulation of telomere length in different species also reveals important information about karyotype evolution and relevance to human illnesses. In another example, the
non-homologous end joining
Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. NHEJ is referred to as "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology direct ...
(NHEJ) protein repairs double-stranded DNA breaks and relies on the Ku70
Ku70 is a protein that, in humans, is encoded by the ''XRCC6'' gene.
Function
Together, Ku70 and Ku80 make up the Ku heterodimer, which binds to DNA double-strand break ends and is required for the non-homologous end joining (NHEJ) pathway of ...
/Ku80
Ku80 is a protein that, in humans, is encoded by the ''XRCC5'' gene. Together, Ku70 and Ku80 make up the Ku heterodimer, which binds to DNA double-strand break ends and is required for the non-homologous end joining (NHEJ) pathway of DNA repair ...
heterodimer
In biochemistry, a protein dimer is a macromolecular complex formed by two protein monomers, or single proteins, which are usually non-covalently bound. Many macromolecules, such as proteins or nucleic acids, form dimers. The word ''dimer'' has ...
to function. Disrupting these proteins causes telomeric shortening, which can be observed by measuring telomere length with FISH. For example, in mice lacking the Ku 80 gene, the telomere lengths are measured by qFISH and are observed to be significantly shorter.Fagagna, F., Hande, MP., Tong, WM., Roth, D., Lansdorp, PM., Wang, ZQ., and Jackson, SP. "Effects of DNA nonhomologous end-joining factors on telomere length and chromosomal stability in mammalian cells." Current Biology 11 (2001) 15:1192-1196.
Q-FISH is commonly used in cancer
Cancer is a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. These contrast with benign tumors, which do not spread. Possible signs and symptoms include a lump, abnormal b ...
research to measure differences in telomere lengths between cancerous and non-cancerous cells. Telomere shortening causes genomic instability and occurs naturally with advanced age, both factors that correlate with possible causes of cancer.Marcondes, AM., Bair, S., Rabinovitch, PS., Gooley, T., Deeg, HJ., and Risques, R. "No telomere shortening in marrow stroma from patients with MDS." Annals of Hematology (2009) 88:623-628.
Advantages of Q-FISH
The greatest advantage of Q-FISH over other FISH techniques is the quantitative ability of the technique. Compared to traditional FISH which uses DNA probes, quantitative information is difficult to acquire because the hybridization probes compete with the renaturation of complementary genomic DNA strands. Therefore, by using PNAs and hybridizing them under very stringent conditions, it allows one to overcome this issue. Similarly, because one is able to denature the chromosomal DNA in the presence of the PNA probe, it simplifies the FISH procedure. In addition, the method provides greater resolution, allowing the user to examine the telomere length of each individual chromosome ( p or q arm) in a particular cell. Also, unlikeSouthern blot
A Southern blot is a method used in molecular biology for detection of a specific DNA sequence in DNA samples. Southern blotting combines transfer of electrophoresis-separated DNA fragments to a filter membrane and subsequent fragment detecti ...
s which need over 105 cells for a blot, less than 30 cells are needed in Q-FISH.
Drawbacks of Q-FISH
Despite its advantages, Q-FISH is quite labor-intensive and is generally not suitable for high throughput analysis. The technique depends on well-prepared metaphase cells and it is vital that the equipment and samples are adjusted/normalized correctly in order for the quantification to be accurate. Also, while only a small quantity of cells are needed, it is difficult to get a sufficient number in metaphase at once. In addition, poor chromosome morphology may result from overexposure to high temperatures during preparation. Similarly, if one is using different cell types, many of the steps in Q-FISH (such as the length of colcemid treatment) will require optimization. A common problem in fluorescence microscopy isphotobleaching
In optics, photobleaching (sometimes termed fading) is the photochemical alteration of a dye or a fluorophore molecule such that it is permanently unable to fluoresce. This is caused by cleaving of covalent bonds or non-specific reactions between t ...
, where the fluorophore
A fluorophore (or fluorochrome, similarly to a chromophore) is a fluorescent chemical compound that can re-emit light upon light excitation. Fluorophores typically contain several combined aromatic groups, or planar or cyclic molecules with se ...
loses its activity as a result of exposure to light. This can lead to the inaccurate measurement of fluorescence intensity. Photobleaching, light source stability, and system variability are all sources of error but can be minimized if the user is able to reduce the acquisition time between samples and include the proper controls.
Classical technique
Prior to the development of Q-FISH and PNAs, the classical technique for measuring telomere length was the use ofSouthern blot
A Southern blot is a method used in molecular biology for detection of a specific DNA sequence in DNA samples. Southern blotting combines transfer of electrophoresis-separated DNA fragments to a filter membrane and subsequent fragment detecti ...
s. In this method, genomic DNA is digested using restriction enzyme
A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class o ...
s and separated by gel electrophoresis
Gel electrophoresis is a method for separation and analysis of biomacromolecules ( DNA, RNA, proteins, etc.) and their fragments, based on their size and charge. It is used in clinical chemistry to separate proteins by charge or size (IEF ...
. The DNA is then transferred onto a membrane and hybridized using radioactive or fluorescent telomeric DNA probes. However, this method is only able to evaluate average telomere length in a cell population and the presence of interstitial telomeric sequences in the genome can yield inaccurate measurements.Poon, SSS. and Lansdorp, PM (2001) "Quantitative Fluorescence in-situ Hybridization." Current Protocols ''Current Protocols'' is a series of laboratory manuals for life scientists. The first title, ''Current Protocols in Molecular Biology'', was established in 1987 by the founding editors Frederick M. Ausubel, Roger Brent, Robert Kingston, David D. M ...
in Cell Biology (University of Southern California, Los Angeles, California, USA.: John Wiley and Sons, Inc.) Chapter 18 (2001) Section 18.4.1-18.4.21.
Variations of Q-FISH
Flow-FISH
Similar to Q-FISH,Flow-FISH
Flow-FISH (fluorescent in-situ hybridization) is a cytogenetic technique to quantify the copy number of RNA or specific repetitive elements in genomic DNA of whole cell populations via the combination of flow cytometry with cytogenetic fluoresc ...
is an adaptation of Q-FISH that combines the use of PNAs with flow cytometry. In this method, Flow-FISH uses interphase
Interphase is the portion of the cell cycle that is not accompanied by visible changes under the microscope, and includes the G1, S and G2 phases. During interphase, the cell grows (G1), replicates its DNA (S) and prepares for mitosis (G2). A c ...
cells rather than metaphase
Metaphase ( and ) is a stage of mitosis in the eukaryotic cell cycle in which chromosomes are at their second-most condensed and coiled stage (they are at their most condensed in anaphase). These chromosomes, carrying genetic information, align ...
chromosomes and hybridizes the PNA probes in suspension. Following hybridization, thousands of cells can be analyzed on a flow cytometer in a relatively short time. However, Flow-FISH only provides an average telomeric length for each cell whereas Q-FISH is able to analyze the telomere length of an individual chromosome.

PNA-FISH
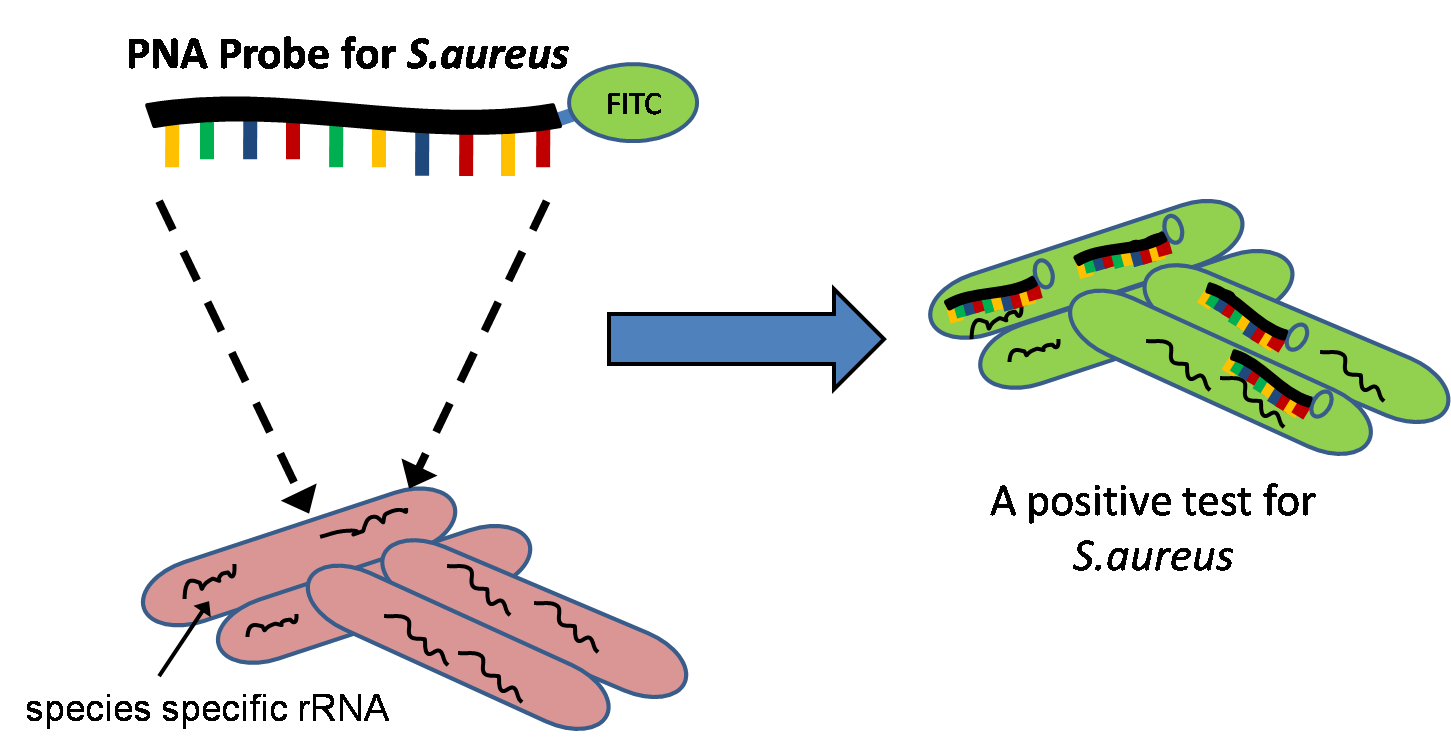
Although the quantitative ability of Q-FISH is most commonly used in telomere research, other fields that only require qualitative data have adopted the use of PNAs with FISH for both research and diagnostic purposes. PNA-FISH assays have been developed for identifying and diagnosing infectious diseases in a rapid manner within the clinic. Combined with traditionalgram staining
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish bac ...
of positive blood cultures, PNAs can be used to target species-specific rRNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosoma ...
(ribosomal RNA) to identify different strains of bacteria or yeast.Stender, H. "PNA-FISH: an intelligent stain for rapid diagnosis of infectious diseases." ''Expert Review in molecular diagnostics'' (2003)5:649-655 Since the test can be performed relatively quickly, the test is being considered for use in hospitals where hospital-acquired infections can occur.
CO-FISH (chromosome orientation-FISH)
Another adaptation that utilizes PNAs and FISH is known as CO-FISH (Chromosome Orientation-FISH) which allows for the labelling of chromosomes with PNAs in a strand specific manner. This method involves the selective removal of newly replicated strands of DNA (through the use of BrdU incorporation), resulting in only single stranded target DNA. By using different colored unidirectional PNA probes, it becomes possible to uniquely label sister chromatids,.Falconer, E.,Chavez, EA., Henderson, A., Poon, SSS., McKinney, S., Brown, L., Huntsman, DG., and Lansdorp, PM."Identification of sister chromatids by DNA template strand sequences." ''Nature'' (2010)463:93-98.References