|
Branched DNA
In biology, a branched DNA assay is a signal amplification assay (as opposed to a target amplification assay) that is used to detect nucleic acid molecules. Method A branched DNA assay begins with a dish or some other solid support (e.g., a plastic dipstick). The dish is peppered with small, single stranded DNA molecules (or chains) that stick out into the solution. These are known as capture probe DNA molecules. Next, an extender DNA molecule is added. Each extender has two domains; one that hybridizes to the capture DNA molecule and one that sticks out above the surface. The purpose of the extender is two-fold. First, it creates more available surface area for target DNA molecules to bind, and second, it allows the assay to be easily adapted to detect a variety of target DNA molecules. Once the capture and extender molecules are in place and they have hybridized, the sample can be added. Target molecules in the sample will bind to the extender molecule. This results in a base p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Nucleic Acid
Nucleic acids are large biomolecules that are crucial in all cells and viruses. They are composed of nucleotides, which are the monomer components: a pentose, 5-carbon sugar, a phosphate group and a nitrogenous base. The two main classes of nucleic acids are deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). If the sugar is ribose, the polymer is RNA; if the sugar is deoxyribose, a variant of ribose, the polymer is DNA. Nucleic acids are chemical compounds that are found in nature. They carry information in cells and make up genetic material. These acids are very common in all living things, where they create, encode, and store information in every living cell of every outline of life forms, life-form on Earth. In turn, they send and express that information inside and outside the cell nucleus. From the inner workings of the cell to the young of a living thing, they contain and provide information via the nucleic acid sequence. This gives the RNA and DNA their unmistakable 'la ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Nucleic Acid Hybridization
In molecular biology, hybridization (or hybridisation) is a phenomenon in which single-stranded deoxyribonucleic acid (DNA) or ribonucleic acid (RNA) molecules anneal to complementary DNA or RNA. Though a double-stranded DNA sequence is generally stable under physiological conditions, changing these conditions in the laboratory (generally by raising the surrounding temperature) will cause the molecules to separate into single strands. These strands are complementary to each other but may also be complementary to other sequences present in their surroundings. Lowering the surrounding temperature allows the single-stranded molecules to anneal or “hybridize” to each other. DNA replication and transcription of DNA into RNA both rely upon nucleotide hybridization, as do molecular biology techniques including Southern blots and Northern blots, the polymerase chain reaction (PCR), and most approaches to DNA sequencing. Applications Hybridization is a basic property of nucleotide ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, Recombinant DNA, research, and Forensic DNA, forensics. Commonly made in the laboratory by Oligonucleotide synthesis, solid-phase chemical synthesis, these small fragments of nucleic acids can be manufactured as single-stranded molecules with any user-specified sequence, and so are vital for artificial gene synthesis, polymerase chain reaction (PCR), DNA sequencing, molecular cloning and as Fluorescence in situ hybridization, molecular probes. In nature, oligonucleotides are usually found as small RNA molecules that function in the regulation of gene expression (e.g. microRNA), or are degradation intermediates derived from the breakdown of larger nucleic acid molecules. Oligonucleotides are characterized by the Nucleic acid sequence, sequence of nucleotide residues that make up the entire molecule. The length of the oligonucleotide is usually denoted by "Monomer, -m ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Alkaline Phosphatase
The enzyme alkaline phosphatase (ALP, alkaline phenyl phosphatase, also abbreviated PhoA) is a phosphatase with the physiological role of dephosphorylating compounds. The enzyme is found across a multitude of organisms, prokaryotes and eukaryotes alike, with the same general function, but in different structural forms suitable to the environment they function in. Alkaline phosphatase is found in the periplasmic space of '' E. coli'' bacteria. This enzyme is heat stable and has its maximum activity at high pH. In humans, it is found in many forms depending on its origin within the body – it plays an integral role in metabolism within the liver and development within the skeleton. Due to its widespread prevalence in these areas, its concentration in the bloodstream is used by diagnosticians as a biomarker in helping determine diagnoses such as hepatitis or osteomalacia. The level of alkaline phosphatase in the blood is checked through the ALP test, which is often par ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Assay
An assay is an investigative (analytic) procedure in laboratory medicine, mining, pharmacology, environmental biology and molecular biology for qualitatively assessing or quantitatively measuring the presence, amount, or functional activity of a target entity. The measured entity is often called the analyte, the measurand, or the target of the assay. The analyte can be a drug, biochemical substance, chemical element or compound, or cell in an organism or organic sample. An assay usually aims to measure an analyte's intensive property and express it in the relevant measurement unit (e.g. molarity, density, functional activity in enzyme international units, degree of effect in comparison to a standard, etc.). If the assay involves exogenous reactants (the reagents), then their quantities are kept fixed (or in excess) so that the quantity and quality of the target are the only limiting factors. The difference in the assay outcome is used to deduce the unknown quality ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Reverse Transcription
A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B virus, hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryote, eukaryotic cells to extend the telomeres at the ends of their Chromosome#Eukaryotes, linear chromosomes. The process does not violate the flows of genetic information as described by the classical central dogma of molecular biology, central dogma, but rather expands it to include transfers of information from RNA to DNA. Retroviral RT has three sequential biochemical activities: RNA-dependent DNA polymerase activity, ribonuclease H (RNase H), and DNA-dependent DNA polymerase activity. Collectively, these activities enable the enzyme to convert single-stranded RNA into double-stranded cDNA. In retroviruses and retrotransposons, this cDNA can then inte ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to make millions to billions of copies of a specific DNA sample rapidly, allowing scientists to amplify a very small sample of DNA (or a part of it) sufficiently to enable detailed study. PCR was invented in 1983 by American biochemist Kary Mullis at Cetus Corporation. Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research and forensic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Northern Blot
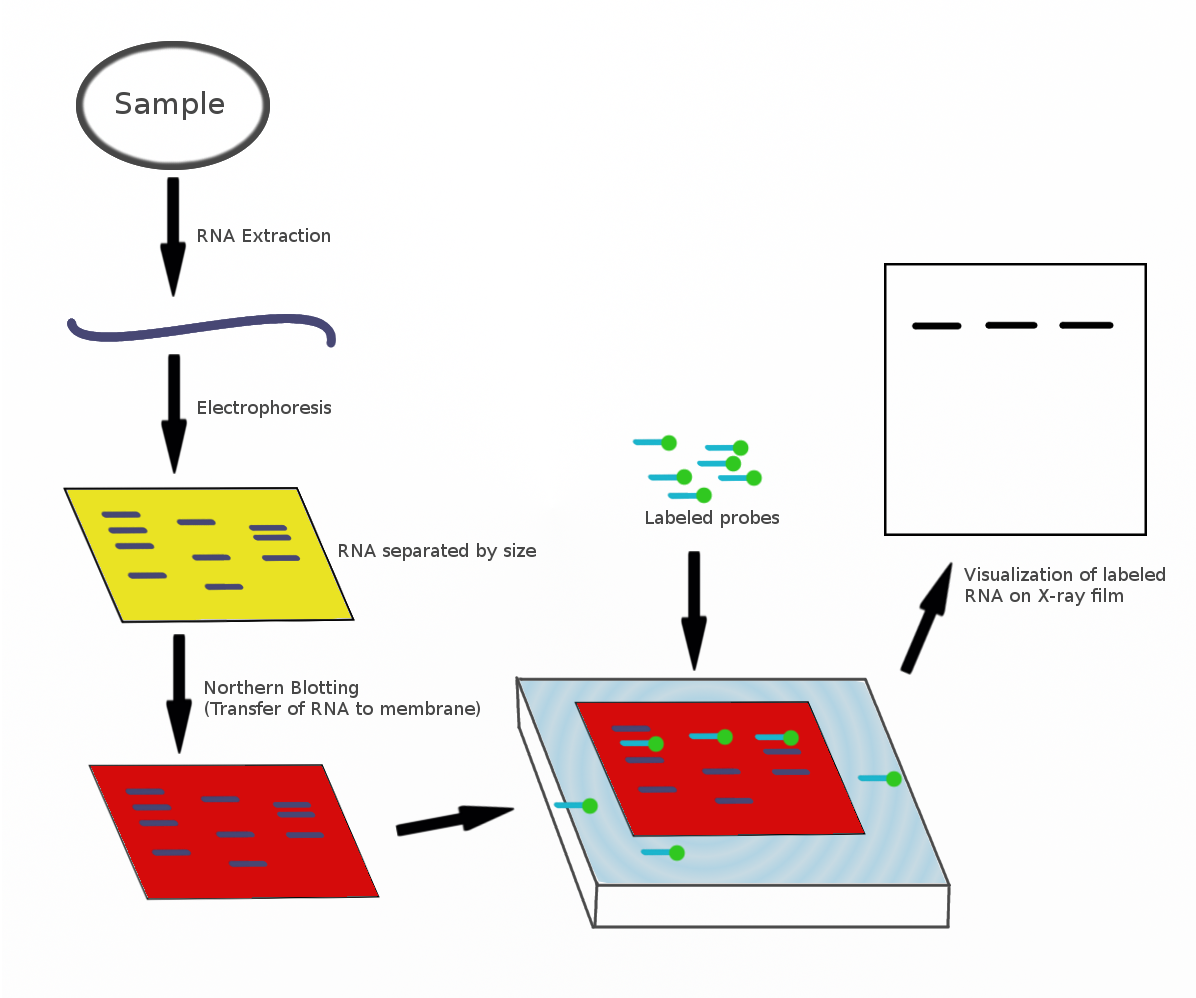
The northern blot, or RNA blot,Gilbert, S. F. (2000) Developmental Biology, 6th Ed. Sunderland MA, Sinauer Associates. is a technique used in molecular biology research to study gene expression by detection of RNA (or isolated mRNA) in a sample.Kevil, C. G., Walsh, L., Laroux, F. S., Kalogeris, T., Grisham, M. B., Alexander, J. S. (1997) An Improved, Rapid Northern Protocol. Biochem. and Biophys. Research Comm. 238:277–279. With northern blotting it is possible to observe cellular control over structure and function by determining the particular gene expression rates during differentiation and morphogenesis, as well as in abnormal or diseased conditions. Northern blotting involves the use of electrophoresis to separate RNA samples by size, and detection with a hybridization probe complementary to part of or the entire target sequence. Strictly speaking, the term 'northern blot' refers specifically to the capillary transfer of RNA from the electrophoresis gel to the blotting m ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Quantitative PCR
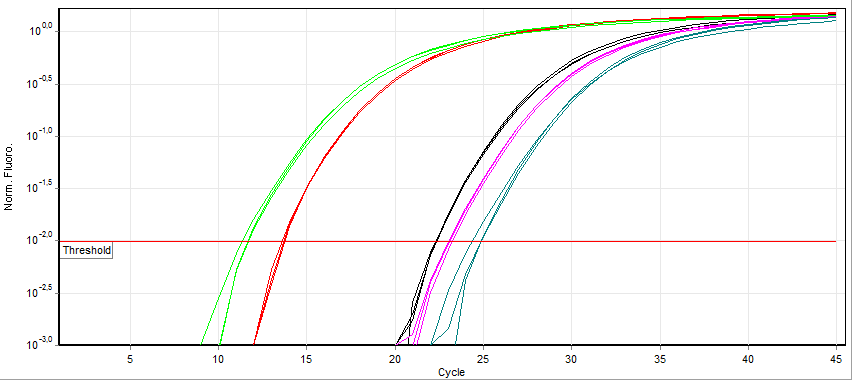
A real-time polymerase chain reaction (real-time PCR, or qPCR when used quantitatively) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real time), not at its end, as in conventional PCR. Real-time PCR can be used quantitatively and semi-quantitatively (i.e., above/below a certain amount of DNA molecules). Two common methods for the detection of PCR products in real-time PCR are (1) non-specific fluorescent dyes that intercalate with any double-stranded DNA and (2) sequence-specific DNA probes consisting of oligonucleotides that are labelled with a fluorescent reporter, which permits detection only after hybridization of the probe with its complementary sequence. The Minimum Information for Publication of Quantitative Real-Time PCR Experiments (MIQE) guidelines propose that the abbreviation ''qPCR'' be used for quantitative real-time PCR and that ' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
CDNA
In genetics, complementary DNA (cDNA) is DNA that was reverse transcribed (via reverse transcriptase) from an RNA (e.g., messenger RNA or microRNA). cDNA exists in both single-stranded and double-stranded forms and in both natural and engineered forms. In engineered forms, it often is a copy (replicate) of the naturally occurring DNA from any particular organism's natural genome; the organism's own mRNA was naturally transcribed from its DNA, and the cDNA is reverse transcribed from the mRNA, yielding a duplicate of the original DNA. Engineered cDNA is often used to express a specific protein in a cell that does not normally express that protein (i.e., heterologous expression), or to sequence or quantify mRNA molecules using DNA based methods (qPCR, RNA-seq). cDNA that codes for a specific protein can be transferred to a recipient cell for expression as part of recombinant DNA, often bacterial or yeast expression systems. cDNA is also generated to analyze transcriptomic pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
CfDNA
Circulating free DNA (cfDNA) (also known as cell-free DNA) are degraded DNA fragments released to body fluids such as blood plasma, urine, cerebrospinal fluid, etc. Typical sizes of cfDNA fragments reflect chromatosome particles (~165bp), as well as multiples of nucleosomes, which protect DNA from digestion by apoptotic nucleases. The term cfDNA can be used to describe various forms of DNA freely circulating in body fluids, including circulating tumor DNA (ctDNA), cell-free mitochondrial DNA (ccf mtDNA), cell-free fetal DNA (cffDNA) and donor-derived cell-free DNA (dd-cfDNA). Elevated levels of cfDNA are observed in cancer, especially in advanced disease. There is evidence that cfDNA becomes increasingly frequent in circulation with the onset of age. cfDNA has been shown to be a useful biomarker for a multitude of ailments other than cancer and fetal medicine. This includes but is not limited to trauma, sepsis, aseptic inflammation, myocardial infarction, stroke, transplantatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |
Dendrimer
Dendrimers are highly ordered, Branching (polymer chemistry), branched molecules, polymeric molecules. Synonymous terms for dendrimer include arborols and cascade molecules. Typically, dendrimers are symmetric about the core, and often adopt a spherical three-dimensional morphology. The word dendron is also encountered frequently. A dendron usually contains a single chemically addressable group called the focal point or core. The difference between dendrons and dendrimers is illustrated in the top figure, but the terms are typically encountered interchangeably. The first dendrimers were made by #Divergent methods, divergent synthesis approaches by Fritz Vögtle in 1978, R.G. Denkewalter at Allied Corporation in 1981, Donald Tomalia at Dow Chemical in 1983 and in 1985, and by George R. Newkome in 1985. In 1990 a #Convergent methods, convergent synthetic approach was introduced by Craig Hawker and Jean Fréchet. Dendrimer popularity then greatly increased, resulting in more tha ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] [Amazon] |