Multiple Sequence Alignment on:
[Wikipedia]
[Google]
[Amazon]
 Multiple sequence alignment (MSA) is the process or the result of
Multiple sequence alignment (MSA) is the process or the result of
 A
A
 Most multiple sequence alignment methods try to minimize the number of insertions/deletions (gaps) and, as a consequence, produce compact alignments. This causes several problems if the sequences to be aligned contain non- homologous regions, if gaps are informative in a
Most multiple sequence alignment methods try to minimize the number of insertions/deletions (gaps) and, as a consequence, produce compact alignments. This causes several problems if the sequences to be aligned contain non- homologous regions, if gaps are informative in a
 Motif finding, also known as profile analysis, is a method of locating
Motif finding, also known as profile analysis, is a method of locating
ExPASy sequence alignment tools
– from the Virtual School of Natural Sciences
– from Pôle Bioinformatique Lyonnais
An entry point to clustal servers and information
An entry point to the main T-Coffee servers
An entry point to the main MergeAlign server and information
*European Bioinformatics Institute servers: *
ClustalW2
– general purpose multiple sequence alignment program for DNA or proteins. *
Muscle
– MUltiple Sequence Comparison by Log-Expectation *
T-coffee
– multiple sequence alignment. *
MAFFT
– Multiple Alignment using Fast Fourier Transform *
KALIGN
– a fast and accurate multiple sequence alignment algorithm.
Multiple sequence alignment lectures
– from the Max Planck Institute for Molecular Genetics
on multiple sequence alignments at the
Molecular Bioinformatics Lecture Notes
{{DEFAULTSORT:Multiple Sequence Alignment Bioinformatics Computational phylogenetics Markov models NP-complete problems
 Multiple sequence alignment (MSA) is the process or the result of
Multiple sequence alignment (MSA) is the process or the result of sequence alignment
In bioinformatics, a sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity that may be a consequence of functional, structural biology, structural, or evolutionary relationships between ...
of three or more biological sequences, generally protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metab ...
, DNA
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of al ...
, or RNA
Ribonucleic acid (RNA) is a polymeric molecule that is essential for most biological functions, either by performing the function itself (non-coding RNA) or by forming a template for the production of proteins (messenger RNA). RNA and deoxyrib ...
. These alignments are used to infer evolutionary
Evolution is the change in the heritable characteristics of biological populations over successive generations. It occurs when evolutionary processes such as natural selection and genetic drift act on genetic variation, resulting in certa ...
relationships via phylogenetic
In biology, phylogenetics () is the study of the evolutionary history of life using observable characteristics of organisms (or genes), which is known as phylogenetic inference. It infers the relationship among organisms based on empirical dat ...
analysis and can highlight homologous features between sequences. Alignments highlight mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, ...
events such as point mutations (single amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 a ...
or nucleotide
Nucleotides are Organic compound, organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both o ...
changes), insertion mutations and deletion mutations, and alignments are used to assess sequence conservation and infer the presence and activity of protein domain
In molecular biology, a protein domain is a region of a protein's Peptide, polypeptide chain that is self-stabilizing and that Protein folding, folds independently from the rest. Each domain forms a compact folded Protein tertiary structure, thre ...
s, tertiary structure
Protein tertiary structure is the three-dimensional shape of a protein. The tertiary structure will have a single polypeptide chain "backbone" with one or more protein secondary structures, the protein domains. Amino acid side chains and the ...
s, secondary structure
Protein secondary structure is the local spatial conformation of the polypeptide backbone excluding the side chains. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta ...
s, and individual amino acids or nucleotides.
Multiple sequence alignments require more sophisticated methodologies than pairwise alignments, as they are more computationally complex. Most multiple sequence alignment programs use heuristic
A heuristic or heuristic technique (''problem solving'', '' mental shortcut'', ''rule of thumb'') is any approach to problem solving that employs a pragmatic method that is not fully optimized, perfected, or rationalized, but is nevertheless ...
methods rather than global optimization
Global optimization is a branch of operations research, applied mathematics, and numerical analysis that attempts to find the global minimum or maximum of a function or a set of functions on a given set. It is usually described as a minimization ...
because identifying the optimal alignment between more than a few sequences of moderate length is prohibitively computationally expensive. However, heuristic methods generally cannot guarantee high-quality solutions and have been shown to fail to yield near-optimal solutions on benchmark test cases.
Problem statement
Given sequences , similar to the form below:A multiple sequence alignment is taken of this set of sequences by inserting any amount of gaps needed into each of the sequences of until the modified sequences, , all conform to length and no values in the sequences of of the same column consists of only gaps. The mathematical form of an MSA of the above sequence set is shown below:
To return from each particular sequence to , remove all gaps.
Graphing approach
A general approach when calculating multiple sequence alignments is to use graphs to identify all of the different alignments. When finding alignments via graph, a ''complete alignment'' is created in a weighted graph that contains a set of vertices and a set of edges. Each of the graph edges has a weight based on a certain heuristic that helps to score each ''alignment'' or subset of the original graph.Tracing alignments
When determining the best suited alignments for each MSA, a ''trace'' is usually generated. A trace is a set of ''realized'', or corresponding and aligned, vertices that has a specific weight based on the edges that are selected between corresponding vertices. When choosing traces for a set of sequences it is necessary to choose a trace with a maximum weight to get the best alignment of the sequences.Alignment methods
There are various alignment methods used within multiple sequence to maximize scores and correctness of alignments. Each is usually based on a certain heuristic with an insight into the evolutionary process. Most try to replicate evolution to get the most realistic alignment possible to best predict relations between sequences.Dynamic programming
A direct method for producing an MSA uses the dynamic programming technique to identify the globally optimal alignment solution. For proteins, this method usually involves two sets of parameters: agap penalty A Gap penalty is a method of scoring alignments of two or more sequences. When aligning sequences, introducing gaps in the sequences can allow an alignment algorithm to match more terms than a gap-less alignment can. However, minimizing gaps in an a ...
and a substitution matrix
In bioinformatics and evolutionary biology, a substitution matrix describes the frequency at which a character in a Nucleic acid sequence, nucleotide sequence or a Protein primary structure, protein sequence changes to other character states ove ...
assigning scores or probabilities to the alignment of each possible pair of amino acids based on the similarity of the amino acids' chemical properties and the evolutionary probability of the mutation. For nucleotide sequences, a similar gap penalty is used, but a much simpler substitution matrix, wherein only identical matches and mismatches are considered, is typical. The scores in the substitution matrix may be either all positive or a mix of positive and negative in the case of a global alignment, but must be both positive and negative, in the case of a local alignment.
For ''n'' individual sequences, the naive method requires constructing the ''n''-dimensional equivalent of the matrix formed in standard pairwise sequence alignment
In bioinformatics, a sequence alignment is a way of arranging the sequences of DNA, RNA, or protein to identify regions of similarity that may be a consequence of functional, structural biology, structural, or evolutionary relationships between ...
. The search space thus increases exponentially with increasing ''n'' and is also strongly dependent on sequence length. Expressed with the big O notation
Big ''O'' notation is a mathematical notation that describes the asymptotic analysis, limiting behavior of a function (mathematics), function when the Argument of a function, argument tends towards a particular value or infinity. Big O is a memb ...
commonly used to measure computational complexity
In computer science, the computational complexity or simply complexity of an algorithm is the amount of resources required to run it. Particular focus is given to computation time (generally measured by the number of needed elementary operations ...
, a naïve
Naivety (also spelled naïvety), naiveness, or naïveté is the state of being naive. It refers to an apparent or actual lack of experience and sophistication, often describing a neglect of pragmatism in favor of moral idealism. A ''naïve'' may ...
MSA takes ''O(LengthNseqs)'' time to produce. To find the global optimum for ''n'' sequences this way has been shown to be an NP-complete
In computational complexity theory, NP-complete problems are the hardest of the problems to which ''solutions'' can be verified ''quickly''.
Somewhat more precisely, a problem is NP-complete when:
# It is a decision problem, meaning that for any ...
problem. In 1989, based on Carrillo-Lipman Algorithm, Altschul introduced a practical method that uses pairwise alignments to constrain the n-dimensional search space. In this approach pairwise dynamic programming alignments are performed on each pair of sequences in the query set, and only the space near the n-dimensional intersection of these alignments is searched for the n-way alignment. The MSA program optimizes the sum of all of the pairs of characters at each position in the alignment (the so-called ''sum of pair'' score) and has been implemented in a software program for constructing multiple sequence alignments. In 2019, Hosseininasab and van Hoeve showed that by using decision diagrams, MSA may be modeled in polynomial space complexity.
Progressive alignment construction
The most widely used approach to multiple sequence alignments uses a heuristic search known as progressive technique (also known as the hierarchical or tree method) developed by Da-Fei Feng and Doolittle in 1987. Progressive alignment builds up a final MSA by combining pairwise alignments beginning with the most similar pair and progressing to the most distantly related. All progressive alignment methods require two stages: a first stage in which the relationships between the sequences are represented as aphylogenetic tree
A phylogenetic tree or phylogeny is a graphical representation which shows the evolutionary history between a set of species or taxa during a specific time.Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA. In ...
, called a ''guide tree'', and a second step in which the MSA is built by adding the sequences sequentially to the growing MSA according to the guide tree. The initial ''guide tree'' is determined by an efficient clustering method such as neighbor-joining or ''unweighted pair group method with arithmetic mean'' (UPGMA
UPGMA (unweighted pair group method with arithmetic mean) is a simple agglomerative (bottom-up) hierarchical clustering method. It also has a weighted variant, WPGMA, and they are generally attributed to Sokal and Michener.
Note that the unwei ...
), and may use distances based on the number of identical two-letter sub-sequences (as in FASTA rather than a dynamic programming alignment).
Progressive alignments are not guaranteed to be globally optimal. The primary problem is that when errors are made at any stage in growing the MSA, these errors are then propagated through to the final result. Performance is also particularly bad when all of the sequences in the set are rather distantly related. Most modern progressive methods modify their scoring function with a secondary weighting function that assigns scaling factors to individual members of the query set in a nonlinear fashion based on their phylogenetic distance from their nearest neighbors. This corrects for non-random selection of the sequences given to the alignment program.
Progressive alignment methods are efficient enough to implement on a large scale for many (100s to 1000s) sequences. A popular progressive alignment method has been the Clustal family. ClustalW is used extensively for phylogenetic tree construction, in spite of the author's explicit warnings that unedited alignments should not be used in such studies and as input for protein structure prediction
Protein structure prediction is the inference of the three-dimensional structure of a protein from its amino acid sequence—that is, the prediction of its Protein secondary structure, secondary and Protein tertiary structure, tertiary structure ...
by homology modeling. European Bioinformatics Institute
The European Bioinformatics Institute (EMBL-EBI) is an intergovernmental organization (IGO) which, as part of the European Molecular Biology Laboratory (EMBL) family, focuses on research and services in bioinformatics. It is located on the Wel ...
(EMBL-EBI) announced that CLustalW2 will expire in August 2015. They recommend Clustal Omega which performs based on seeded guide trees and HMM profile-profile techniques for protein alignments. An alternative tool for progressive DNA alignments is ''multiple alignment using fast Fourier transform'' ( MAFFT).
Another common progressive alignment method named T-Coffee is slower than Clustal and its derivatives but generally produces more accurate alignments for distantly related sequence sets. T-Coffee calculates pairwise alignments by combining the direct alignment of the pair with indirect alignments that aligns each sequence of the pair to a third sequence. It uses the output from Clustal as well as another local alignment program LALIGN, which finds multiple regions of local alignment between two sequences. The resulting alignment and phylogenetic tree are used as a guide to produce new and more accurate weighting factors.
Because progressive methods are heuristics that are not guaranteed to converge to a global optimum, alignment quality can be difficult to evaluate and their true biological significance can be obscure. A semi-progressive method that improves alignment quality and does not use a lossy heuristic while running in polynomial time
In theoretical computer science, the time complexity is the computational complexity that describes the amount of computer time it takes to run an algorithm. Time complexity is commonly estimated by counting the number of elementary operations p ...
has been implemented in the program PSAlign.
Iterative methods
A set of methods to produce MSAs while reducing the errors inherent in progressive methods are classified as "iterative" because they work similarly to progressive methods but repeatedly realign the initial sequences as well as adding new sequences to the growing MSA. One reason progressive methods are so strongly dependent on a high-quality initial alignment is the fact that these alignments are always incorporated into the final result – that is, once a sequence has been aligned into the MSA, its alignment is not considered further. This approximation improves efficiency at the cost of accuracy. By contrast, iterative methods can return to previously calculated pairwise alignments or sub-MSAs incorporating subsets of the query sequence as a means of optimizing a generalobjective function
In mathematical optimization and decision theory, a loss function or cost function (sometimes also called an error function) is a function that maps an event or values of one or more variables onto a real number intuitively representing some "cost ...
such as finding a high-quality alignment score.
A variety of subtly different iteration methods have been implemented and made available in software packages; reviews and comparisons have been useful but generally refrain from choosing a "best" technique. The software package PRRN/PRRP uses a hill-climbing algorithm to optimize its MSA alignment score and iteratively corrects both alignment weights and locally divergent or "gappy" regions of the growing MSA.Mount DM. (2004). Bioinformatics: Sequence and Genome Analysis 2nd ed. Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY. PRRP performs best when refining an alignment previously constructed by a faster method.
Another iterative program, DIALIGN, takes an unusual approach of focusing narrowly on local alignments between sub-segments or sequence motif
In biology, a sequence motif is a nucleotide or amino-acid sequence pattern that is widespread and usually assumed to be related to biological function of the macromolecule. For example, an ''N''-glycosylation site motif can be defined as ''A ...
s without introducing a gap penalty. The alignment of individual motifs is then achieved with a matrix representation similar to a dot-matrix plot in a pairwise alignment. An alternative method that uses fast local alignments as anchor points or ''seeds'' for a slower global-alignment procedure is implemented in the CHAOS/DIALIGN suite.
A third popular iteration-based method named MUSCLE
Muscle is a soft tissue, one of the four basic types of animal tissue. There are three types of muscle tissue in vertebrates: skeletal muscle, cardiac muscle, and smooth muscle. Muscle tissue gives skeletal muscles the ability to muscle contra ...
(multiple sequence alignment by log-expectation) improves on progressive methods with a more accurate distance measure to assess the relatedness of two sequences. The distance measure is updated between iteration stages (although, in its original form, MUSCLE contained only 2-3 iterations depending on whether refinement was enabled).
Consensus methods
Consensus methods attempt to find the optimal multiple sequence alignment given multiple different alignments of the same set of sequences. There are two commonly used consensus methods, M-COFFEE and MergeAlign. M-COFFEE uses multiple sequence alignments generated by seven different methods to generate consensus alignments. MergeAlign is capable of generating consensus alignments from any number of input alignments generated using different models of sequence evolution or different methods of multiple sequence alignment. The default option for MergeAlign is to infer a consensus alignment using alignments generated using 91 different models of protein sequence evolution.Hidden Markov models
 A
A hidden Markov model
A hidden Markov model (HMM) is a Markov model in which the observations are dependent on a latent (or ''hidden'') Markov process (referred to as X). An HMM requires that there be an observable process Y whose outcomes depend on the outcomes of X ...
(HMM) is a probabilistic model that can assign likelihoods to all possible combinations of gaps, matches, and mismatches, to determine the most likely MSA or set of possible MSAs. HMMs can produce a single highest-scoring output but can also generate a family of possible alignments that can then be evaluated for biological significance. HMMs can produce both global and local alignments. Although HMM-based methods have been developed relatively recently, they offer significant improvements in computational speed, especially for sequences that contain overlapping regions.
Typical HMM-based methods work by representing an MSA as a form of directed acyclic graph
In mathematics, particularly graph theory, and computer science, a directed acyclic graph (DAG) is a directed graph with no directed cycles. That is, it consists of vertices and edges (also called ''arcs''), with each edge directed from one ...
known as a partial-order graph, which consists of a series of nodes representing possible entries in the columns of an MSA. In this representation a column that is absolutely conserved (that is, that all the sequences in the MSA share a particular character at a particular position) is coded as a single node with as many outgoing connections as there are possible characters in the next column of the alignment. In the terms of a typical hidden Markov model, the observed states are the individual alignment columns and the "hidden" states represent the presumed ancestral sequence from which the sequences in the query set are hypothesized to have descended. An efficient search variant of the dynamic programming method, named the Viterbi algorithm, is generally used to successively align the growing MSA to the next sequence in the query set to produce a new MSA. This is distinct from progressive alignment methods because the alignment of prior sequences is updated at each new sequence addition. However, like progressive methods, this technique can be influenced by the order in which the sequences in the query set are integrated into the alignment, especially when the sequences are distantly related.
Several software programs are available in which variants of HMM-based methods have been implemented and which are noted for their scalability and efficiency, although properly using an HMM method is more complex than using more common progressive methods. The simplest is Partial-Order Alignment (POA), and a similar more general method is implemented in the Sequence Alignment and Modeling System (SAM) software package.Hughey R, Krogh A. SAM: Sequence alignment and modeling software system. Technical Report UCSC-CRL-96-22, University of California, Santa Cruz, CA, September 1996. and HMMER.Durbin R, Eddy S, Krogh A, Mitchison G. (1998). Biological sequence analysis: probabilistic models of proteins and nucleic acids, Cambridge University Press, 1998.
SAM has been used as a source of alignments for protein structure prediction
Protein structure prediction is the inference of the three-dimensional structure of a protein from its amino acid sequence—that is, the prediction of its Protein secondary structure, secondary and Protein tertiary structure, tertiary structure ...
to participate in the Critical Assessment of Structure Prediction (CASP
Critical Assessment of Structure Prediction (CASP), sometimes called Critical Assessment of Protein Structure Prediction, is a community-wide, worldwide experiment for protein structure prediction taking place every two years since 1994. CASP pro ...
) structure prediction experiment and to develop a database of predicted proteins in the yeast
Yeasts are eukaryotic, single-celled microorganisms classified as members of the fungus kingdom (biology), kingdom. The first yeast originated hundreds of millions of years ago, and at least 1,500 species are currently recognized. They are est ...
species '' S. cerevisiae''. HHsearch is a software package for the detection of remotely related protein sequences based on the pairwise comparison of HMMs. A server running HHsearch ( HHpred) was the fastest of 10 automatic structure prediction servers in the CASP7 and CASP8 structure prediction competitions.
Phylogeny-aware methods
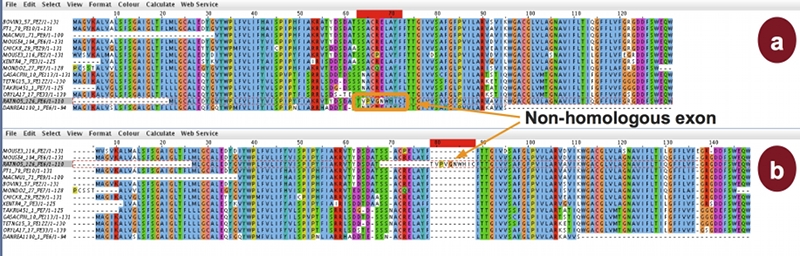
 Most multiple sequence alignment methods try to minimize the number of insertions/deletions (gaps) and, as a consequence, produce compact alignments. This causes several problems if the sequences to be aligned contain non- homologous regions, if gaps are informative in a
Most multiple sequence alignment methods try to minimize the number of insertions/deletions (gaps) and, as a consequence, produce compact alignments. This causes several problems if the sequences to be aligned contain non- homologous regions, if gaps are informative in a phylogeny
A phylogenetic tree or phylogeny is a graphical representation which shows the evolutionary history between a set of species or Taxon, taxa during a specific time.Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, M ...
analysis. These problems are common in newly produced sequences that are poorly annotated and may contain frame-shifts, wrong domains or non-homologous spliced exon
An exon is any part of a gene that will form a part of the final mature RNA produced by that gene after introns have been removed by RNA splicing. The term ''exon'' refers to both the DNA sequence within a gene and to the corresponding sequence ...
s. The first such method was developed in 2005 by Löytynoja and Goldman. The same authors released a software package called ''PRANK'' in 2008. PRANK improves alignments when insertions are present. Nevertheless, it runs slowly compared to progressive and/or iterative methods which have been developed for several years.
In 2012, two new phylogeny-aware tools appeared. One is called ''PAGAN'' that was developed by the same team as PRANK. The other is ''ProGraphMSA'' developed by Szalkowski. Both software packages were developed independently but share common features, notably the use of graph algorithms to improve the recognition of non-homologous regions, and an improvement in code making these software faster than PRANK.
Motif finding
 Motif finding, also known as profile analysis, is a method of locating
Motif finding, also known as profile analysis, is a method of locating sequence motif
In biology, a sequence motif is a nucleotide or amino-acid sequence pattern that is widespread and usually assumed to be related to biological function of the macromolecule. For example, an ''N''-glycosylation site motif can be defined as ''A ...
s in global MSAs that is both a means of producing a better MSA and a means of producing a scoring matrix for use in searching other sequences for similar motifs. A variety of methods for isolating the motifs have been developed, but all are based on identifying short highly conserved patterns within the larger alignment and constructing a matrix similar to a substitution matrix that reflects the amino acid or nucleotide composition of each position in the putative motif. The alignment can then be refined using these matrices. In standard profile analysis, the matrix includes entries for each possible character as well as entries for gaps. Alternatively, statistical pattern-finding algorithms can identify motifs as a precursor to an MSA rather than as a derivation. In many cases when the query set contains only a small number of sequences or contains only highly related sequences, pseudocounts are added to normalize the distribution reflected in the scoring matrix. In particular, this corrects zero-probability entries in the matrix to values that are small but nonzero.
Blocks analysis is a method of motif finding that restricts motifs to ungapped regions in the alignment. Blocks can be generated from an MSA or they can be extracted from unaligned sequences using a precalculated set of common motifs previously generated from known gene families. Block scoring generally relies on the spacing of high-frequency characters rather than on the calculation of an explicit substitution matrix.
Statistical pattern-matching has been implemented using both the expectation-maximization algorithm and the Gibbs sampler. One of the most common motif-finding tools, named Multiple EM for Motif Elicitation (MEME), uses expectation maximization and hidden Markov methods to generate motifs that are then used as search tools by its companion MAST in the combined suite MEME/MAST.
Non-coding multiple sequence alignment
Non-coding DNA regions, especiallytranscription factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription (genetics), transcription of genetics, genetic information from DNA to messenger RNA, by binding t ...
binding sites (TFBSs), are conserved, but not necessarily evolutionarily related, and may have converged from non-common ancestors. Thus, the assumptions used to align protein sequences and DNA coding regions are inherently different from those that hold for TFBS sequences. Although it is meaningful to align DNA coding regions for homologous sequences using mutation operators, alignment of binding site sequences for the same transcription factor cannot rely on evolutionary related mutation operations. Similarly, the evolutionary operator of point mutations can be used to define an edit distance for coding sequences, but this has little meaning for TFBS sequences because any sequence variation has to maintain a certain level of specificity for the binding site to function. This becomes specifically important when trying to align known TFBS sequences to build supervised models to predict unknown locations of the same TFBS. Hence, Multiple Sequence Alignment methods need to adjust the underlying evolutionary hypothesis and the operators used as in the work published incorporating neighbouring base thermodynamic information to align the binding sites searching for the lowest thermodynamic alignment conserving specificity of the binding site.
Optimization
Genetic algorithms and simulated annealing
Standard optimization techniques in computer science – both of which were inspired by, but do not directly reproduce, physical processes – have also been used in an attempt to more efficiently produce quality MSAs. One such technique,genetic algorithm
In computer science and operations research, a genetic algorithm (GA) is a metaheuristic inspired by the process of natural selection that belongs to the larger class of evolutionary algorithms (EA). Genetic algorithms are commonly used to g ...
s, has been used for MSA production in an attempt to broadly simulate the hypothesized evolutionary process that gave rise to the divergence in the query set. The method works by breaking a series of possible MSAs into fragments and repeatedly rearranging those fragments with the introduction of gaps at varying positions. A general objective function
In mathematical optimization and decision theory, a loss function or cost function (sometimes also called an error function) is a function that maps an event or values of one or more variables onto a real number intuitively representing some "cost ...
is optimized during the simulation, most generally the "sum of pairs" maximization function introduced in dynamic programming-based MSA methods. A technique for protein sequences has been implemented in the software program SAGA (Sequence Alignment by Genetic Algorithm) and its equivalent in RNA is called RAGA.
The technique of simulated annealing
Simulated annealing (SA) is a probabilistic technique for approximating the global optimum of a given function. Specifically, it is a metaheuristic to approximate global optimization in a large search space for an optimization problem. ...
, by which an existing MSA produced by another method is refined by a series of rearrangements designed to find better regions of alignment space than the one the input alignment already occupies. Like the genetic algorithm method, simulated annealing maximizes an objective function like the sum-of-pairs function. Simulated annealing uses a metaphorical "temperature factor" that determines the rate at which rearrangements proceed and the likelihood of each rearrangement; typical usage alternates periods of high rearrangement rates with relatively low likelihood (to explore more distant regions of alignment space) with periods of lower rates and higher likelihoods to more thoroughly explore local minima near the newly "colonized" regions. This approach has been implemented in the program MSASA (Multiple Sequence Alignment by Simulated Annealing).
Mathematical programming and exact solution algorithms
Mathematical programming
Mathematical optimization (alternatively spelled ''optimisation'') or mathematical programming is the selection of a best element, with regard to some criteria, from some set of available alternatives. It is generally divided into two subfiel ...
and in particular mixed integer programming models are another approach to solve MSA problems. The advantage of such optimization models is that they can be used to find the optimal MSA solution more efficiently compared to the traditional DP approach. This is due in part, to the applicability of decomposition techniques for mathematical programs, where the MSA model is decomposed into smaller parts and iteratively solved until the optimal solution is found. Example algorithms used to solve mixed integer programming models of MSA include branch and price
In applied mathematics, branch and price is a method of combinatorial optimization for solving integer linear programming (ILP) and mixed integer linear programming (MILP) problems with many variables. The method is a hybrid of branch and bound an ...
and Benders decomposition. Although exact approaches are computationally slow compared to heuristic algorithms for MSA, they are guaranteed to reach the optimal solution eventually, even for large-size problems.
Simulated quantum computing
In January 2017, D-Wave Systems announced that its qbsolv open-sourcequantum computing
A quantum computer is a computer that exploits quantum mechanical phenomena. On small scales, physical matter exhibits properties of wave-particle duality, both particles and waves, and quantum computing takes advantage of this behavior using s ...
software had been successfully used to find a faster solution to the MSA problem.
Alignment visualization and quality control
The necessary use of heuristics for multiple alignment means that for an arbitrary set of proteins, there is always a good chance that an alignment will contain errors. For example, an evaluation of several leading alignment programs using the BAliBase benchmark found that at least 24% of all pairs of aligned amino acids were incorrectly aligned. These errors can arise because of unique insertions into one or more regions of sequences, or through some more complex evolutionary process leading to proteins that do not align easily by sequence alone. As the number of sequence and their divergence increases many more errors will be made simply because of the heuristic nature of MSA algorithms. Multiple sequence alignment viewers enable alignments to be visually reviewed, often by inspecting the quality of alignment for annotated functional sites on two or more sequences. Many also enable the alignment to be edited to correct these (usually minor) errors, in order to obtain an optimal 'curated' alignment suitable for use in phylogenetic analysis or comparative modeling. However, as the number of sequences increases and especially in genome-wide studies that involve many MSAs it is impossible to manually curate all alignments. Furthermore, manual curation is subjective. And finally, even the best expert cannot confidently align the more ambiguous cases of highly diverged sequences. In such cases it is common practice to use automatic procedures to exclude unreliably aligned regions from the MSA. For the purpose of phylogeny reconstruction (see below) the Gblocks program is widely used to remove alignment blocks suspect of low quality, according to various cutoffs on the number of gapped sequences in alignment columns. However, these criteria may excessively filter out regions with insertion/deletion events that may still be aligned reliably, and these regions might be desirable for other purposes such as detection of positive selection. A few alignment algorithms output site-specific scores that allow the selection of high-confidence regions. Such a service was first offered by the SOAP program, which tests the robustness of each column to perturbation in the parameters of the popular alignment program CLUSTALW. The T-Coffee program uses a library of alignments in the construction of the final MSA, and its output MSA is colored according to confidence scores that reflect the agreement between different alignments in the library regarding each aligned residue. Its extension, Transitive Consistency Score (TCS), uses T-Coffeelibraries
A library is a collection of Book, books, and possibly other Document, materials and Media (communication), media, that is accessible for use by its members and members of allied institutions. Libraries provide physical (hard copies) or electron ...
of pairwise alignments to evaluate any third party MSA. Pairwise projections can be produced using fast or slow methods, thus allowing a trade-off between speed and accuracy. Another alignment program that can output an MSA with confidence scores is FSA, which uses a statistical model that allows calculation of the uncertainty in the alignment. The HoT (Heads-Or-Tails) score can be used as a measure of site-specific alignment uncertainty due to the existence of multiple co-optimal solutions. The GUIDANCE program calculates a similar site-specific confidence measure based on the robustness of the alignment to uncertainty in the guide tree that is used in progressive alignment programs. An alternative, more statistically justified approach to assess alignment uncertainty is the use of probabilistic evolutionary models for joint estimation of phylogeny and alignment. A Bayesian approach allows calculation of posterior probabilities of estimated phylogeny and alignment, which is a measure of the confidence in these estimates. In this case, a posterior probability can be calculated for each site in the alignment. Such an approach was implemented in the program BAli-Phy.
There are free programs available for visualization of multiple sequence alignments, for example Jalview and UGENE
UGENE is computer software for bioinformatics. It helps biologists to analyze various biological genetics data, such as sequences, annotations, multiple alignments, phylogenetic trees, NGS assemblies, and others. UGENE integrates dozens of well- ...
.
Phylogenetic use
Multiple sequence alignments can be used to create aphylogenetic tree
A phylogenetic tree or phylogeny is a graphical representation which shows the evolutionary history between a set of species or taxa during a specific time.Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA. In ...
. This is made possible by two reasons. The first is because functional domains that are known in annotated sequences can be used for alignment in non-annotated sequences. The other is that conserved regions known to be functionally important can be found. This makes it possible for multiple sequence alignments to be used to analyze and find evolutionary relationships through homology between sequences. Point mutations and insertion or deletion events (called indels) can be detected.
Multiple sequence alignments can also be used to identify functionally important sites, such as binding sites, active sites, or sites corresponding to other key functions, by locating conserved domains. When looking at multiple sequence alignments, it is useful to consider different aspects of the sequences when comparing sequences. These aspects include identity, similarity, and homology. Identity means that the sequences have identical residues at their respective positions. On the other hand, similarity has to do with the sequences being compared having similar residues quantitatively. For example, in terms of nucleotide sequences, pyrimidines are considered similar to each other, as are purines. Similarity ultimately leads to homology, in that the more similar sequences are, the closer they are to being homologous. This similarity in sequences can then go on to help find common ancestry.
See also
* Alignment-free sequence analysis *Cladistics
Cladistics ( ; from Ancient Greek 'branch') is an approach to Taxonomy (biology), biological classification in which organisms are categorized in groups ("clades") based on hypotheses of most recent common ancestry. The evidence for hypothesiz ...
* Generalized tree alignment
* Multiple sequence alignment viewers
*PANDIT
A pandit (; ; also spelled pundit, pronounced ; abbreviated Pt. or Pdt.) is an individual with specialised knowledge or a teacher of any field of knowledge in Hinduism, particularly the Vedic scriptures, dharma, or Hindu philosophy; in colonial-e ...
, a biological database covering protein domains
*Phylogenetics
In biology, phylogenetics () is the study of the evolutionary history of life using observable characteristics of organisms (or genes), which is known as phylogenetic inference. It infers the relationship among organisms based on empirical dat ...
* Sequence alignment software
* Structural alignment
References
Survey articles
* * * * *External links
ExPASy sequence alignment tools
– from the Virtual School of Natural Sciences
– from Pôle Bioinformatique Lyonnais
An entry point to clustal servers and information
An entry point to the main T-Coffee servers
An entry point to the main MergeAlign server and information
*European Bioinformatics Institute servers: *
ClustalW2
– general purpose multiple sequence alignment program for DNA or proteins. *
Muscle
– MUltiple Sequence Comparison by Log-Expectation *
T-coffee
– multiple sequence alignment. *
MAFFT
– Multiple Alignment using Fast Fourier Transform *
KALIGN
– a fast and accurate multiple sequence alignment algorithm.
Lecture notes, tutorials, and courses
Multiple sequence alignment lectures
– from the Max Planck Institute for Molecular Genetics
on multiple sequence alignments at the
European Molecular Biology Laboratory
The European Molecular Biology Laboratory (EMBL) is an intergovernmental organization dedicated to molecular biology research and is supported by 29 member states, two prospect member states, and one associate member state. EMBL was created in ...
(EMBL)
Molecular Bioinformatics Lecture Notes
{{DEFAULTSORT:Multiple Sequence Alignment Bioinformatics Computational phylogenetics Markov models NP-complete problems