|
H2AX
H2A histone family member X (usually abbreviated as H2AX) is a type of histone protein from the H2A family encoded by the ''H2AFX'' gene. An important phosphorylated form is γH2AX (S139), which forms when double-strand breaks appear. In humans and other eukaryotes, the DNA is wrapped around histone octamers, consisting of core histones H2A, H2B, H3 and H4, to form chromatin. H2AX contributes to nucleosome-formation, chromatin-remodeling and DNA repair, and is also used ''in vitro'' as an assay for double-strand breaks in dsDNA. Formation of γH2AX H2AX becomes phosphorylated on serine 139, then called γH2AX, as a reaction on DNA double-strand breaks (DSB). The kinases of the PI3-family (Ataxia telangiectasia mutated, ATR and DNA-PKcs) are responsible for this phosphorylation, especially ATM. The modification can happen accidentally during replication fork collapse or in the response to ionizing radiation but also during controlled physiological processes such as V(D)J re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Repair
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages (interstrand crosslinks or ICLs). This can eventually lead to malignant ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin Remodeling
Chromatin remodeling is the dynamic modification of chromatin architecture to allow access of condensed genomic DNA to the regulatory transcription machinery proteins, and thereby control gene expression. Such remodeling is principally carried out by 1) covalent histone modifications by specific enzymes, e.g., histone acetyltransferases (HATs), deacetylases, methyltransferases, and kinases, and 2) ATP-dependent chromatin remodeling complexes which either move, eject or restructure nucleosomes. Besides actively regulating gene expression, dynamic remodeling of chromatin imparts an epigenetic regulatory role in several key biological processes, egg cells DNA replication and repair; apoptosis; chromosome segregation as well as development and pluripotency. Aberrations in chromatin remodeling proteins are found to be associated with human diseases, including cancer. Targeting chromatin remodeling pathways is currently evolving as a major therapeutic strategy in the treatment of several ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone
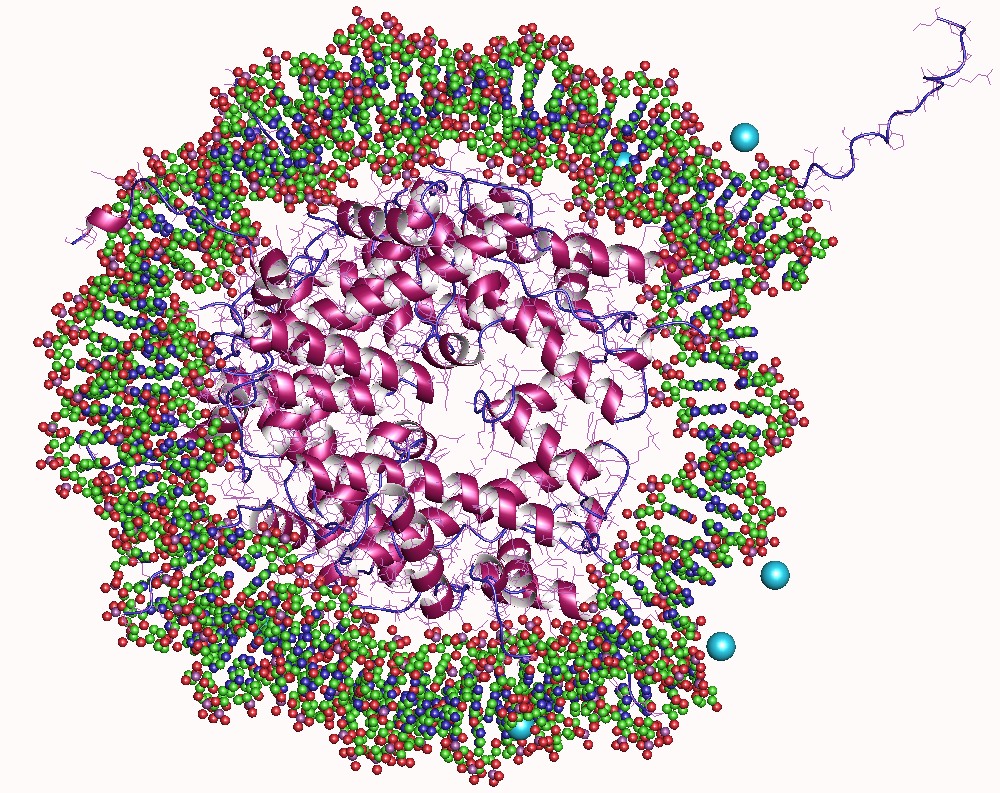
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wrapped into 30-nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 90 micrometers (0.09 mm) of 30 nm diameter chromatin fibers. There are five families of histones which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B dimers and a H3-H4 tetramer. The tight wrapping of DNA around histones ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Core Histones
In biology, histones are highly Base (chemistry), basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wrapped into 30-nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 90 micrometers (0.09 mm) of 30 nm diameter chromatin fibers. There are five families of histones which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B protein dimer, dimers and a H3-H4 tetrameric protein, te ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNF8
E3 ubiquitin-protein ligase RNF8 is an enzyme that in humans is encoded by the ''RNF8'' gene. RNF8 has activity both in immune system functions and in DNA repair. Function The protein encoded by this gene contains a RING finger motif and an FHA domain. This protein has been shown to interact with several class II ubiquitin-conjugating enzymes (E2), including UBE2E1/UBCH6, UBE2E2, and UBE2E3, and may act as a ubiquitin ligase (E3) in the ubiquitination of certain nuclear proteins. Alternatively spliced transcript variants encoding distinct isoforms have been reported. RNF8 promotes repair of DNA damage through three DNA repair pathways: homologous recombinational repair (HRR), non-homologous end joining (NHEJ), and nucleotide excision repair (NER). DNA damage is considered to be the primary cause of cancer, and deficiency in DNA repair can cause mutations leading to cancer. A deficiency in RNF8 predisposes mice to cancer. Chromatin remodeling After the occurrence of a do ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin
Chromatin is a complex of DNA and protein found in eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important roles in reinforcing the DNA during cell division, preventing DNA damage, and regulating gene expression and DNA replication. During mitosis and meiosis, chromatin facilitates proper segregation of the chromosomes in anaphase; the characteristic shapes of chromosomes visible during this stage are the result of DNA being coiled into highly condensed chromatin. The primary protein components of chromatin are histones. An octamer of two sets of four histone cores (Histone H2A, Histone H2B, Histone H3, and Histone H4) bind to DNA and function as "anchors" around which the strands are wound.Maeshima, K., Ide, S., & Babokhov, M. (2019). Dynamic chromatin organization without the 30-nm fiber. ''Current opinion in cell biology, 58,'' 95–104. https://doi.o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MDC1
Mediator of DNA damage checkpoint protein 1 is a 2080 amino acid long protein that in humans is encoded by the ''MDC1'' gene located on the short arm (p) of chromosome 6. MDC1 protein is a regulator of the Intra-S phase and the G2/M cell cycle checkpoints and recruits repair proteins to the site of DNA damage. It is involved in determining cell survival fate in association with tumor suppressor protein p53. This protein also goes by the name Nuclear Factor with BRCT Domain 1 (NFBD1). Function Role in DNA damage response The ''MDC1'' gene encodes the MDC1 nuclear protein which is part of the DNA damage response (DDR) pathway, the mechanism through which eukaryotic cells respond to damaged DNA, specifically DNA double-strand breaks (DSB) that are caused by ionizing radiation or chemical clastogens. The DDR of mammalian cells is made up of kinases, and mediator/adaptors factors. In mammalian cells the DDR is a network of pathways made up of proteins that function as either kinas ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ataxia Telangiectasia Mutated
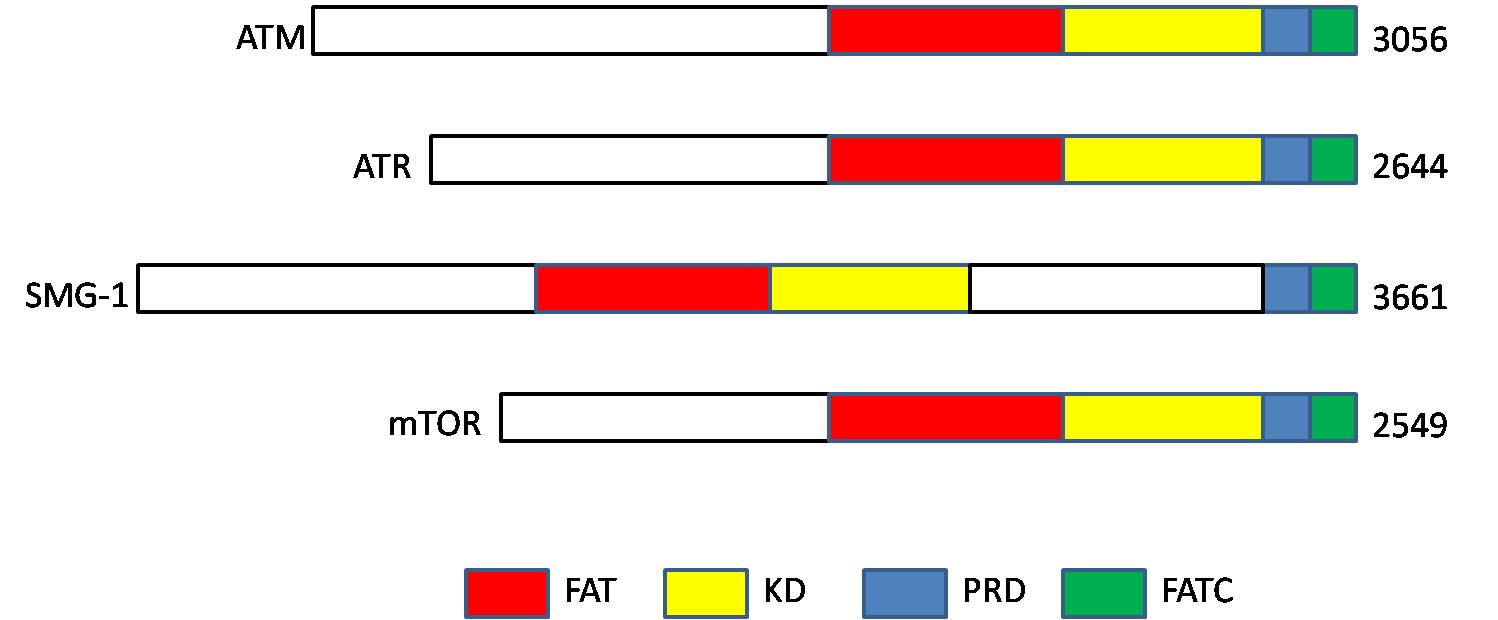
ATM serine/threonine kinase or Ataxia-telangiectasia mutated, symbol ATM, is a serine/threonine protein kinase that is recruited and activated by DNA double-strand breaks. It phosphorylates several key proteins that initiate activation of the DNA damage checkpoint, leading to cell cycle arrest, DNA repair or apoptosis. Several of these targets, including p53, CHK2, BRCA1, NBS1 and H2AX are tumor suppressors. In 1995, the gene was discovered by Yosef Shiloh who named its product ATM since he found that its mutations are responsible for the disorder ataxia–telangiectasia#Cause, ataxia–telangiectasia. In 1998, the Shiloh and Michael B. Kastan, Kastan laboratories independently showed that ATM is a protein kinase whose activity is enhanced by DNA damage. Introduction Throughout the cell cycle DNA is monitored for damage. Damages result from errors during DNA replication, replication, by-products of metabolism, general toxic drugs or ionizing radiation. The cell cycle has diffe ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone H2A
Histone H2A is one of the five main histone proteins involved in the structure of chromatin in eukaryotic cells. The other histone proteins are: H1, H2B, H3 and H4. Background Histones are proteins that package DNA into nucleosomes. Histones are responsible for maintaining the shape and structure of a nucleosome. One chromatin molecule is composed of at least one of each core histones per 100 base pairs of DNA. There are five families of histones known to date; these histones are termed H1/H5, H2A, H2B, H3, and H4. H2A is considered a core histone, along with H2B, H3 and H4. Core formation first occurs through the interaction of two H2A molecules. Then, H2A forms a dimer with H2B; the core molecule is complete when H3-H4 also attaches to form a tetramer. Sequence variants Histone H2A is composed of non-allelic variants. The term "Histone H2A" is intentionally non-specific and refers to a variety of closely related proteins that vary often by only a few amino acids. Apart ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ATM Serine/threonine Kinase
ATM serine/threonine kinase or Ataxia-telangiectasia mutated, symbol ATM, is a serine/ threonine protein kinase that is recruited and activated by DNA double-strand breaks. It phosphorylates several key proteins that initiate activation of the DNA damage checkpoint, leading to cell cycle arrest, DNA repair or apoptosis. Several of these targets, including p53, CHK2, BRCA1, NBS1 and H2AX are tumor suppressors. In 1995, the gene was discovered by Yosef Shiloh who named its product ATM since he found that its mutations are responsible for the disorder ataxia–telangiectasia. In 1998, the Shiloh and Kastan laboratories independently showed that ATM is a protein kinase whose activity is enhanced by DNA damage. Introduction Throughout the cell cycle DNA is monitored for damage. Damages result from errors during replication, by-products of metabolism, general toxic drugs or ionizing radiation. The cell cycle has different DNA damage checkpoints, which inhibit the next or maint ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nibrin
Nibrin, also known as NBN or NBS1, is a protein which in humans is encoded by the ''NBN'' gene. Function Nibrin is a protein associated with the repair of double strand breaks (DSBs) which pose serious damage to a genome. It is a 754 amino acid protein identified as a member of the NBS1/hMre11/RAD50(N/M/R, more commonly referred to as MRN) double strand DNA break repair complex. This complex recognizes DNA damage and rapidly relocates to DSB sites and forms nuclear foci. It also has a role in regulation of N/M/R (MRN) protein complex activity which includes end-processing of both physiological and mutagenic DNA double strand breaks (DSBs). Cellular response to DSBs Cellular response is performed by damage sensors, effectors of lesion repair and signal transduction. The central role is carried out by ataxia telangiectasia mutated (ATM) by activating the DSB signaling cascade, phosphorylating downstream substrates such as histone H2AX and NBS1. NBS1 relocates to DSB sites ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleosome
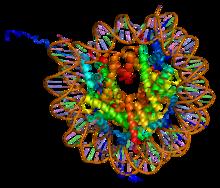
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundamental subunit of chromatin. Each nucleosome is composed of a little less than two turns of DNA wrapped around a set of eight proteins called histones, which are known as a histone octamer. Each histone octamer is composed of two copies each of the histone proteins H2A, H2B, H3, and H4. DNA must be compacted into nucleosomes to fit within the cell nucleus. In addition to nucleosome wrapping, eukaryotic chromatin is further compacted by being folded into a series of more complex structures, eventually forming a chromosome. Each human cell contains about 30 million nucleosomes. Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. Nucleosome positions in the gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |



.png)