Protein structure on:
[Wikipedia]
[Google]
[Amazon]
Protein structure is the three-dimensional arrangement of atoms in an


 Proteins are frequently described as consisting of several structural units. These units include domains, motifs, and folds. Despite the fact that there are about 100,000 different proteins expressed in
Proteins are frequently described as consisting of several structural units. These units include domains, motifs, and folds. Despite the fact that there are about 100,000 different proteins expressed in
 Proteins are often thought of as relatively stable tertiary structures that experience conformational changes after being affected by interactions with other proteins or as a part of enzymatic activity. However, proteins may have varying degrees of stability, and some of the less stable variants are intrinsically disordered proteins. These proteins exist and function in a relatively 'disordered' state lacking a stable
Proteins are often thought of as relatively stable tertiary structures that experience conformational changes after being affected by interactions with other proteins or as a part of enzymatic activity. However, proteins may have varying degrees of stability, and some of the less stable variants are intrinsically disordered proteins. These proteins exist and function in a relatively 'disordered' state lacking a stable
Protein Ensemble Database
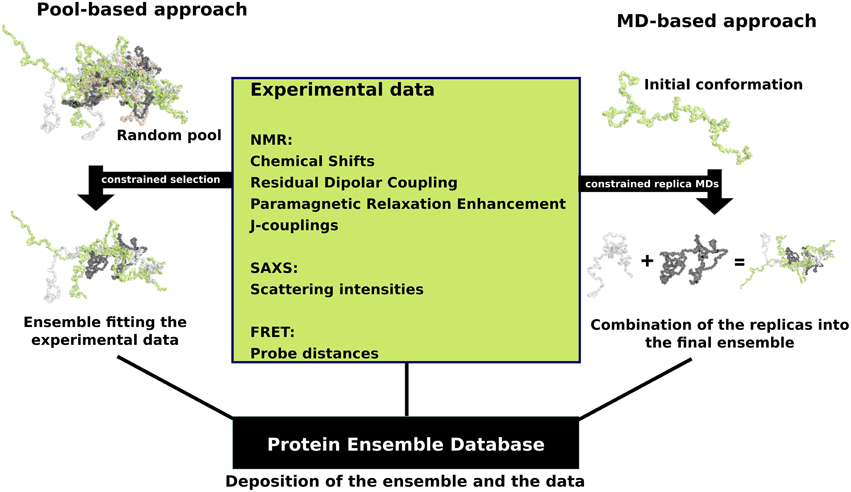
that fall into two general methodologies – pool and molecular dynamics (MD) approaches (diagrammed in the figure). The pool based approach uses the protein’s amino acid sequence to create a massive pool of random conformations. This pool is then subjected to more computational processing that creates a set of theoretical parameters for each conformation based on the structure. Conformational subsets from this pool whose average theoretical parameters closely match known experimental data for this protein are selected. The alternative molecular dynamics approach takes multiple random conformations at a time and subjects all of them to experimental data. Here the experimental data is serving as limitations to be placed on the conformations (e.g. known distances between atoms). Only conformations that manage to remain within the limits set by the experimental data are accepted. This approach often applies large amounts of experimental data to the conformations which is a very computationally demanding task. The conformational ensembles were generated for a number of highly dynamic and partially unfolded proteins, such as Sic1/ Cdc4, p15 PAF, MKK7, Beta-synuclein and P27
 Around 90% of the protein structures available in the
Around 90% of the protein structures available in the
50 Years of Protein Structure Determination Timeline - HTML Version - National Institute of General Medical Sciences
at NIH
amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha ...
-chain molecule
A molecule is a group of two or more atoms held together by attractive forces known as chemical bonds; depending on context, the term may or may not include ions which satisfy this criterion. In quantum physics, organic chemistry, and bio ...
. Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respon ...
s are polymer
A polymer (; Greek ''poly-'', "many" + '' -mer'', "part")
is a substance or material consisting of very large molecules called macromolecules, composed of many repeating subunits. Due to their broad spectrum of properties, both synthetic and ...
s specifically polypeptide
Peptides (, ) are short chains of amino acids linked by peptide bonds. Long chains of amino acids are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides ...
s formed from sequences of amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha ...
s, the monomers of the polymer. A single amino acid monomer may also be called a ''residue'' indicating a repeating unit of a polymer. Proteins form by amino acids undergoing condensation reactions, in which the amino acids lose one water molecule per reaction in order to attach to one another with a peptide bond
In organic chemistry, a peptide bond is an amide type of covalent chemical bond linking two consecutive alpha-amino acids from C1 (carbon number one) of one alpha-amino acid and N2 ( nitrogen number two) of another, along a peptide or protein c ...
. By convention, a chain under 30 amino acids is often identified as a peptide
Peptides (, ) are short chains of amino acids linked by peptide bonds. Long chains of amino acids are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides. ...
, rather than a protein. To be able to perform their biological function, proteins fold into one or more specific spatial conformations driven by a number of non-covalent interactions such as hydrogen bonding
In chemistry, a hydrogen bond (or H-bond) is a primarily electrostatic force of attraction between a hydrogen (H) atom which is covalently bound to a more electronegative "donor" atom or group (Dn), and another electronegative atom bearing ...
, ionic interactions, Van der Waals forces
In molecular physics, the van der Waals force is a distance-dependent interaction between atoms or molecules. Unlike ionic or covalent bonds, these attractions do not result from a chemical electronic bond; they are comparatively weak and t ...
, and hydrophobic
In chemistry, hydrophobicity is the physical property of a molecule that is seemingly repelled from a mass of water (known as a hydrophobe). In contrast, hydrophiles are attracted to water.
Hydrophobic molecules tend to be nonpolar and, ...
packing. To understand the functions of proteins at a molecular level, it is often necessary to determine their three-dimensional structure. This is the topic of the scientific field of structural biology
Structural biology is a field that is many centuries old which, and as defined by the Journal of Structural Biology, deals with structural analysis of living material (formed, composed of, and/or maintained and refined by living cells) at every le ...
, which employs techniques such as X-ray crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angle ...
, NMR spectroscopy, cryo electron microscopy (cryo-EM) and dual polarisation interferometry to determine the structure of proteins.
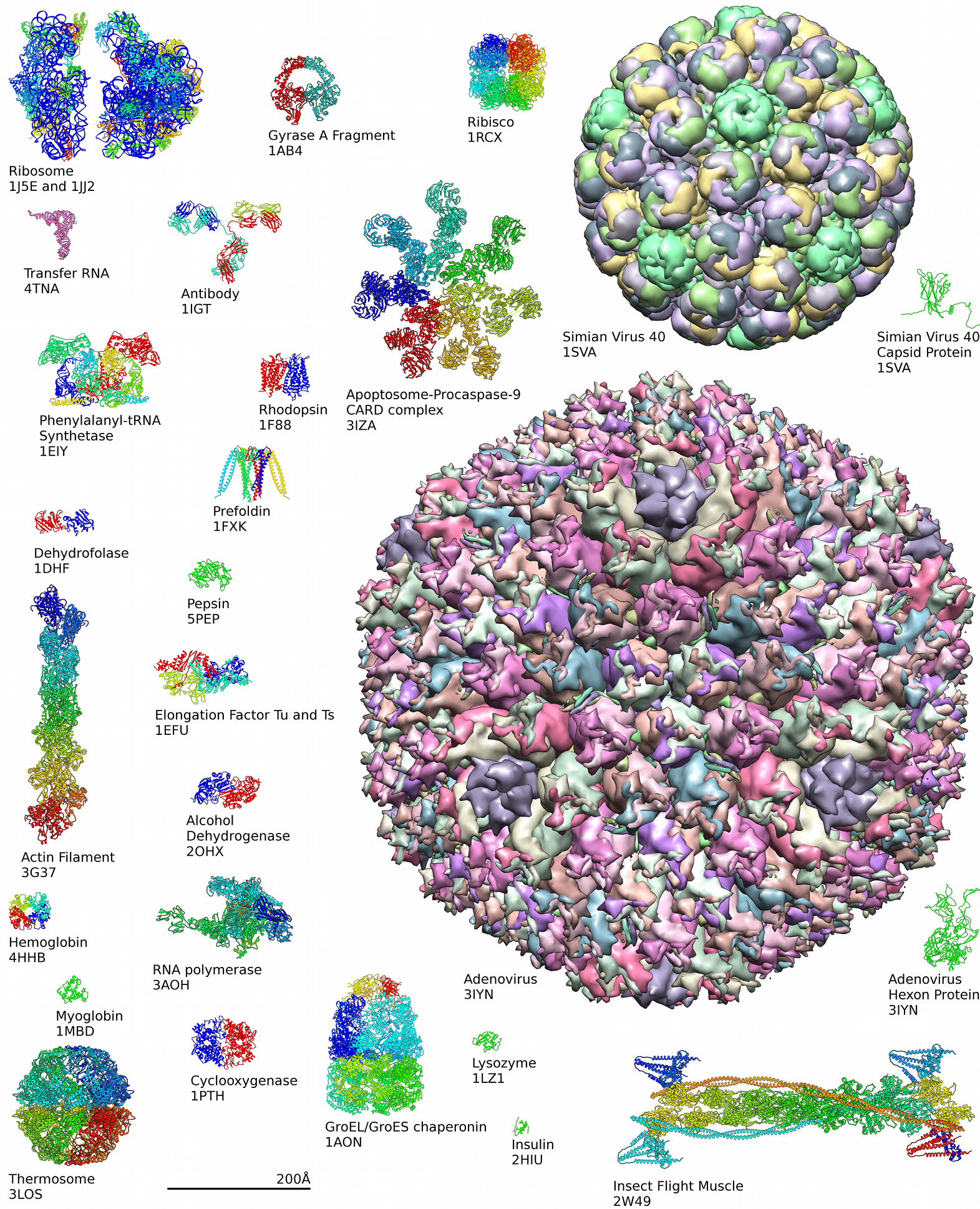
Protein structures range in size from tens to several thousand amino acids. By physical size, proteins are classified as nanoparticle
A nanoparticle or ultrafine particle is usually defined as a particle of matter that is between 1 and 100 nanometres (nm) in diameter. The term is sometimes used for larger particles, up to 500 nm, or fibers and tubes that are less than 10 ...
s, between 1–100 nm. Very large protein complexes can be formed from protein subunit
In structural biology, a protein subunit is a polypeptide chain or single protein molecule that assembles (or "''coassembles''") with others to form a protein complex.
Large assemblies of proteins such as viruses often use a small number of ...
s. For example, many thousands of actin
Actin is a protein family, family of Globular protein, globular multi-functional proteins that form microfilaments in the cytoskeleton, and the thin filaments in myofibril, muscle fibrils. It is found in essentially all Eukaryote, eukaryotic cel ...
molecules assemble into a microfilament.
A protein usually undergoes reversible structural changes in performing its biological function. The alternative structures of the same protein are referred to as different conformations, and transitions between them are called conformational change
In biochemistry, a conformational change is a change in the shape of a macromolecule, often induced by environmental factors.
A macromolecule is usually flexible and dynamic. Its shape can change in response to changes in its environment or oth ...
s.
Levels of protein structure
There are four distinct levels of protein structure.
Primary structure
Theprimary structure
Protein primary structure is the linear sequence of amino acids in a peptide or protein. By convention, the primary structure of a protein is reported starting from the amino-terminal (N) end to the carboxyl-terminal (C) end. Protein biosynthes ...
of a protein refers to the sequence of amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha ...
s in the polypeptide chain. The primary structure is held together by peptide bonds that are made during the process of protein biosynthesis
Protein biosynthesis (or protein synthesis) is a core biological process, occurring inside cells, balancing the loss of cellular proteins (via degradation or export) through the production of new proteins. Proteins perform a number of critical ...
. The two ends of the polypeptide chain
Peptides (, ) are short chains of amino acids linked by peptide bonds. Long chains of amino acids are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides ...
are referred to as the carboxyl terminus (C-terminus) and the amino terminus (N-terminus) based on the nature of the free group on each extremity. Counting of residues always starts at the N-terminal end (NH2-group), which is the end where the amino group is not involved in a peptide bond. The primary structure of a protein is determined by the gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
corresponding to the protein. A specific sequence of nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecul ...
s in DNA is transcribed into mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
, which is read by the ribosome in a process called translation
Translation is the communication of the Meaning (linguistic), meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The ...
. The sequence of amino acids in insulin was discovered by Frederick Sanger
Frederick Sanger (; 13 August 1918 – 19 November 2013) was an English biochemist who received the Nobel Prize in Chemistry twice.
He won the 1958 Chemistry Prize for determining the amino acid sequence of insulin and numerous other pr ...
, establishing that proteins have defining amino acid sequences. The sequence of a protein is unique to that protein, and defines the structure and function of the protein. The sequence of a protein can be determined by methods such as Edman degradation or tandem mass spectrometry. Often, however, it is read directly from the sequence of the gene using the genetic code
The genetic code is the set of rules used by living cells to translate information encoded within genetic material ( DNA or RNA sequences of nucleotide triplets, or codons) into proteins. Translation is accomplished by the ribosome, which links ...
. It is strictly recommended to use the words "amino acid residues" when discussing proteins because when a peptide bond is formed, a water molecule is lost, and therefore proteins are made up of amino acid residues. Post-translational modification
Post-translational modification (PTM) is the covalent and generally enzymatic modification of proteins following protein biosynthesis. This process occurs in the endoplasmic reticulum and the golgi apparatus. Proteins are synthesized by ribos ...
s such as phosphorylation
In chemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology and could be driven by natural selection. Text was copied from this source, ...
s and glycosylation
Glycosylation is the reaction in which a carbohydrate (or 'glycan'), i.e. a glycosyl donor, is attached to a hydroxyl or other functional group of another molecule (a glycosyl acceptor) in order to form a glycoconjugate. In biology (but not ...
s are usually also considered a part of the primary structure, and cannot be read from the gene. For example, insulin
Insulin (, from Latin ''insula'', 'island') is a peptide hormone produced by beta cells of the pancreatic islets encoded in humans by the ''INS'' gene. It is considered to be the main anabolic hormone of the body. It regulates the metabol ...
is composed of 51 amino acids in 2 chains. One chain has 31 amino acids, and the other has 20 amino acids.
Secondary structure

Secondary structure
Protein secondary structure is the three dimensional form of ''local segments'' of proteins. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary struct ...
refers to highly regular local sub-structures on the actual polypeptide backbone chain. Two main types of secondary structure, the α-helix and the β-strand or β-sheets, were suggested in 1951 by Linus Pauling
Linus Carl Pauling (; February 28, 1901August 19, 1994) was an American chemist, biochemist, chemical engineer, peace activist, author, and educator. He published more than 1,200 papers and books, of which about 850 dealt with scientific top ...
et al. These secondary structures are defined by patterns of hydrogen bonds between the main-chain peptide groups. They have a regular geometry, being constrained to specific values of the dihedral angles ψ and φ on the Ramachandran plot. Both the α-helix and the β-sheet represent a way of saturating all the hydrogen bond donors and acceptors in the peptide backbone. Some parts of the protein are ordered but do not form any regular structures. They should not be confused with random coil, an unfolded polypeptide chain lacking any fixed three-dimensional structure. Several sequential secondary structures may form a " supersecondary unit".
Tertiary structure
Tertiary structure
Protein tertiary structure is the three dimensional shape of a protein. The tertiary structure will have a single polypeptide chain "backbone" with one or more protein secondary structures, the protein domains. Amino acid side chains may int ...
refers to the three-dimensional structure created by a single protein molecule (a single polypeptide chain
Peptides (, ) are short chains of amino acids linked by peptide bonds. Long chains of amino acids are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides ...
). It may include one or several domains. The α-helices and β-pleated-sheets are folded into a compact globular structure
In biochemistry, globular proteins or spheroproteins are spherical ("globe-like") proteins and are one of the common protein types (the others being fibrous, disordered and membrane proteins). Globular proteins are somewhat water-soluble (for ...
. The folding is driven by the ''non-specific'' hydrophobic interactions, the burial of hydrophobic residues from water
Water (chemical formula ) is an inorganic, transparent, tasteless, odorless, and nearly colorless chemical substance, which is the main constituent of Earth's hydrosphere and the fluids of all known living organisms (in which it acts as ...
, but the structure is stable only when the parts of a protein domain
In molecular biology, a protein domain is a region of a protein's polypeptide chain that is self-stabilizing and that folds independently from the rest. Each domain forms a compact folded three-dimensional structure. Many proteins consist o ...
are locked into place by ''specific'' tertiary interactions, such as salt bridges, hydrogen bonds, and the tight packing of side chains and disulfide bond
In biochemistry, a disulfide (or disulphide in British English) refers to a functional group with the structure . The linkage is also called an SS-bond or sometimes a disulfide bridge and is usually derived by the coupling of two thiol groups ...
s. The disulfide bonds are extremely rare in cytosolic proteins, since the cytosol
The cytosol, also known as cytoplasmic matrix or groundplasm, is one of the liquids found inside cells ( intracellular fluid (ICF)). It is separated into compartments by membranes. For example, the mitochondrial matrix separates the mitochondri ...
(intracellular fluid) is generally a reducing environment.
Quaternary structure
Quaternary structure is the three-dimensional structure consisting of the aggregation of two or more individual polypeptide chains (subunits) that operate as a single functional unit ( multimer). The resulting multimer is stabilized by the same non-covalent interactions and disulfide bonds as in tertiary structure. There are many possible quaternary structure organisations. Complexes of two or more polypeptides (i.e. multiple subunits) are called multimers. Specifically it would be called a dimer if it contains two subunits, a trimer if it contains three subunits, a tetramer if it contains four subunits, and a pentamer if it contains five subunits. The subunits are frequently related to one another by symmetry operations, such as a 2-fold axis in a dimer. Multimers made up of identical subunits are referred to with a prefix of "homo-" and those made up of different subunits are referred to with a prefix of "hetero-", for example, a heterotetramer, such as the two alpha and two beta chains ofhemoglobin
Hemoglobin (haemoglobin BrE) (from the Greek word αἷμα, ''haîma'' 'blood' + Latin ''globus'' 'ball, sphere' + ''-in'') (), abbreviated Hb or Hgb, is the iron-containing oxygen-transport metalloprotein present in red blood cells (erythrocyte ...
.
Domains, motifs, and folds in protein structure
 Proteins are frequently described as consisting of several structural units. These units include domains, motifs, and folds. Despite the fact that there are about 100,000 different proteins expressed in
Proteins are frequently described as consisting of several structural units. These units include domains, motifs, and folds. Despite the fact that there are about 100,000 different proteins expressed in eukaryotic
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bact ...
systems, there are many fewer different domains, structural motifs and folds.
Structural domain
Astructural domain
In molecular biology, a protein domain is a region of a protein's polypeptide chain that is self-stabilizing and that folds independently from the rest. Each domain forms a compact folded three-dimensional structure. Many proteins consist of se ...
is an element of the protein's overall structure that is self-stabilizing and often folds
Benjamin Scott Folds (born September 12, 1966) is an American singer-songwriter, musician, and composer, who is the first artistic advisor to the National Symphony Orchestra at the John F. Kennedy Center for the Performing Arts, Kennedy Center in ...
independently of the rest of the protein chain. Many domains are not unique to the protein products of one gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
or one gene family but instead appear in a variety of proteins. Domains often are named and singled out because they figure prominently in the biological function of the protein they belong to; for example, the "calcium
Calcium is a chemical element with the symbol Ca and atomic number 20. As an alkaline earth metal, calcium is a reactive metal that forms a dark oxide-nitride layer when exposed to air. Its physical and chemical properties are most similar t ...
-binding domain of calmodulin". Because they are independently stable, domains can be "swapped" by genetic engineering
Genetic engineering, also called genetic modification or genetic manipulation, is the modification and manipulation of an organism's genes using technology. It is a set of technologies used to change the genetic makeup of cells, including ...
between one protein and another to make chimera proteins. A conservative combination of several domains that occur in different proteins, such as protein tyrosine phosphatase domain and C2 domain pair, was called "a superdomain" that may evolve as a single unit.
Structural and sequence motifs
Thestructural
A structure is an arrangement and organization of interrelated elements in a material object or system, or the object or system so organized. Material structures include man-made objects such as buildings and machines and natural objects such as ...
and sequence motifs refer to short segments of protein three-dimensional structure or amino acid sequence that were found in a large number of different proteins
Supersecondary structure
Tertiary protein structures can have multiple secondary elements on the same polypeptide chain. The supersecondary structure refers to a specific combination ofsecondary structure
Protein secondary structure is the three dimensional form of ''local segments'' of proteins. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary struct ...
elements, such as β-α-β units or a helix-turn-helix
Helix-turn-helix is a DNA-binding protein (DBP). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove o ...
motif. Some of them may be also referred to as structural motifs.
Protein fold
A protein fold refers to the general protein architecture, like ahelix bundle A helix bundle is a small protein fold composed of several alpha helices that are usually nearly parallel or antiparallel to each other.
Three-helix bundles
Three-helix bundles are among the smallest and fastest known cooperatively folding struct ...
, β-barrel
In protein structures, a beta barrel is a beta sheet composed of tandem repeats that twists and coils to form a closed toroidal structure in which the first strand is bonded to the last strand (hydrogen bond). Beta-strands in many beta-barrels are ...
, Rossmann fold or different "folds" provided in the Structural Classification of Proteins database. A related concept is protein topology.
Protein dynamics and conformational ensembles
Proteins are not static objects, but rather populate ensembles of conformational states. Transitions between these states typically occur on nanoscales, and have been linked to functionally relevant phenomena such as allosteric signaling andenzyme catalysis
Enzyme catalysis is the increase in the rate of a process by a biological molecule, an "enzyme". Most enzymes are proteins, and most such processes are chemical reactions. Within the enzyme, generally catalysis occurs at a localized site, called ...
. Protein dynamics Proteins are generally thought to adopt unique structures determined by their amino acid sequences. However, proteins are not strictly static objects, but rather populate ensembles of (sometimes similar) conformations. Transitions between these stat ...
and conformational change
In biochemistry, a conformational change is a change in the shape of a macromolecule, often induced by environmental factors.
A macromolecule is usually flexible and dynamic. Its shape can change in response to changes in its environment or oth ...
s allow proteins to function as nanoscale biological machines within cells, often in the form of multi-protein complexes. Examples include motor proteins, such as myosin, which is responsible for muscle
Skeletal muscles (commonly referred to as muscles) are Organ (biology), organs of the vertebrate muscular system and typically are attached by tendons to bones of a skeleton. The muscle cells of skeletal muscles are much longer than in the other ...
contraction, kinesin, which moves cargo inside cells away from the nucleus along microtubules
Microtubules are polymers of tubulin that form part of the cytoskeleton and provide structure and shape to eukaryotic cells. Microtubules can be as long as 50 micrometres, as wide as 23 to 27 nm and have an inner diameter between 11 a ...
, and dynein, which moves cargo inside cells towards the nucleus and produces the axonemal beating of motile cilia and flagella. " effect, the otile ciliumis a nanomachine composed of perhaps over 600 proteins in molecular complexes, many of which also function independently as nanomachines... Flexible linkers allow the mobile protein domains connected by them to recruit their binding partners and induce long-range allostery
In biochemistry, allosteric regulation (or allosteric control) is the regulation of an enzyme by binding an effector molecule at a site other than the enzyme's active site.
The site to which the effector binds is termed the ''allosteric sit ...
via protein domain dynamics. "
 Proteins are often thought of as relatively stable tertiary structures that experience conformational changes after being affected by interactions with other proteins or as a part of enzymatic activity. However, proteins may have varying degrees of stability, and some of the less stable variants are intrinsically disordered proteins. These proteins exist and function in a relatively 'disordered' state lacking a stable
Proteins are often thought of as relatively stable tertiary structures that experience conformational changes after being affected by interactions with other proteins or as a part of enzymatic activity. However, proteins may have varying degrees of stability, and some of the less stable variants are intrinsically disordered proteins. These proteins exist and function in a relatively 'disordered' state lacking a stable tertiary structure
Protein tertiary structure is the three dimensional shape of a protein. The tertiary structure will have a single polypeptide chain "backbone" with one or more protein secondary structures, the protein domains. Amino acid side chains may int ...
. As a result, they are difficult to describe by a single fixed tertiary structure
Protein tertiary structure is the three dimensional shape of a protein. The tertiary structure will have a single polypeptide chain "backbone" with one or more protein secondary structures, the protein domains. Amino acid side chains may int ...
. Conformational ensembles
In computational chemistry, conformational ensembles, also known as structural ensembles, are experimentally constrained computational models describing the structure of intrinsically unstructured proteins. Such proteins are flexible in nature, ...
have been devised as a way to provide a more accurate and 'dynamic' representation of the conformational state of intrinsically disordered proteins.
Protein ensemble files are a representation of a protein that can be considered to have a flexible structure. Creating these files requires determining which of the various theoretically possible protein conformations actually exist. One approach is to apply computational algorithms to the protein data in order to try to determine the most likely set of conformations for an ensemble file. There are multiple methods for preparing data for thProtein Ensemble Database
that fall into two general methodologies – pool and molecular dynamics (MD) approaches (diagrammed in the figure). The pool based approach uses the protein’s amino acid sequence to create a massive pool of random conformations. This pool is then subjected to more computational processing that creates a set of theoretical parameters for each conformation based on the structure. Conformational subsets from this pool whose average theoretical parameters closely match known experimental data for this protein are selected. The alternative molecular dynamics approach takes multiple random conformations at a time and subjects all of them to experimental data. Here the experimental data is serving as limitations to be placed on the conformations (e.g. known distances between atoms). Only conformations that manage to remain within the limits set by the experimental data are accepted. This approach often applies large amounts of experimental data to the conformations which is a very computationally demanding task. The conformational ensembles were generated for a number of highly dynamic and partially unfolded proteins, such as Sic1/ Cdc4, p15 PAF, MKK7, Beta-synuclein and P27
Protein folding
As it is translated, polypeptides exit the ribosome mostly as a random coil and folds into its native state. The final structure of the protein chain is generally assumed to be determined by its amino acid sequence ( Anfinsen's dogma).Protein stability
Thermodynamic stability of proteins represents the free energy difference between the folded and unfolded protein states. This free energy difference is very sensitive to temperature, hence a change in temperature may result in unfolding or denaturation. Protein denaturation may result in loss of function, and loss of native state. The free energy of stabilization of soluble globular proteins typically does not exceed 50 kJ/mol. Taking into consideration the large number of hydrogen bonds that take place for the stabilization of secondary structures, and the stabilization of the inner core through hydrophobic interactions, the free energy of stabilization emerges as small difference between large numbers.Protein structure determination

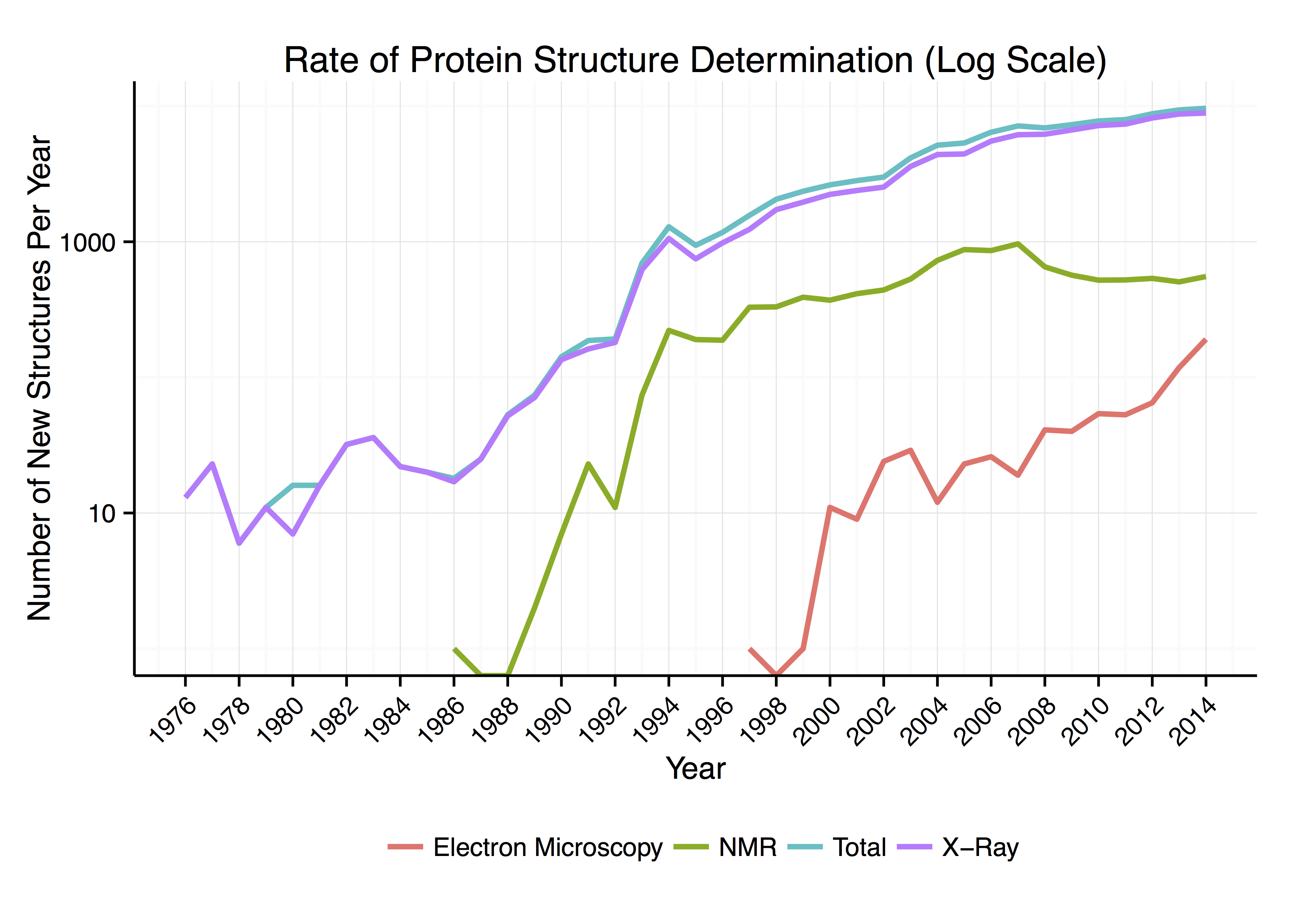
 Around 90% of the protein structures available in the
Around 90% of the protein structures available in the Protein Data Bank
The Protein Data Bank (PDB) is a database for the three-dimensional structural data of large biological molecules, such as proteins and nucleic acids. The data, typically obtained by X-ray crystallography, NMR spectroscopy, or, increasingly, c ...
have been determined by X-ray crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angle ...
. This method allows one to measure the three-dimensional (3-D) density distribution of electron
The electron (, or in nuclear reactions) is a subatomic particle with a negative one elementary electric charge. Electrons belong to the first generation of the lepton particle family,
and are generally thought to be elementary partic ...
s in the protein, in the crystallized state, and thereby infer
Inferences are steps in reasoning, moving from premises to logical consequences; etymologically, the word '' infer'' means to "carry forward". Inference is theoretically traditionally divided into deduction and induction, a distinction that ...
the 3-D coordinates of all the atom
Every atom is composed of a nucleus and one or more electrons bound to the nucleus. The nucleus is made of one or more protons and a number of neutrons. Only the most common variety of hydrogen has no neutrons.
Every solid, liquid, gas ...
s to be determined to a certain resolution. Roughly 7% of the known protein structures have been obtained by nuclear magnetic resonance
Nuclear magnetic resonance (NMR) is a physical phenomenon in which nuclei in a strong constant magnetic field are perturbed by a weak oscillating magnetic field (in the near field) and respond by producing an electromagnetic signal with a ...
(NMR) techniques. For larger protein complexes, cryo-electron microscopy can determine protein structures. The resolution is typically lower than that of X-ray crystallography, or NMR, but the maximum resolution is steadily increasing. This technique is still a particularly valuable for very large protein complexes such as virus coat protein
A capsid is the protein shell of a virus, enclosing its genetic material. It consists of several oligomeric (repeating) structural subunits made of protein called protomers. The observable 3-dimensional morphological subunits, which may or may ...
s and amyloid fibers.
General secondary structure composition can be determined via circular dichroism. Vibrational spectroscopy
Infrared spectroscopy (IR spectroscopy or vibrational spectroscopy) is the measurement of the interaction of infrared radiation with matter by absorption, emission, or reflection. It is used to study and identify chemical substances or functi ...
can also be used to characterize the conformation of peptides, polypeptides, and proteins. Two-dimensional infrared spectroscopy has become a valuable method to investigate the structures of flexible peptides and proteins that cannot be studied with other methods. A more qualitative picture of protein structure is often obtained by proteolysis
Proteolysis is the breakdown of proteins into smaller polypeptides or amino acids. Uncatalysed, the hydrolysis of peptide bonds is extremely slow, taking hundreds of years. Proteolysis is typically catalysed by cellular enzymes called protease ...
, which is also useful to screen for more crystallizable protein samples. Novel implementations of this approach, including fast parallel proteolysis (FASTpp)
Fast parallel proteolysis (FASTpp) is a method to determine the thermostability of proteins by measuring which fraction of protein resists rapid proteolytic digestion.
History and background
Proteolysis is widely used in biochemistry and cell b ...
, can probe the structured fraction and its stability without the need for purification. Once a protein's structure has been experimentally determined, further detailed studies can be done computationally, using molecular dynamic simulations of that structure.
Protein structure databases
A protein structure database is a database that is modeled around the various experimentally determined protein structures. The aim of most protein structure databases is to organize and annotate the protein structures, providing the biological community access to the experimental data in a useful way. Data included in protein structure databases often includes 3D coordinates as well as experimental information, such as unit cell dimensions and angles forx-ray crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angle ...
determined structures. Though most instances, in this case either proteins or a specific structure determinations of a protein, also contain sequence information and some databases even provide means for performing sequence based queries, the primary attribute of a structure database is structural information, whereas sequence databases focus on sequence information, and contain no structural information for the majority of entries. Protein structure databases are critical for many efforts in computational biology
Computational biology refers to the use of data analysis, mathematical modeling and computational simulations to understand biological systems and relationships. An intersection of computer science, biology, and big data, the field also has fo ...
such as structure based drug design, both in developing the computational methods used and in providing a large experimental dataset used by some methods to provide insights about the function of a protein.
Structural classifications of proteins
Protein structures can be grouped based on their structural similarity, topological class or a commonevolution
Evolution is change in the heritable characteristics of biological populations over successive generations. These characteristics are the expressions of genes, which are passed on from parent to offspring during reproduction. Variation ...
ary origin. The Structural Classification of Proteins database and CATH
The CATH Protein Structure Classification database is a free, publicly available online resource that provides information on the evolutionary relationships of protein domains. It was created in the mid-1990s by Professor Christine Orengo and coll ...
database provide two different structural classifications of proteins. When the structural similarity is large the two proteins have possibly diverged from a common ancestor, and shared structure between proteins is considered evidence of homology. Structure similarity can then be used to group proteins together into protein superfamilies. If shared structure is significant but the fraction shared is small, the fragment shared may be the consequence of a more dramatic evolutionary event such as horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offspring ( reproduction). ...
, and joining proteins sharing these fragments into protein superfamilies is no longer justified. Topology of a protein can be used to classify proteins as well. Knot theory and circuit topology are two topology frameworks developed for classification of protein folds based on chain crossing and intrachain contacts respectively.
Computational prediction of protein structure
The generation of a protein sequence is much easier than the determination of a protein structure. However, the structure of a protein gives much more insight in the function of the protein than its sequence. Therefore, a number of methods for the computational prediction of protein structure from its sequence have been developed. ''Ab initio'' prediction methods use just the sequence of the protein. Threading andhomology modeling
Homology modeling, also known as comparative modeling of protein, refers to constructing an atomic-resolution model of the "''target''" protein from its amino acid sequence and an experimental three-dimensional structure of a related homologous p ...
methods can build a 3-D model for a protein of unknown structure from experimental structures of evolutionarily-related proteins, called a protein family.
See also
* Biomolecular structure * Gene structure * Nucleic acid structure * PCRPi-DB *Ribbon diagram
Ribbon diagrams, also known as Richardson diagrams, are 3D schematic representations of protein structure and are one of the most common methods of protein depiction used today. The ribbon shows the overall path and organization of the protein b ...
3D schematic representation of proteins
References
Further reading
50 Years of Protein Structure Determination Timeline - HTML Version - National Institute of General Medical Sciences
at NIH
External links
* {{DEFAULTSORT:Protein Structure