rRNA on:
[Wikipedia]
[Google]
[Amazon]
Ribosomal ribonucleic acid (rRNA) is a type of
 Ribosomal RNA organizes into two types of major ribosomal subunit: the large subunit (LSU) and the small subunit (SSU). One of each type come together to form a functioning ribosome. The subunits are at times referred to by their size-sedimentation measurements (a number with an "S" suffix). In prokaryotes, the LSU and SSU are called the 50S and 30S subunits, respectively. In eukaryotes, they are a little larger; the LSU and SSU of eukaryotes are termed the 60S and 40S subunits, respectively.
In the ribosomes of prokaryotes such as
Ribosomal RNA organizes into two types of major ribosomal subunit: the large subunit (LSU) and the small subunit (SSU). One of each type come together to form a functioning ribosome. The subunits are at times referred to by their size-sedimentation measurements (a number with an "S" suffix). In prokaryotes, the LSU and SSU are called the 50S and 30S subunits, respectively. In eukaryotes, they are a little larger; the LSU and SSU of eukaryotes are termed the 60S and 40S subunits, respectively.
In the ribosomes of prokaryotes such as
 Universally conserved secondary structural elements in rRNA among different species show that these sequences are some of the oldest discovered. They serve critical roles in forming the catalytic sites of translation of mRNA. During translation of mRNA, rRNA functions to bind both mRNA and tRNA to facilitate the process of translating mRNA's codon sequence into amino acids. rRNA initiates the catalysis of protein synthesis when tRNA is sandwiched between the SSU and LSU. In the SSU, the mRNA interacts with the anticodons of the tRNA. In the LSU, the amino acid acceptor stem of the tRNA interacts with the LSU rRNA. The ribosome catalyzes ester-amide exchange, transferring the C-terminus of a nascent peptide from a tRNA to the amine of an amino acid. These processes are able to occur due to sites within the ribosome in which these molecules can bind, formed by the rRNA stem-loops. A ribosome has three of these binding sites called the A, P and E sites:
* In general, the A (aminoacyl) site contains an aminoacyl-tRNA (a
Universally conserved secondary structural elements in rRNA among different species show that these sequences are some of the oldest discovered. They serve critical roles in forming the catalytic sites of translation of mRNA. During translation of mRNA, rRNA functions to bind both mRNA and tRNA to facilitate the process of translating mRNA's codon sequence into amino acids. rRNA initiates the catalysis of protein synthesis when tRNA is sandwiched between the SSU and LSU. In the SSU, the mRNA interacts with the anticodons of the tRNA. In the LSU, the amino acid acceptor stem of the tRNA interacts with the LSU rRNA. The ribosome catalyzes ester-amide exchange, transferring the C-terminus of a nascent peptide from a tRNA to the amine of an amino acid. These processes are able to occur due to sites within the ribosome in which these molecules can bind, formed by the rRNA stem-loops. A ribosome has three of these binding sites called the A, P and E sites:
* In general, the A (aminoacyl) site contains an aminoacyl-tRNA (a
 In contrast, eukaryotes generally have many copies of the rRNA genes organized in
In contrast, eukaryotes generally have many copies of the rRNA genes organized in
 Ribosomal RNA characteristics are important in evolution, thus taxonomy and medicine.
* rRNA is one of only a few gene products present in all cells. For this reason, genes that encode the rRNA ( rDNA) are sequenced to identify an organism's Taxonomy (biology), taxonomic group, calculate related groups, and estimate rates of Genetic divergence, species divergence. As a result, many thousands of rRNA sequences are known and stored in specialized databases such as RDP-II and SILVA.
* Alterations to rRNA are what allow certain disease-causing
Ribosomal RNA characteristics are important in evolution, thus taxonomy and medicine.
* rRNA is one of only a few gene products present in all cells. For this reason, genes that encode the rRNA ( rDNA) are sequenced to identify an organism's Taxonomy (biology), taxonomic group, calculate related groups, and estimate rates of Genetic divergence, species divergence. As a result, many thousands of rRNA sequences are known and stored in specialized databases such as RDP-II and SILVA.
* Alterations to rRNA are what allow certain disease-causing
16S rRNA, BioMineWiki
Ribosomal Database Project II
*
SILVA rRNA Database Project
(also includes Eukaryotes (18S) and LSU (23S/28S))
''Halococcus morrhuae'' (archaebacterium) 5S rRNA
{{Ribosome subunits Ribosomal RNA, Protein biosynthesis RNA Non-coding RNA Ribozymes
non-coding RNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not Translation (genetics), translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally im ...
which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozymes demons ...
which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal DNA
Ribosomal DNA (rDNA) is a DNA sequence that codes for ribosomal RNA. These sequences regulate transcription initiation and amplification, and contain both transcribed and non-transcribed spacer segments.
In the human genome there are 5 chromo ...
(rDNA) and then bound to ribosomal protein
A ribosomal protein (r-protein or rProtein) is any of the proteins that, in conjunction with rRNA, make up the ribosomal subunits involved in the cellular process of translation. ''E. coli'', other bacteria and Archaea have a 30S small subunit an ...
s to form small and large
Large means of great size.
Large may also refer to:
Mathematics
* Arbitrarily large, a phrase in mathematics
* Large cardinal, a property of certain transfinite numbers
* Large category, a category with a proper class of objects and morphisms ( ...
ribosome subunits. rRNA is the physical and mechanical factor of the ribosome that forces transfer RNA (tRNA) and messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the p ...
(mRNA) to process and translate
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transl ...
the latter into proteins. Ribosomal RNA is the predominant form of RNA found in most cells; it makes up about 80% of cellular RNA despite never being translated into proteins itself. Ribosomes are composed of approximately 60% rRNA and 40% ribosomal proteins by mass.
Structure
Although the primary structure of rRNA sequences can vary across organisms,base-pairing
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA ...
within these sequences commonly forms stem-loop configurations. The length and position of these rRNA stem-loops allow them to create three-dimensional rRNA structures that are similar across species
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriate s ...
. Because of these configurations, rRNA can form tight and specific interactions with ribosomal proteins to form ribosomal subunits. These ribosomal proteins contain basic residues (as opposed to acidic residues) and aromatic residues (i.e. phenylalanine, tyrosine
-Tyrosine or tyrosine (symbol Tyr or Y) or 4-hydroxyphenylalanine is one of the 20 standard amino acids that are used by cells to synthesize proteins. It is a non-essential amino acid with a polar side group. The word "tyrosine" is from the G ...
and tryptophan
Tryptophan (symbol Trp or W)
is an α-amino acid that is used in the biosynthesis of proteins. Tryptophan contains an α-amino group, an α- carboxylic acid group, and a side chain indole, making it a polar molecule with a non-polar aromatic ...
) allowing them to form chemical interactions with their associated RNA regions, such as stacking interactions. Ribosomal proteins can also cross-link to the sugar-phosphate backbone of rRNA with binding sites that consist of basic residues (i.e. lysine and arginine). All ribosomal proteins (including the specific sequences that bind to rRNA) have been identified. These interactions along with the association of the small and large ribosomal subunits result in a functioning ribosome capable of synthesizing protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s.
 Ribosomal RNA organizes into two types of major ribosomal subunit: the large subunit (LSU) and the small subunit (SSU). One of each type come together to form a functioning ribosome. The subunits are at times referred to by their size-sedimentation measurements (a number with an "S" suffix). In prokaryotes, the LSU and SSU are called the 50S and 30S subunits, respectively. In eukaryotes, they are a little larger; the LSU and SSU of eukaryotes are termed the 60S and 40S subunits, respectively.
In the ribosomes of prokaryotes such as
Ribosomal RNA organizes into two types of major ribosomal subunit: the large subunit (LSU) and the small subunit (SSU). One of each type come together to form a functioning ribosome. The subunits are at times referred to by their size-sedimentation measurements (a number with an "S" suffix). In prokaryotes, the LSU and SSU are called the 50S and 30S subunits, respectively. In eukaryotes, they are a little larger; the LSU and SSU of eukaryotes are termed the 60S and 40S subunits, respectively.
In the ribosomes of prokaryotes such as bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
, the SSU contains a single small rRNA molecule (~1500 nucleotides) while the LSU contains one single small rRNA and a single large rRNA molecule (~3000 nucleotides). These are combined with ~50 ribosomal proteins to form ribosomal subunits. There are three types of rRNA found in prokaryotic ribosomes: 23S and 5S rRNA in the LSU and 16S rRNA in the SSU.
In the ribosomes of eukaryotes such as human
Humans (''Homo sapiens'') are the most abundant and widespread species of primate, characterized by bipedalism and exceptional cognitive skills due to a large and complex brain. This has enabled the development of advanced tools, culture, ...
s, the SSU contains a single small rRNA (~1800 nucleotides) while the LSU contains two small rRNAs and one molecule of large rRNA (~5000 nucleotides). Eukaryotic rRNA has over 70 ribosomal protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s which interact to form larger and more polymorphic ribosomal units in comparison to prokaryotes. There are four types of rRNA in eukaryotes: 3 species in the LSU and 1 in the SSU. Yeast
Yeasts are eukaryotic, single-celled microorganisms classified as members of the fungus kingdom. The first yeast originated hundreds of millions of years ago, and at least 1,500 species are currently recognized. They are estimated to constitut ...
has been the traditional model for observation of eukaryotic
Eukaryotes () are organisms whose Cell (biology), cells have a cell nucleus, nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the ...
rRNA behavior and processes, leading to a deficit in diversification of research. It has only been within the last decade that technical advances (specifically in the field of Cryo-EM
Cryogenic electron microscopy (cryo-EM) is a cryomicroscopy technique applied on samples cooled to cryogenic temperatures. For biological specimens, the structure is preserved by embedding in an environment of vitreous ice. An aqueous sample so ...
) have allowed for preliminary investigation into ribosomal behavior in other eukaryotes. In yeast
Yeasts are eukaryotic, single-celled microorganisms classified as members of the fungus kingdom. The first yeast originated hundreds of millions of years ago, and at least 1,500 species are currently recognized. They are estimated to constitut ...
, the LSU contains the 5S, 5.8S and 28S rRNAs. The combined 5.8S and 28S are roughly equivalent in size and function to the prokaryotic 23S rRNA subtype, minus expansion segments (ESs) that are localized to the surface of the ribosome which were thought to occur only in eukaryotes. However recently, the Asgard
In Nordic mythology, Asgard (Old Norse: ''Ásgarðr'' ; "enclosure of the Æsir") is a location associated with the gods. It appears in a multitude of Old Norse sagas and mythological texts. It is described as the fortified home of the Æsir ...
phyla, namely, Lokiarchaeota
Lokiarchaeota is a proposed phylum of the Archaea. The phylum includes all members of the group previously named Deep Sea Archaeal Group (DSAG), also known as Marine Benthic Group B (MBG-B). Lokiarchaeota is part of the superphylum Asgard contai ...
and Heimdallarchaeota
Asgard or Asgardarchaeota is a proposed superphylum consisting of a group of archaea that includes Lokiarchaeota, Thorarchaeota, Odinarchaeota, and Heimdallarchaeota. It appears the eukaryotes emerged within the Asgard, in a branch containin ...
, considered the closest archaeal relatives to Eukarya
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bacte ...
, were reported to possess two supersized ESs in their 23S rRNAs. Likewise, the 5S rRNA contains a 108‐nucleotide insertion in the ribosomes of the halophilic archaeon ''Halococcus morrhuae''.
A eukaryotic SSU contains the 18S rRNA subunit, which also contains ESs. SSU ESs are generally smaller than LSU ESs.
SSU and LSU rRNA sequences are widely used for study of evolutionary relationships
Evolution is change in the heritable characteristics of biological populations over successive generations. These characteristics are the expressions of genes, which are passed on from parent to offspring during reproduction. Variation t ...
among organisms, since they are of ancient origin, are found in all known forms of life and are resistant to horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). H ...
. rRNA sequences are conserved (unchanged) over time due to their crucial role in the function of the ribosome. Phylogenic
In biology, phylogenetics (; from Greek φυλή/ φῦλον [] "tribe, clan, race", and wikt:γενετικός, γενετικός [] "origin, source, birth") is the study of the evolutionary history and relationships among or within groups o ...
information derived from the 16s rRNA is currently used as the main method of delineation between similar prokaryotic species by calculating nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules wi ...
similarity. The canonical tree of life is the lineage of the translation system.
LSU rRNA subtypes have been called ribozyme
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozymes demons ...
s because ribosomal proteins cannot bind to the catalytic site of the ribosome in this area (specifically the peptidyl transferase
The peptidyl transferase is an aminoacyltransferase () as well as the primary enzymatic function of the ribosome, which forms peptide bonds between adjacent amino acids using tRNAs during the translation process of protein biosynthesis. The subs ...
center, or PTC).
The SSU rRNA subtypes decode mRNA in its decoding center (DC). Ribosomal proteins cannot enter the DC.
The structure of rRNA is able to drastically change to affect tRNA binding to the ribosome during translation of other mRNAs. In 16S rRNA, this is thought to occur when certain nucleotides in the rRNA appear to alternate base pairing between one nucleotide or another, forming a "switch" that alters the rRNA's conformation. This process is able to affect the structure of the LSU and SSU, suggesting that this conformational switch in the rRNA structure affects the entire ribosome in its ability to match a codon with its anticodon in tRNA selection as well as decode mRNA.
Assembly
Ribosomal RNA's integration and assembly into ribosomes begins with their folding, modification, processing and assembly withribosomal protein
A ribosomal protein (r-protein or rProtein) is any of the proteins that, in conjunction with rRNA, make up the ribosomal subunits involved in the cellular process of translation. ''E. coli'', other bacteria and Archaea have a 30S small subunit an ...
s to form the two ribosomal subunits, the LSU and the SSU. In Prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s, rRNA incorporation occurs in the cytoplasm due to the lack of membrane-bound organelles. In Eukaryotes, however, this process primarily takes place in the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
and is initiated by the synthesis of pre-RNA. This requires the presence of all three RNA polymerases. In fact, the transcription of pre-RNA by RNA polymerase I accounts for about 60% of cell's total cellular RNA transcription. This is followed by the folding of the pre-RNA so that it can be assembled with ribosomal proteins. This folding is catalyzed by endo- and exonucleases, RNA helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separatin ...
s, GTPase
GTPases are a large family of hydrolase enzymes that bind to the nucleotide guanosine triphosphate (GTP) and hydrolyze it to guanosine diphosphate (GDP). The GTP binding and hydrolysis takes place in the highly conserved P-loop "G domain", a pro ...
s and ATPases. The rRNA subsequently undergoes endo- and exonucleolytic processing to remove external
External may refer to:
* External (mathematics), a concept in abstract algebra
* Externality
In economics, an externality or external cost is an indirect cost or benefit to an uninvolved third party that arises as an effect of another party' ...
and internal transcribed spacer
Internal transcribed spacer (ITS) is the spacer DNA situated between the small-subunit ribosomal RNA (rRNA) and large-subunit rRNA genes in the chromosome or the corresponding transcribed region in the polycistronic rRNA precursor transcript.
I ...
s. The pre-RNA then undergoes modifications such as methylation or pseudouridinylation before ribosome assembly factors and ribosomal proteins assemble with the pre-RNA to form pre-ribosomal particles. Upon going under more maturation steps and subsequent exit from the nucleolus into the cytoplasm, these particles combine to form the ribosomes. The basic and aromatic
In chemistry, aromaticity is a chemical property of cyclic ( ring-shaped), ''typically'' planar (flat) molecular structures with pi bonds in resonance (those containing delocalized electrons) that gives increased stability compared to satur ...
residues found within the primary structure of rRNA allow for favorable stacking interactions and attraction to ribosomal proteins, creating a cross-linking effect between the backbone of rRNA and other components of the ribosomal unit. More detail on the initiation and beginning portion of these processes can be found in the "Biosynthesis" section.
Function
 Universally conserved secondary structural elements in rRNA among different species show that these sequences are some of the oldest discovered. They serve critical roles in forming the catalytic sites of translation of mRNA. During translation of mRNA, rRNA functions to bind both mRNA and tRNA to facilitate the process of translating mRNA's codon sequence into amino acids. rRNA initiates the catalysis of protein synthesis when tRNA is sandwiched between the SSU and LSU. In the SSU, the mRNA interacts with the anticodons of the tRNA. In the LSU, the amino acid acceptor stem of the tRNA interacts with the LSU rRNA. The ribosome catalyzes ester-amide exchange, transferring the C-terminus of a nascent peptide from a tRNA to the amine of an amino acid. These processes are able to occur due to sites within the ribosome in which these molecules can bind, formed by the rRNA stem-loops. A ribosome has three of these binding sites called the A, P and E sites:
* In general, the A (aminoacyl) site contains an aminoacyl-tRNA (a
Universally conserved secondary structural elements in rRNA among different species show that these sequences are some of the oldest discovered. They serve critical roles in forming the catalytic sites of translation of mRNA. During translation of mRNA, rRNA functions to bind both mRNA and tRNA to facilitate the process of translating mRNA's codon sequence into amino acids. rRNA initiates the catalysis of protein synthesis when tRNA is sandwiched between the SSU and LSU. In the SSU, the mRNA interacts with the anticodons of the tRNA. In the LSU, the amino acid acceptor stem of the tRNA interacts with the LSU rRNA. The ribosome catalyzes ester-amide exchange, transferring the C-terminus of a nascent peptide from a tRNA to the amine of an amino acid. These processes are able to occur due to sites within the ribosome in which these molecules can bind, formed by the rRNA stem-loops. A ribosome has three of these binding sites called the A, P and E sites:
* In general, the A (aminoacyl) site contains an aminoacyl-tRNA (a tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
esterified to an amino acid on the 3' end).
* The P (peptidyl) site contains a tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
esterified
In chemistry, an ester is a compound derived from an oxoacid (organic or inorganic) in which at least one hydroxyl group () is replaced by an alkoxy group (), as in the substitution reaction of a carboxylic acid and an alcohol. Glycerides ar ...
to the nascent peptide. The free amino (NH2) group of the A site tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
attacks the ester linkage of P site tRNA, causing transfer of the nascent peptide to the amino acid in the A site. This reaction is takes place in the peptidyl transferase center
* The E (exit) site contains a tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
that has been discharged, with a free 3' end (with no amino acid or nascent peptide).
A single mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
can be translated simultaneously by multiple ribosomes. This is called a polysome
A polyribosome (or polysome or ergosome) is a group of ribosomes bound to an mRNA molecule like “beads” on a “thread”. It consists of a complex of an mRNA molecule and two or more ribosomes that act to translate mRNA instructions into pol ...
.
In prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s, much work has been done to further identify the importance of rRNA in translation of mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
. For example, it has been found that the A site consists primarily of 16S rRNA. Apart from various protein elements that interact with tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
at this site, it is hypothesized that if these proteins were removed without altering ribosomal structure, the site would continue to function normally. In the P site, through the observation of crystal structure
In crystallography, crystal structure is a description of the ordered arrangement of atoms, ions or molecules in a crystal, crystalline material. Ordered structures occur from the intrinsic nature of the constituent particles to form symmetric pat ...
s it has been shown the 3' end of 16s rRNA can fold into the site as if a molecule of mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
. This results in intermolecular interactions that stabilize the subunits. Similarly, like the A site, the P site primarily contains rRNA with few protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s. The peptidyl transferase
The peptidyl transferase is an aminoacyltransferase () as well as the primary enzymatic function of the ribosome, which forms peptide bonds between adjacent amino acids using tRNAs during the translation process of protein biosynthesis. The subs ...
center, for example, is formed by nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules wi ...
s from the 23S rRNA subunit. In fact, studies have shown that the peptidyl transferase
The peptidyl transferase is an aminoacyltransferase () as well as the primary enzymatic function of the ribosome, which forms peptide bonds between adjacent amino acids using tRNAs during the translation process of protein biosynthesis. The subs ...
center contains no proteins, and is entirely initiated by the presence of rRNA. Unlike the A and P sites, the E site contains more protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s. Because protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s are not essential for the functioning of the A and P sites, the E site molecular composition shows that it is perhaps evolved later. In primitive ribosomes, it is likely that tRNAs
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino a ...
exited from the P site. Additionally, it has been shown that E-site tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
bind with both the 16S and 23S rRNA subunits.
Subunits and associated ribosomal RNA
Bothprokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
and eukaryotic
Eukaryotes () are organisms whose Cell (biology), cells have a cell nucleus, nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the ...
ribosomes can be broken down into two subunits, one large and one small. The exemplary species used in the table below for their respective rRNAs are the bacterium ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escher ...
'' (prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
) and human ( eukaryote). Note that "nt" represents the length of the rRNA type in nucleotides and the "S" (such as in "16S) represents Svedberg
A Svedberg unit or svedberg (symbol S, sometimes Sv) is a non- SI metric unit for sedimentation coefficients. The Svedberg unit offers a measure of a particle's size indirectly based on its sedimentation rate under acceleration (i.e. how fast a ...
units.
S units of the subunits (or the rRNAs) cannot simply be added because they represent measures of sedimentation rate rather than of mass. The sedimentation rate of each subunit is affected by its shape, as well as by its mass. The nt units can be added as these represent the integer number of units in the linear rRNA polymers (for example, the total length of the human rRNA = 7216 nt).
Gene cluster
A gene family is a set of homologous genes within one organism. A gene cluster is a group of two or more genes found within an organism's DNA that encode similar polypeptides, or proteins, which collectively share a generalized function and are ...
s coding for rRNA are commonly called "ribosomal DNA
Ribosomal DNA (rDNA) is a DNA sequence that codes for ribosomal RNA. These sequences regulate transcription initiation and amplification, and contain both transcribed and non-transcribed spacer segments.
In the human genome there are 5 chromo ...
" or rDNA (note that the term seems to imply that ribosomes contain DNA, which is not the case).
In prokaryotes
Inprokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s a small 30S ribosomal subunit contains the 16S ribosomal RNA. The large 50S ribosomal subunit contains two rRNA species (the 5S and 23S ribosomal RNAs). Therefore it can be deduced that in both bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
and archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaebac ...
there is one rRNA gene that codes for all three rRNA types :16S, 23S and 5S.
Bacterial 16S ribosomal RNA, 23S ribosomal RNA, and 5S rRNA genes are typically organized as a co-transcribed operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
. As shown by the image in this section, there is an internal transcribed spacer
Internal transcribed spacer (ITS) is the spacer DNA situated between the small-subunit ribosomal RNA (rRNA) and large-subunit rRNA genes in the chromosome or the corresponding transcribed region in the polycistronic rRNA precursor transcript.
I ...
between 16S and 23S rRNA gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a ba ...
s. There may be one or more copies of the operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
dispersed in the genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
(for example, ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escher ...
'' has seven). Typically in bacteria there are between one and fifteen copies.
Archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaebac ...
contains either a single rRNA gene operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
or up to four copies of the same operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
.
The 3' end of the 16S ribosomal RNA (in a ribosome) recognizes a sequence on the 5' end of mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
called the Shine-Dalgarno sequence.
In eukaryotes
 In contrast, eukaryotes generally have many copies of the rRNA genes organized in
In contrast, eukaryotes generally have many copies of the rRNA genes organized in tandem repeats
Tandem repeats occur in DNA when a pattern of one or more nucleotides is repeated and the repetitions are directly adjacent to each other. Several protein domains also form tandem repeats within their amino acid primary structure, such as armadil ...
. In humans, approximately 300–400 repeats are present in five clusters, located on chromosomes
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
13 (RNR1
RNA, ribosomal 1, also known as RNR1, is a human gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can ha ...
), 14 ( RNR2), 15 ( RNR3), 21 (RNR4
RNA, ribosomal 4, also known as RNR4, is a human gene.
References
Further reading
* Nucleolus organizer region ]
Nucleolus organizer regions (NORs) are chromosome, chromosomal regions crucial for the formation of the nucleolus. In humans, th ...
) and 22 (RNR5
RNA, ribosomal 5, also known as RNR5, is a human gene. Genes for ribosomal RNA are clustered on the short arms of chromosomes 13, 14, 15, 20, 21. The gene for RNR5 exists in multiple copies on chromosome 22. Each gene cluster contains 30–4 ...
). Diploid humans have 10 clusters of genomic rDNA which in total make up less than 0.5% of the human genome
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the ...
.
It was previously accepted that repeat rDNA sequences were identical and served as redundancies or failsafes to account for natural replication errors and point mutation
A point mutation is a genetic mutation where a single nucleotide base is changed, inserted or deleted from a DNA or RNA sequence of an organism's genome. Point mutations have a variety of effects on the downstream protein product—consequence ...
s. However, sequence variation in rDNA (and subsequently rRNA) in humans across multiple chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
s has been observed, both within and between human individuals. Many of these variations are palindromic sequence
A palindromic sequence is a nucleic acid sequence in a double-stranded DNA or RNA molecule whereby reading in a certain direction (e.g. 5' to 3') on one strand is identical to the sequence in the same direction (e.g. 5' to 3') on the comple ...
s and potential errors due to replication. Certain variants are also expressed in a tissue-specific manner in mice.
Mammalian cells have 2 mitochondrial ( 12S and 16S) rRNA molecules and 4 types of cytoplasmic rRNA (the 28S, 5.8S, 18S, and 5S subunits). The 28S, 5.8S, and 18S rRNAs are encoded by a single transcription unit (45S) separated by 2 internally transcribed spacers. The first spacer corresponds to the one found in bacteria and archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaebac ...
, and the other spacer is an insertion into what was the 23S rRNA in prokaryotes. The 45S rDNA is organized into 5 clusters (each has 30–40 repeats) on chromosomes 13, 14, 15, 21, and 22. These are transcribed by RNA polymerase I
RNA polymerase 1 (also known as Pol I) is, in higher eukaryotes, the polymerase that only transcribes ribosomal RNA (but not 5S rRNA, which is synthesized by RNA polymerase III), a type of RNA that accounts for over 50% of the total RNA synthesize ...
. The DNA for the 5S subunit occurs in tandem arrays (~200–300 true 5S genes and many dispersed pseudogenes), the largest one on the chromosome 1q41-42. 5S rRNA is transcribed by RNA polymerase III
In eukaryote cells, RNA polymerase III (also called Pol III) is a protein that transcribes DNA to synthesize ribosomal 5S rRNA, tRNA and other small RNAs.
The genes transcribed by RNA Pol III fall in the category of "housekeeping" genes whose e ...
. The 18S rRNA in most eukaryotes is in the small ribosomal subunit, and the large subunit contains three rRNA species (the 5S, 5.8S and 28S
28S ribosomal RNA is the structural ribosomal RNA (rRNA) for the LSU rRNA, large subunit (LSU) of eukaryotic cytoplasmic ribosomes, and thus one of the basic components of all eukaryotic cells. It has a size of 25S in plants and 28S in mammals, ...
in mammals, 25S in plants, rRNAs).
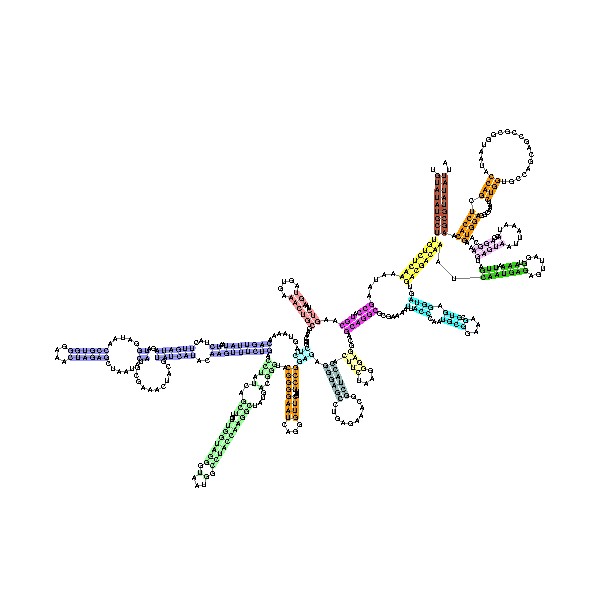
The tertiary structure of the small subunit ribosomal RNA (SSU rRNA) has been resolved by X-ray crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angles ...
. The secondary structure of SSU rRNA contains 4 distinct domains—the 5', central, 3' major and 3' minor domains. A model of the secondary structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta ...
for the 5' domain (500-800 nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules wi ...
s) is shown.
Biosynthesis
In eukaryotes
As the building-blocks for the organelle, production of rRNA is ultimately therate-limiting step
In chemical kinetics, the overall rate of a reaction is often approximately determined by the slowest step, known as the rate-determining step (RDS or RD-step or r/d step) or rate-limiting step. For a given reaction mechanism, the prediction of th ...
in the synthesis of a ribosome. In the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
, rRNA is synthesized by RNA polymerase I
RNA polymerase 1 (also known as Pol I) is, in higher eukaryotes, the polymerase that only transcribes ribosomal RNA (but not 5S rRNA, which is synthesized by RNA polymerase III), a type of RNA that accounts for over 50% of the total RNA synthesize ...
using the specialty genes ( rDNA) that encode for it, which are found repeatedly throughout the genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
. The genes coding for 18S, 28S and 5.8S rRNA are located in the nucleolus organizer region ]
Nucleolus organizer regions (NORs) are chromosome, chromosomal regions crucial for the formation of the nucleolus. In humans, the NORs are located on the short arms of the acrocentric chromosomes 13, 14, 15, 21 and 22, the genes RNR1, RNR2, RNR ...
and are transcribed into large precursor rRNA (pre-rRNA) molecules by RNA polymerase I
RNA polymerase 1 (also known as Pol I) is, in higher eukaryotes, the polymerase that only transcribes ribosomal RNA (but not 5S rRNA, which is synthesized by RNA polymerase III), a type of RNA that accounts for over 50% of the total RNA synthesize ...
. These pre-rRNA molecules are separated by external and internal spacer sequences and then Methylation, methylated, which is key for later assembly and folding
Fold, folding or foldable may refer to:
Arts, entertainment, and media
* ''Fold'' (album), the debut release by Australian rock band Epicure
* Fold (poker), in the game of poker, to discard one's hand and forfeit interest in the current pot
*Abov ...
. After separation and release as individual molecules, assembly proteins bind to each naked rRNA strand and fold it into its functional form using cooperative assembly and progressive addition of more folding proteins as needed. The exact details of how the folding proteins bind to the rRNA and how correct folding is achieved remains unknown. The rRNA complexes are then further processed by reactions involving exo- and endo-nucleolytic cleavages guided by snoRNA (small nucleolar RNAs) in complex with proteins. As these complexes are compacted together to form a cohesive unit, interactions between rRNA and surrounding ribosomal protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s are constantly remodeled throughout assembly in order to provide stability and protect binding site
In biochemistry and molecular biology, a binding site is a region on a macromolecule such as a protein that binds to another molecule with specificity. The binding partner of the macromolecule is often referred to as a ligand. Ligands may includ ...
s. This process is referred to as the "maturation" phase of the rRNA lifecycle. The modifications that occur during maturation of rRNA have been found to contribute directly to control of gene expression by providing physical regulation of translational access of tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
and mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
. Some studies have found that extensive methylation of various rRNA types is also necessary during this time to maintain ribosome stability.
The genes for 5S rRNA are located inside the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
and are transcribed into pre-5S rRNA by RNA polymerase III
In eukaryote cells, RNA polymerase III (also called Pol III) is a protein that transcribes DNA to synthesize ribosomal 5S rRNA, tRNA and other small RNAs.
The genes transcribed by RNA Pol III fall in the category of "housekeeping" genes whose e ...
. The pre-5S rRNA enters the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
for processing and assembly with 28S and 5.8S rRNA to form the LSU. 18S rRNA forms the SSUs by combining with numerous ribosomal protein
A ribosomal protein (r-protein or rProtein) is any of the proteins that, in conjunction with rRNA, make up the ribosomal subunits involved in the cellular process of translation. ''E. coli'', other bacteria and Archaea have a 30S small subunit an ...
s. Once both subunits are assembled, they are individually exported into the cytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
to form the 80S unit and begin initiation of translation
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transla ...
of mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
.
Ribosomal RNA is non-coding
Non-coding DNA (ncDNA) sequences are components of an organism's DNA that do not encode protein sequences. Some non-coding DNA is transcribed into functional non-coding RNA molecules (e.g. transfer RNA, microRNA, piRNA, ribosomal RNA, and regula ...
and is never translated into protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s of any kind: rRNA is only transcribed from rDNA and then matured for use as a structural building block for ribosomes. Transcribed rRNA is bound to ribosomal protein
A ribosomal protein (r-protein or rProtein) is any of the proteins that, in conjunction with rRNA, make up the ribosomal subunits involved in the cellular process of translation. ''E. coli'', other bacteria and Archaea have a 30S small subunit an ...
s to form the subunits of ribosomes and acts as the physical structure that pushes mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
and tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
through the ribosome to process and translate them.
Eukaryotic regulation
Synthesis of rRNA is up-regulated and down-regulated to maintainhomeostasis
In biology, homeostasis (British also homoeostasis) (/hɒmɪə(ʊ)ˈsteɪsɪs/) is the state of steady internal, physical, and chemical conditions maintained by living systems. This is the condition of optimal functioning for the organism and ...
by a variety of processes and interactions:
* The kinase AKT
Protein kinase B (PKB), also known as Akt, is the collective name of a set of three serine/threonine-specific protein kinases that play key roles in multiple cellular processes such as glucose metabolism, apoptosis, cell proliferation, tran ...
indirectly promotes synthesis of rRNA as RNA
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. RNA and deoxyribonucleic acid ( DNA) are nucleic acids. Along with lipids, proteins, and carbohydra ...
polymerase I is AKT-dependent.
* Certain angiogenic ribonuclease
Ribonuclease (commonly abbreviated RNase) is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within ...
s, such as angiogenin
Angiogenin (ANG) also known as ribonuclease 5 is a small 123 amino acid protein that in humans is encoded by the ''ANG'' gene. Angiogenin is a potent stimulator of new blood vessels through the process of angiogenesis. Ang hydrolyzes cellular ...
(ANG), can translocate and accumulate in the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
. When the concentration of ANG becomes too high, some studies have found that ANG can bind to the promoter region of rDNA and unnecessarily increase rRNA transcription. This can be damaging to the nucleolus and can even lead to unchecked transcription and cancer
Cancer is a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. These contrast with benign tumors, which do not spread. Possible signs and symptoms include a lump, abnormal b ...
.
* During times of cellular glucose restriction, AMP-activated protein kinase
5' AMP-activated protein kinase or AMPK or 5' adenosine monophosphate-activated protein kinase is an enzyme (EC 2.7.11.31) that plays a role in cellular energy homeostasis, largely to activate glucose and fatty acid uptake and oxidation when cell ...
(AMPK) discourages metabolic processes that consume energy but are non-essential. As a result, it is capable of phosphorylating RNA polymerase I
RNA polymerase 1 (also known as Pol I) is, in higher eukaryotes, the polymerase that only transcribes ribosomal RNA (but not 5S rRNA, which is synthesized by RNA polymerase III), a type of RNA that accounts for over 50% of the total RNA synthesize ...
(at the Ser-635 site) in order to down-regulate rRNA synthesis by disrupting transcription initiation
Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules called ...
.
* Impairment or removal of more than one pseudouridine
Pseudouridine (abbreviated by the Greek letter psi- Ψ) is an isomer of the nucleoside uridine in which the uracil is attached via a carbon-carbon instead of a nitrogen-carbon glycosidic bond. (In this configuration, uracil is sometimes referred ...
or 29-O-methylation regions from the ribosome decoding center significantly reduces rate of rRNA transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
by reducing the rate of incorporation of new amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha a ...
s.
* Formation of heterochromatin is essential to silencing rRNA transcription, without which ribosomal RNA is synthesized unchecked and greatly decreases the lifespan of the organism.
In prokaryotes
Similar to eukaryotes, the production of rRNA is therate-limiting step
In chemical kinetics, the overall rate of a reaction is often approximately determined by the slowest step, known as the rate-determining step (RDS or RD-step or r/d step) or rate-limiting step. For a given reaction mechanism, the prediction of th ...
in the prokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
synthesis of a ribosome. In '' E. coli,'' it has been found that rRNA is transcribed from the two promoters P1 and P2 found within seven different ''rrn'' operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
s. The P1 promoter is specifically responsible for regulating rRNA synthesis during moderate to high bacterial growth rates. Because the transcriptional activity of this promoter is directly proportional to the growth rate, it is primarily responsible for rRNA regulation
Regulation is the management of complex systems according to a set of rules and trends. In systems theory, these types of rules exist in various fields of biology and society, but the term has slightly different meanings according to context. Fo ...
. An increased rRNA concentration serves as a negative feedback mechanism to ribosome synthesis. High NTP concentration has been found to be required for efficient transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of the ''rrn'' P1 promoters. They are thought to form stabilizing complexes with RNA polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template.
Using the enzyme helicase, RNAP locally opens the ...
and the promoters. In bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
specifically, this association of high NTP concentration with increased rRNA synthesis provides a molecular explanation as to why ribosomal and thus protein synthesis is dependent on growth-rate. A low growth-rate yields lower rRNA / ribosomal synthesis rates while a higher growth rate yields a higher rRNA / ribosomal synthesis rate. This allows a cell to save energy or increase its metabolic activity
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of life-sustaining chemical reactions in organisms. The three main functions of metabolism are: the conversion of the energy in food to energy available to run cell ...
dependent on its needs and available resources.
In prokaryotic cells
A prokaryote () is a Unicellular organism, single-celled organism that lacks a cell nucleus, nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek language, Greek wikt:πρό#Ancient Greek, πρό (, 'before') a ...
, each rRNA gene or operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
is transcribed into a single RNA precursor that includes 16S, 23S, 5S rRNA and tRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino ac ...
sequences along with transcribed spacers. The RNA processing then begins before the transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
is complete. During processing reactions, the rRNAs and tRNAs
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino a ...
are released as separate molecules.
Prokaryotic regulation
Because of the vital role rRNA plays in thecell physiology
Cell physiology is the biological study of the activities that take place in a cell to keep it alive. The term ''physiology'' refers to normal functions in a living organism. Animal cells, plant cells and microorganism cells show similarities in ...
of prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s, there is much overlap in rRNA regulation
Regulation is the management of complex systems according to a set of rules and trends. In systems theory, these types of rules exist in various fields of biology and society, but the term has slightly different meanings according to context. Fo ...
mechanisms. At the transcriptional level, there are both positive and negative effectors of rRNA transcription that facilitate a cell's maintenance of homeostasis
In biology, homeostasis (British also homoeostasis) (/hɒmɪə(ʊ)ˈsteɪsɪs/) is the state of steady internal, physical, and chemical conditions maintained by living systems. This is the condition of optimal functioning for the organism and ...
:
* An UP element upstream of the ''rrn'' P1 promoter can bind a subunit of RNA polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template.
Using the enzyme helicase, RNAP locally opens the ...
, thus promoting transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of rRNA.
* Transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
factors such as FIS bind upstream of the promoter and interact with RNA polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template.
Using the enzyme helicase, RNAP locally opens the ...
which facilitates transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
.
* Anti-termination factors bind downstream of the ''rrn'' P2 promoter, preventing premature transcription termination.
* Due to the stringent response
The stringent response, also called stringent control, is a stress response of bacteria and plant chloroplasts in reaction to amino-acid starvation, fatty acid limitation, iron limitation, heat shock and other stress conditions. The stringent resp ...
, when the availability of amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha a ...
s is low, ppGpp (a negative effector) can inhibit transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
from both the P1 and P2 promoters.
Degradation
Ribosomal RNA is quite stable in comparison to other common types of RNA and persists for longer periods of time in a healthy cellular environment. Once assembled into functional units, ribosomal RNA within ribosomes are stable in the stationary phase of the cell life cycle for many hours. Degradation can be triggered via "stalling" of a ribosome, a state that occurs when the ribosome recognizes faulty mRNA or encounters other processing difficulties that causes translation by the ribosome to cease. Once a ribosome stalls, a specialized pathway on the ribosome is initiated to target the entire complex for disassembly.In eukaryotes
As with anyprotein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
or RNA
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. RNA and deoxyribonucleic acid ( DNA) are nucleic acids. Along with lipids, proteins, and carbohydra ...
, rRNA production is prone to errors resulting in the production of non-functional rRNA. To correct this, the cell allows for degradation of rRNA through the non-functional rRNA decay (NRD) pathway. Much of the research in this topic was conducted on eukaryotic cells, specifically ''Saccharomyces cerevisiae
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have b ...
'' yeast. Currently, only a basic understanding of how cells are able to target functionally defective ribosomes for ubiquination and degradation in eukaryotes is available.
* The NRD pathway for the 40S subunit may be independent or separate from the NRD pathway for the 60S subunit. It has been observed that certain gene
In biology, the word gene (from , ; "...Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a ba ...
s were able to affect degradation of certain pre-RNAs, but not others.
* Numerous protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
s are involved in the NRD pathway, such as Mms1p and Rtt101p, which are believed to complex together to target ribosomes for degradation. Mms1p and Rtt101p are found to bind together and Rtt101p is believed to recruit a ubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Fo ...
E3 ligase
In biochemistry, a ligase is an enzyme that can catalyze the joining (ligation) of two large molecules by forming a new chemical bond. This is typically via hydrolysis of a small pendant chemical group on one of the larger molecules or the enzym ...
complex, allowing for the non-functional ribosomes to be ubiquinated before being degraded.
** Prokaryotes lack a homolog
In biology, homology is similarity due to shared ancestry between a pair of structures or genes in different taxa. A common example of homologous structures is the forelimbs of vertebrates, where the wings of bats and birds, the arms of prima ...
for Mms1, so it is unclear how prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s are able to degrade non-functional rRNAs.
* The growth rate of eukaryotic cells
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bact ...
did not seem to be significantly affected by the accumulation of non-functional rRNAs.
In prokaryotes
Although there is far less research available on ribosomal RNA degradation inprokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s in comparison to eukaryotes, there has still been interest on whether bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
follow a similar degradation scheme in comparison to the NRD in eukaryotes. Much of the research done for prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s has been conducted on ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escher ...
''. Many differences were found between eukaryotic and prokaryotic rRNA degradation, leading researchers to believe that the two degrade using different pathways.
* Certain mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA replication, DNA or viral repl ...
s in rRNA that were able to trigger rRNA degradation in eukaryotes were unable to do so in prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
s.
* Point mutations in a 23S rRNA would cause both 23S and 16S rRNAs to be degraded, in comparison to eukaryotes, in which mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA replication, DNA or viral repl ...
s in one subunit would only cause that subunit to be degraded.
* Researchers found that removal of a whole helix structure (H69) from the 23S rRNA did not trigger its degradation. This led them to believe that H69 was critical for endonucleases to recognize and degrade the mutated rRNA.
Sequence conservation and stability
Due to the prevalent and unwavering nature of rRNA across all organisms, the study of its resistance to gene transfer,mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA replication, DNA or viral repl ...
, and alteration without destruction of the organism has become a popular field of interest. Ribosomal RNA genes have been found to be tolerant to modification and incursion. When rRNA sequencing ''is'' altered, cells have been found to become compromised and quickly cease normal function. These key traits of rRNA have become especially important for gene database projects (comprehensive online resources such as SILVA or SINA) where alignment of ribosomal RNA sequences from across the different biologic domains greatly eases "Taxonomy (biology), taxonomic assignment, Phylogenetics, phylogenetic analysis and the investigation of microbial diversity."
Examples of resilience:
* Addition of large, nonsensical RNA fragments into many parts of the 16S rRNA unit does not observably alter the function of the Ribosome, ribosomal unit as a whole.
* Non-coding RNARD7 has the capability to alter processing of rRNA to make the molecules resistant to degradation by carboxylic acid. This is a crucial mechanism in maintaining rRNA concentrations during active growth when acid build-up (due to the substrate phosphorylation required to produce Adenosine triphosphate, ATP) can become toxic to intracellular functions.
* Insertion of hammerhead ribozymes that are capable of cis-cleavages along 16S rRNA greatly inhibit function and diminish stability.
* While most cellular functions degrade heavily after only short period of exposure to Hypoxic tissue, hypoxic environments, rRNA remains un-degraded and resolved after six days of prolonged hypoxia. Only after such an extended period of time do rRNA intermediates (indicative of degradation finally occurring) begin to present themselves.
Significance
 Ribosomal RNA characteristics are important in evolution, thus taxonomy and medicine.
* rRNA is one of only a few gene products present in all cells. For this reason, genes that encode the rRNA ( rDNA) are sequenced to identify an organism's Taxonomy (biology), taxonomic group, calculate related groups, and estimate rates of Genetic divergence, species divergence. As a result, many thousands of rRNA sequences are known and stored in specialized databases such as RDP-II and SILVA.
* Alterations to rRNA are what allow certain disease-causing
Ribosomal RNA characteristics are important in evolution, thus taxonomy and medicine.
* rRNA is one of only a few gene products present in all cells. For this reason, genes that encode the rRNA ( rDNA) are sequenced to identify an organism's Taxonomy (biology), taxonomic group, calculate related groups, and estimate rates of Genetic divergence, species divergence. As a result, many thousands of rRNA sequences are known and stored in specialized databases such as RDP-II and SILVA.
* Alterations to rRNA are what allow certain disease-causing bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among ...
, such as ''Mycobacterium tuberculosis'' (the bacterium that causes tuberculosis) to develop extreme drug resistance. Due to similar issues, this has become a prevalent problem in veterinary medicine where the main method for handling bacterial infection in pets is administration of drugs that attack the Peptidyl transferase, peptidyl-transferase centre (PTC) of the Bacterial ribosomes, bacterial ribosome. Mutations in 23S rRNA have created perfect resistance to these drugs as they operate together in an unknown fashion to bypass the PTC enitrely.
* rRNA is the target of numerous clinically relevant antibiotics: chloramphenicol, erythromycin, kasugamycin, Thiopeptide, micrococcin, paromomycin, ricin, alpha-sarcin, spectinomycin, streptomycin, and thiostrepton.
* rRNA have been shown to be the origin of species-specific microRNAs, like Mir-663 microRNA precursor family, miR-663 in humans and MicroRNA#miRNA-712, miR-712 in mice. These particular MicroRNA, miRNAs originate from the internal transcribed spacer
Internal transcribed spacer (ITS) is the spacer DNA situated between the small-subunit ribosomal RNA (rRNA) and large-subunit rRNA genes in the chromosome or the corresponding transcribed region in the polycistronic rRNA precursor transcript.
I ...
s of the rRNA.
Human genes
* 45S:RNR1
RNA, ribosomal 1, also known as RNR1, is a human gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can ha ...
, RNR2, RNR3, RNR4
RNA, ribosomal 4, also known as RNR4, is a human gene.
References
Further reading
* Nucleolus organizer region ]
Nucleolus organizer regions (NORs) are chromosome, chromosomal regions crucial for the formation of the nucleolus. In humans, th ...
, RNR5
RNA, ribosomal 5, also known as RNR5, is a human gene. Genes for ribosomal RNA are clustered on the short arms of chromosomes 13, 14, 15, 20, 21. The gene for RNR5 exists in multiple copies on chromosome 22. Each gene cluster contains 30–4 ...
; (unclustered) RNA18SN1, RNA18SN2, RNA18SN3, RNA18SN4, RNA18SN5, RNA28SN1, RNA28SN2, RNA28SN3, RNA28SN4, RNA28SN5, RNA45SN1, RNA45SN2, RNA45SN3, RNA45SN4, RNA45SN5, RNA5-8SN1, RNA5-8SN2, RNA5-8SN3, RNA5-8SN4, RNA5-8SN5
* 5S: RNA5S1, RNA5S2, RNA5S3, RNA5S4, RNA5S5, RNA5S6, RNA5S7, RNA5S8, RNA5S9, RNA5S10, RNA5S11, RNA5S12, RNA5S13, RNA5S14, RNA5S15, RNA5S16, RNA5S17
* Mt: MT-RNR1, MT-TV (mitochondrial), MT-TV (co-opted), MT-RNR2
See also
* Ribotyping * Diazaborine B, a maturation inhibitor of rRNAs for the large ribosomal subunitReferences
External links
16S rRNA, BioMineWiki
Ribosomal Database Project II
*
SILVA rRNA Database Project
(also includes Eukaryotes (18S) and LSU (23S/28S))
''Halococcus morrhuae'' (archaebacterium) 5S rRNA
{{Ribosome subunits Ribosomal RNA, Protein biosynthesis RNA Non-coding RNA Ribozymes