|
Nucleosome Structure-2
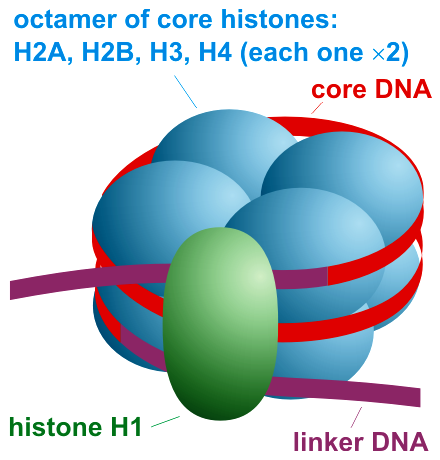
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundamental subunit of chromatin. Each nucleosome is composed of a little less than two turns of DNA wrapped around a set of eight proteins called histones, which are known as a histone octamer. Each histone octamer is composed of two copies each of the histone proteins H2A, H2B, H3, and H4. DNA must be compacted into nucleosomes to fit within the cell nucleus. In addition to nucleosome wrapping, eukaryotic chromatin is further compacted by being folded into a series of more complex structures, eventually forming a chromosome. Each human cell contains about 30 million nucleosomes. Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. Nucleosome positions in the ge ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Basic Units Of Chromatin Structure
BASIC (Beginners' All-purpose Symbolic Instruction Code) is a family of general-purpose, high-level programming languages designed for ease of use. The original version was created by John G. Kemeny and Thomas E. Kurtz at Dartmouth College in 1963. They wanted to enable students in non-scientific fields to use computers. At the time, nearly all computers required writing custom software, which only scientists and mathematicians tended to learn. In addition to the program language, Kemeny and Kurtz developed the Dartmouth Time Sharing System (DTSS), which allowed multiple users to edit and run BASIC programs simultaneously on remote terminals. This general model became very popular on minicomputer systems like the PDP-11 and Data General Nova in the late 1960s and early 1970s. Hewlett-Packard produced an entire computer line for this method of operation, introducing the HP2000 series in the late 1960s and continuing sales into the 1980s. Many early video games trace their h ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Supercoil
DNA supercoiling refers to the amount of twist in a particular DNA strand, which determines the amount of strain on it. A given strand may be "positively supercoiled" or "negatively supercoiled" (more or less tightly wound). The amount of a strand’s supercoiling affects a number of biological processes, such as compacting DNA and regulating access to the genetic code (which strongly affects DNA metabolism and possibly gene expression). Certain enzymes, such as topoisomerases, change the amount of DNA supercoiling to facilitate functions such as DNA replication and transcription. The amount of supercoiling in a given strand is described by a mathematical formula that compares it to a reference state known as "relaxed B-form" DNA. Overview In a "relaxed" double-helical segment of B-DNA, the two strands twist around the helical axis once every 10.4–10.5 base pairs of sequence. Adding or subtracting twists, as some enzymes do, imposes strain. If a DNA segment under twist ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleosome Core Particle 1EQZ V
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundamental subunit of chromatin. Each nucleosome is composed of a little less than two turns of DNA wrapped around a set of eight proteins called histones, which are known as a histone octamer. Each histone octamer is composed of two copies each of the histone proteins H2A, H2B, H3, and H4. DNA must be compacted into nucleosomes to fit within the cell nucleus. In addition to nucleosome wrapping, eukaryotic chromatin is further compacted by being folded into a series of more complex structures, eventually forming a chromosome. Each human cell contains about 30 million nucleosomes. Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. Nucleosome positions in the geno ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleosome Organization
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundamental subunit of chromatin. Each nucleosome is composed of a little less than two turns of DNA wrapped around a set of eight proteins called histones, which are known as a histone octamer. Each histone octamer is composed of two copies each of the histone proteins H2A, H2B, H3, and H4. DNA must be compacted into nucleosomes to fit within the cell nucleus. In addition to nucleosome wrapping, eukaryotic chromatin is further compacted by being folded into a series of more complex structures, eventually forming a chromosome. Each human cell contains about 30 million nucleosomes. Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. Nucleosome positions in the geno ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
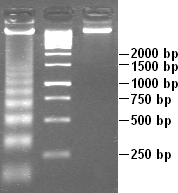
Apoptotic DNA Laddering
Apoptosis (from grc, ἀπόπτωσις, apóptōsis, 'falling off') is a form of programmed cell death that occurs in multicellular organisms. Biochemical events lead to characteristic cell changes (morphology) and death. These changes include blebbing, cell shrinkage, nuclear fragmentation, chromatin condensation, DNA fragmentation, and mRNA decay. The average adult human loses between 50 and 70 billion cells each day due to apoptosis. For an average human child between eight and fourteen years old, approximately twenty to thirty billion cells die per day. In contrast to necrosis, which is a form of traumatic cell death that results from acute cellular injury, apoptosis is a highly regulated and controlled process that confers advantages during an organism's life cycle. For example, the separation of fingers and toes in a developing human embryo occurs because cells between the digits undergo apoptosis. Unlike necrosis, apoptosis produces cell fragments called apoptotic bod ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Root Mean Square Deviation
The root-mean-square deviation (RMSD) or root-mean-square error (RMSE) is a frequently used measure of the differences between values (sample or population values) predicted by a model or an estimator and the values observed. The RMSD represents the square root of the second sample moment of the differences between predicted values and observed values or the quadratic mean of these differences. These deviations are called '' residuals'' when the calculations are performed over the data sample that was used for estimation and are called ''errors'' (or prediction errors) when computed out-of-sample. The RMSD serves to aggregate the magnitudes of the errors in predictions for various data points into a single measure of predictive power. RMSD is a measure of accuracy, to compare forecasting errors of different models for a particular dataset and not between datasets, as it is scale-dependent. RMSD is always non-negative, and a value of 0 (almost never achieved in practice) would ind ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Palindromic Sequence
A palindromic sequence is a nucleic acid sequence in a double-stranded DNA or RNA molecule whereby reading in a certain direction (e.g. 5' to 3') on one strand is identical to the sequence in the same direction (e.g. 5' to 3') on the complementary strand. This definition of palindrome thus depends on complementary strands being palindromic of each other. The meaning of palindrome in the context of genetics is slightly different from the definition used for words and sentences. Since a double helix is formed by two paired antiparallel strands of nucleotides that run in opposite directions, and the nucleotides always pair in the same way (adenine (A) with thymine (T) in DNA or uracil (U) in RNA; cytosine (C) with guanine (G)), a (single-stranded) nucleotide sequence is said to be a palindrome if it is equal to its reverse complement. For example, the DNA sequence ACCTAGGT is palindromic with its nucleotide-by-nucleotide complement TGGATCCA because reversing the order of the nuc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Centromere
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers attach to the centromere via the kinetochore. The physical role of the centromere is to act as the site of assembly of the kinetochores – a highly complex multiprotein structure that is responsible for the actual events of chromosome segregation – i.e. binding microtubules and signaling to the cell cycle machinery when all chromosomes have adopted correct attachments to the spindle, so that it is safe for cell division to proceed to completion and for cells to enter anaphase. There are, broadly speaking, two types of centromeres. "Point centromeres" bind to specific proteins that recognize particular DNA sequences with high efficiency. Any piece of DNA with the point centromere DNA sequence on it will typically form a centromere if pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Crystal Structure
In crystallography, crystal structure is a description of the ordered arrangement of atoms, ions or molecules in a crystal, crystalline material. Ordered structures occur from the intrinsic nature of the constituent particles to form symmetric patterns that repeat along the principal directions of Three-dimensional space (mathematics), three-dimensional space in matter. The smallest group of particles in the material that constitutes this repeating pattern is the unit cell of the structure. The unit cell completely reflects the symmetry and structure of the entire crystal, which is built up by repetitive Translation (geometry), translation of the unit cell along its principal axes. The translation vectors define the nodes of the Bravais lattice. The lengths of the principal axes, or edges, of the unit cell and the angles between them are the lattice constants, also called ''lattice parameters'' or ''cell parameters''. The symmetry properties of the crystal are described by the con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleosome 1KX5 Colour Coded
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundamental subunit of chromatin. Each nucleosome is composed of a little less than two turns of DNA wrapped around a set of eight proteins called histones, which are known as a histone octamer. Each histone octamer is composed of two copies each of the histone proteins H2A, H2B, H3, and H4. DNA must be compacted into nucleosomes to fit within the cell nucleus. In addition to nucleosome wrapping, eukaryotic chromatin is further compacted by being folded into a series of more complex structures, eventually forming a chromosome. Each human cell contains about 30 million nucleosomes. Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. Nucleosome positions in the geno ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaebacteria kingdom), but this term has fallen out of use. Archaeal cells have unique properties separating them from the other two domains, Bacteria and Eukaryota. Archaea are further divided into multiple recognized phyla. Classification is difficult because most have not been isolated in a laboratory and have been detected only by their gene sequences in environmental samples. Archaea and bacteria are generally similar in size and shape, although a few archaea have very different shapes, such as the flat, square cells of ''Haloquadratum walsbyi''. Despite this morphological similarity to bacteria, archaea possess genes and several metabolic pathways that are more closely related to those of eukaryotes, notably for the enzymes involved ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protamines
Protamines are small, arginine-rich, nuclear proteins that replace histones late in the haploid phase of spermatogenesis and are believed essential for sperm head condensation and DNA stabilization. They may allow for denser packaging of DNA in the spermatozoon than histones, but they must be decompressed before the genetic data can be used for protein synthesis. However, in humans and maybe other primates, 10-15% of the sperm's genome is packaged by histones thought to bind genes that are essential for early embryonic development. Protamine and protamine-like (PL) proteins are collectively known as the sperm-specific nuclear basic proteins (SNBPs). The PL proteins are intermediate in structure between protamine and Histone H1. The C-terminal domain of PL could be the precursor of vertebrate protamine. Spermatogenesis During the formation of sperm, protamine binds to the phosphate backbone of DNA using the arginine-rich domain as an anchor. DNA is then folded into a toroid, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |





.jpg)
