CRISPR activation on:
[Wikipedia]
[Google]
[Amazon]
CRISPR activation (CRISPRa) is a type of


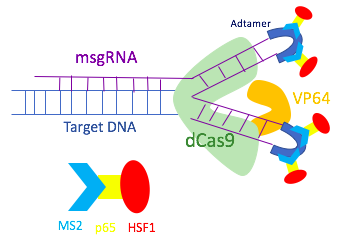 In order to assemble different transcriptional activators, the dCas9-SAM system uses a modified single guide RNA (sgRNA) that has binding sites for the MS2 protein. Hairpin aptamers are attached to the tetra loop and the stem loop 2 of the sgRNA to become binding sites for dimerized MS2 bacteriophage coat proteins. As the hairpins are exposed outside of the dCas9-sgRNA complex, other transcriptional factors can bind to the MS2 protein without disrupting the dCas9-sgRNA complex. Thus, the MS2 protein is engineered to include p65 and HSF1 proteins. The MS2-p65-HSF1 fusion protein interacts with the dCas9-VP64 to recruit more transcriptional factors onto the promoter of the target genes.
Employing the dCas-SAM system, Zhang et al. (2015) successfully reactivated the latent HIV gene to over-express viral proteins from the HIV host cells. They were able to over-express viral proteins substantially to trigger apoptosis of HIV-1 latent cells due to the toxicity of viral proteins. In another dCas-SAM system experiment, Konermann et al. (2015) found genes in melanoma cells that give resistance to a BRAF inhibitor through activating candidate genes via dCas system. Thus, the dCas9-SAM system can further be employed to activate latent genes, develop gene therapies, and discover new genes.
In order to assemble different transcriptional activators, the dCas9-SAM system uses a modified single guide RNA (sgRNA) that has binding sites for the MS2 protein. Hairpin aptamers are attached to the tetra loop and the stem loop 2 of the sgRNA to become binding sites for dimerized MS2 bacteriophage coat proteins. As the hairpins are exposed outside of the dCas9-sgRNA complex, other transcriptional factors can bind to the MS2 protein without disrupting the dCas9-sgRNA complex. Thus, the MS2 protein is engineered to include p65 and HSF1 proteins. The MS2-p65-HSF1 fusion protein interacts with the dCas9-VP64 to recruit more transcriptional factors onto the promoter of the target genes.
Employing the dCas-SAM system, Zhang et al. (2015) successfully reactivated the latent HIV gene to over-express viral proteins from the HIV host cells. They were able to over-express viral proteins substantially to trigger apoptosis of HIV-1 latent cells due to the toxicity of viral proteins. In another dCas-SAM system experiment, Konermann et al. (2015) found genes in melanoma cells that give resistance to a BRAF inhibitor through activating candidate genes via dCas system. Thus, the dCas9-SAM system can further be employed to activate latent genes, develop gene therapies, and discover new genes.

CRISPR
CRISPR () (an acronym for clustered regularly interspaced short palindromic repeats) is a family of DNA sequences found in the genomes of prokaryotic organisms such as bacteria and archaea. These sequences are derived from DNA fragments of bacte ...
tool that uses modified versions of CRISPR effectors without endonuclease
Endonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain. Some, such as deoxyribonuclease I, cut DNA relatively nonspecifically (without regard to sequence), while many, typically called restriction endonucleases ...
activity, with added transcriptional
Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules calle ...
activators on dCas9 or the guide RNA
A guide RNA (gRNA) is a piece of RNA that functions as a guide for RNA- or DNA-targeting enzymes, with which it forms complexes. Very often these enzymes will delete, insert or otherwise alter the targeted RNA or DNA. They occur naturally, serv ...
s (gRNAs).
Like for CRISPR interference
CRISPR interference (CRISPRi) is a genetic perturbation technique that allows for sequence-specific repression of gene expression in prokaryotic and eukaryotic cells. It was first developed by Stanley Qi and colleagues in the laboratories of Wen ...
, the CRISPR effector is guided to the target by a complementary guide RNA. However, CRISPR activation systems are fused to transcriptional activators to increase expression of genes of interest. Such systems are usable for many purposes including but not limited to, genetic screens and overexpression of proteins of interest.
The most commonly-used effector is based on Cas9
Cas9 (CRISPR associated protein 9, formerly called Cas5, Csn1, or Csx12) is a 160 kilodalton protein which plays a vital role in the immunological defense of certain bacteria against DNA viruses and plasmids, and is heavily utilized in genetic e ...
(from Type II systems), but other effectors like Cas12a
Cas12a (CRISPR associated protein 12a, previously known as Cpf1) is an RNA-guided endonuclease of that forms part of the CRISPR system in some bacteria and is used by scientists to modify DNA. It originates as part of a bacterial immune mechan ...
(Type V) have been used as well.
Components
dCas9
Cas9 Endonuclease Dead, also known as dead Cas9 or dCas9, is a mutant form ofCas9
Cas9 (CRISPR associated protein 9, formerly called Cas5, Csn1, or Csx12) is a 160 kilodalton protein which plays a vital role in the immunological defense of certain bacteria against DNA viruses and plasmids, and is heavily utilized in genetic e ...
whose endonuclease activity is removed through point mutations in its endonuclease domains. Similar to its unmutated form, dCas9 is used in CRISPR systems along with gRNAs to target specific genes or nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules wi ...
s complementary to the gRNA with PAM sequences that allow Cas9 to bind. Cas9 ordinarily has 2 endonuclease domains called the RuvC and HNH domains. The point mutations D10A and H840A change 2 important residues for endonuclease activity that ultimately results in its deactivation. Although dCas9 lacks endonuclease activity, it is still capable of binding to its guide RNA and the DNA strand that is being targeted because such binding is managed by other domains. This alone is often enough to attenuate if not outright block transcription of the targeted gene if the gRNA positions dCas9 in a way that prevents transcriptional factors and RNA polymerase from accessing the DNA. However, this ability to bind DNA can also be exploited for activation since dCas9 has modifiable regions, typically the N and C terminus of the protein, that can be used to attach transcriptional activators.
Guide RNA
See:Guide RNA
A guide RNA (gRNA) is a piece of RNA that functions as a guide for RNA- or DNA-targeting enzymes, with which it forms complexes. Very often these enzymes will delete, insert or otherwise alter the targeted RNA or DNA. They occur naturally, serv ...
, CRISPR
CRISPR () (an acronym for clustered regularly interspaced short palindromic repeats) is a family of DNA sequences found in the genomes of prokaryotic organisms such as bacteria and archaea. These sequences are derived from DNA fragments of bacte ...
A small guide RNA (sgRNA), or gRNA is an RNA with around 20 nucleotides used to direct Cas9 or dCas9 to their targets. gRNAs contain two major regions of importance for CRISPR systems: the scaffold and spacer regions. The spacer region has nucleotides that are complementary to those found on the target genes, often in the promoter region. The scaffold region is responsible for formation of a complex with (d)Cas9. Together, they bind (d)Cas9 and direct it to the gene(s) of interest. Since the spacer region of a gRNA can be modified for any potential sequence, they give CRISPR systems much more flexibility as any genes and nucleotides with a sequence complementary to the spacer region can become possible targets.
Transcriptional activators
See: Transcriptional Activator,Transcription Factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. The fu ...
Transcriptional Activators are protein domains or whole proteins linked to dCas9 or sgRNAs that assist in the recruitment of important co-factors as well as RNA Polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template.
Using the enzyme helicase, RNAP locally opens the ...
for transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of the gene(s) targeted by the system. In order for a protein to be made from the gene that encodes it, RNA polymerase must make RNA from the DNA template of the gene during a process called transcription. Transcriptional activators have a DNA binding domain and a domain for activation of transcription. The activation domain can recruit general transcription factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. The fu ...
s or RNA polymerase to the gene sequence. Activation domains can also function by facilitating transcription by stalled RNA polymerases, and in eukaryotes can act to move nucleosome
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins and resembles thread wrapped around a spool. The nucleosome is the fundamen ...
s on the DNA or modify histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wr ...
s to increase gene expression. These activators can be introduced into the system through attachment to dCas9 or to the sgRNA. Some researchers have noted that the extent of transcriptional upregulation can be modulated by using multiple sites for activator attachment in one experiment and by using different variations and combinations of activators at once in a given experiment or sample.
Expression system
Anexpression system
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. The ...
is required for the introduction of the gRNAs and (d)Cas9 proteins into the cells of interest. Typically employed options include but are not limited to plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; how ...
s and viral vectors such as adeno-associated virus
Adeno-associated viruses (AAV) are small viruses that infect humans and some other primate species. They belong to the genus ''Dependoparvovirus'', which in turn belongs to the family ''Parvoviridae''. They are small (approximately 26 nm in di ...
(AAV) vector or lentivirus
''Lentivirus'' is a genus of retroviruses that cause chronic and deadly diseases characterized by long incubation periods, in humans and other mammalian species. The genus includes the human immunodeficiency virus (HIV), which causes AIDS. Lent ...
vector.
Specific activation systems
VP64-p65-Rta
The VP64-p65-Rta, or VPR, dCas9 activator was created by modifying an existing dCas9 activator, in which a Vp64 transcriptional activator is joined to the C terminus of dCas9. In the dCas9-VPR protein, the transcription factors p65 and Rta are added to the C terminus of dCas9-Vp64. Therefore, all three transcription factors are targeted to the same gene. The use of three transcription factors, as opposed to solely Vp64, results in increased expression of targeted genes. When different genes were targeted by dCas9, they all showed significantly greater expression with dCas9-VPR than with dCas9-VP64. It has also been demonstrated that dCas9-VPR can be used to increase expression of multiple genes within the same cell by putting multiple sgRNAs into the same cell. dCas9-VPR has been used to activate the neurogenin 2 (link) and neurogenic differentiation 1 (link) genes, resulting in differentiation of induced pluripotent stem cells into inducedneuron
A neuron, neurone, or nerve cell is an electrically excitable cell that communicates with other cells via specialized connections called synapses. The neuron is the main component of nervous tissue in all animals except sponges and placozoa. N ...
s. A study comparing dCas9 activators found that the VPR, SAM, and Suntag activators worked best with dCas9 to increase gene expression in a variety of fruit fly, mouse
A mouse ( : mice) is a small rodent. Characteristically, mice are known to have a pointed snout, small rounded ears, a body-length scaly tail, and a high breeding rate. The best known mouse species is the common house mouse (''Mus musculus' ...
, and human
Humans (''Homo sapiens'') are the most abundant and widespread species of primate, characterized by bipedalism and exceptional cognitive skills due to a large and complex brain. This has enabled the development of advanced tools, culture, ...
cell types.
Synergistic activation mediator
To overcome the limitation of the dCas9-VP64 gene activation system, the dCas9-SAM system was developed to incorporate multiple transcriptional factors. Utilizing MS2, p65, andHSF1
Heat shock factor 1 (HSF1) is a protein that in humans is encoded by the ''HSF1'' gene. HSF1 is highly conserved in eukaryotes and is the primary mediator of transcriptional responses to proteotoxic stress with important roles in non-stress regul ...
proteins, dCas9-SAM system recruits various transcriptional factors working synergistically to activate the gene of interest. In order to assemble different transcriptional activators, the dCas9-SAM system uses a modified single guide RNA (sgRNA) that has binding sites for the MS2 protein. Hairpin aptamers are attached to the tetra loop and the stem loop 2 of the sgRNA to become binding sites for dimerized MS2 bacteriophage coat proteins. As the hairpins are exposed outside of the dCas9-sgRNA complex, other transcriptional factors can bind to the MS2 protein without disrupting the dCas9-sgRNA complex. Thus, the MS2 protein is engineered to include p65 and HSF1 proteins. The MS2-p65-HSF1 fusion protein interacts with the dCas9-VP64 to recruit more transcriptional factors onto the promoter of the target genes.
Employing the dCas-SAM system, Zhang et al. (2015) successfully reactivated the latent HIV gene to over-express viral proteins from the HIV host cells. They were able to over-express viral proteins substantially to trigger apoptosis of HIV-1 latent cells due to the toxicity of viral proteins. In another dCas-SAM system experiment, Konermann et al. (2015) found genes in melanoma cells that give resistance to a BRAF inhibitor through activating candidate genes via dCas system. Thus, the dCas9-SAM system can further be employed to activate latent genes, develop gene therapies, and discover new genes.
In order to assemble different transcriptional activators, the dCas9-SAM system uses a modified single guide RNA (sgRNA) that has binding sites for the MS2 protein. Hairpin aptamers are attached to the tetra loop and the stem loop 2 of the sgRNA to become binding sites for dimerized MS2 bacteriophage coat proteins. As the hairpins are exposed outside of the dCas9-sgRNA complex, other transcriptional factors can bind to the MS2 protein without disrupting the dCas9-sgRNA complex. Thus, the MS2 protein is engineered to include p65 and HSF1 proteins. The MS2-p65-HSF1 fusion protein interacts with the dCas9-VP64 to recruit more transcriptional factors onto the promoter of the target genes.
Employing the dCas-SAM system, Zhang et al. (2015) successfully reactivated the latent HIV gene to over-express viral proteins from the HIV host cells. They were able to over-express viral proteins substantially to trigger apoptosis of HIV-1 latent cells due to the toxicity of viral proteins. In another dCas-SAM system experiment, Konermann et al. (2015) found genes in melanoma cells that give resistance to a BRAF inhibitor through activating candidate genes via dCas system. Thus, the dCas9-SAM system can further be employed to activate latent genes, develop gene therapies, and discover new genes.
SunTag
The SunTag activator system uses the dCas9 protein, which is modified to be linked with the SunTag. The SunTag is a repeating polypeptide array that can recruit multiple copies ofantibodies
An antibody (Ab), also known as an immunoglobulin (Ig), is a large, Y-shaped protein used by the immune system to identify and neutralize foreign objects such as pathogenic bacteria and viruses. The antibody recognizes a unique molecule of the ...
. Through attaching transcriptional factors on the antibodies, the SunTag dCas9 activating complex amplifies its recruitment of transcriptional factors. In order to guide the dCas9 protein to its target gene, the dCas9 SunTag system uses sgRNA.
Tanenbaum et al.(2014) are credited for creating the dCas9 SunTag system. For the antibodies
An antibody (Ab), also known as an immunoglobulin (Ig), is a large, Y-shaped protein used by the immune system to identify and neutralize foreign objects such as pathogenic bacteria and viruses. The antibody recognizes a unique molecule of the ...
, they employed GCN4 antibodies which was bound to transcriptional factor VP64. In order to transport the antibodies to the nuclei of the cells, they attached NLS tag. To confirm the nuclear localization of the antibodies, sfGFP was used for visualization purpose. Therefore, the GCN4-sfGFP-NLS-VP64 protein was developed to be interact with dCas SunTag system. The antibodies successfully bound to SunTag polypeptides and activated target CXCR4 gene in K562 cell lines. Comparing with the dCas9-VP64 activation complex, they were able to increase the CXCR4 gene expression 5-25 times greater in K562 cell lines. Not only was there a greater CXCR4 protein overexpression but also CXCR4 proteins were active to further travel on the transwell migration assay. Thus, the dCas9-SunTag system can be used to activate genes that are present latently such as virus genes.

Applications
The dCas9 activation system allows a desired gene or multiple genes in the same cell to be expressed. It is possible to study genes involved in a certain process using agenome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
wide screen that involves activating expression of genes. Examining which sgRNAs yield a phenotype suggests which genes are involved in a specific pathway. The dCas9 activation system can be used to control exactly which cells are activated and at what time activation occurs. dCas9 constructs have been made that turn on a dCas9-activator fusion protein in the presence of light or chemicals. Cells can also be reprogrammed or differentiated from one cell type into another by increasing the expression of certain genes important for the formation or maintenance of a cell type.
Greater control over gene expression
One research group used a system in which dCas9 was fused to a particular domain, C1B1. When blue light is shined on the cell, the cryptochrome 2 (Cry2) domain binds to C1B1. The Cry2 domain is fused to a transcriptional activator, so blue light targets the activator to the spot where dCas9 is bound. The use of light allows a great deal of control over when the targeted gene is activated. Removing the light from the cell results in only dCas9 remaining at the target gene, so expression is not increased. In this way, the system is reversible. A similar system was developed using chemical control. In this system, dCas9 recruits an MS2 fusion protein that contains the domain FKBP. In the presence of the chemical RAP, an FRB domain fused to a chromatin modifying complex binds to FKBP. Whenever RAP is added to the cells, a specific chromatin modifier complex can be targeted to the gene. That allows scientists to examine how specificchromatin
Chromatin is a complex of DNA and protein found in eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important roles in r ...
modifications affect the expression of a gene. The dCAs9-VPR system is used as an activator by targeting it to the promoter of a gene upstream of the coding region. A study used various sgRNAs to target different portions of the gene, finding that the dCas9-VPR activator can act as an activator or a repressor, depending on the location it binds. In a cell, sgRNAs targeting the promoter could allow dCas9-VPR to increase expression, while sgRNAs targeting the coding region of the gene result in dCas9-VPR decreasing expression.
Genome wide activation
The versatility of sgRNAs allows dCas9 activators to increase the expression of any gene within an organism's genome. That could be used to increase expression of a protein coding gene or a transcribed RNA. A paper demonstrated that genome wide activation could be used to determine which proteins are involved in mediated resistance to a specific drug. Another paper used genome wide activation of long, noncoding RNAs and observed that increasing the expression of certain long noncoding RNAs conferred resistance to the drug vemurafenib. In both cases, the cells that survive the drug could be studied to determine which sgRNAs they contain. That allows researchers to determine which gene was activated in each surviving cell, which suggests which genes are important for resistance to that drug.Use in organisms
A dCas9 fusion with VP64, p65, andHSF1
Heat shock factor 1 (HSF1) is a protein that in humans is encoded by the ''HSF1'' gene. HSF1 is highly conserved in eukaryotes and is the primary mediator of transcriptional responses to proteotoxic stress with important roles in non-stress regul ...
(heat shock factor 1) allowed researchers to target genes in Arabidopsis thaliana
''Arabidopsis thaliana'', the thale cress, mouse-ear cress or arabidopsis, is a small flowering plant native to Eurasia and Africa. ''A. thaliana'' is considered a weed; it is found along the shoulders of roads and in disturbed land.
A winter a ...
and increase transcription to a similar level as when the gene itself is inserted into the plant's genome. For one of the two genes tested, the dCas9 activator changes the number and size of leaves and made the plants better able to handle drought. The authors conclude that the dCas9 activator can create phenotypes in plants that are similar to those observed when a transgene is inserted for overexpression. Researchers have used multiple guide RNAs to target dCas9 activation system to multiple genes in a specific mouse strain in which dCas9 can be turned on in specific cell lines using the Cre recombinase
Cre recombinase is a tyrosine recombinase enzyme derived from the P1 bacteriophage. The enzyme uses a topoisomerase I-like mechanism to carry out site specific recombination events. The enzyme (38kDa) is a member of the integrase family of site ...
system. Scientists used the targeting and increased expression of several genes to examine the processes involved in regeneration
Regeneration may refer to:
Science and technology
* Regeneration (biology), the ability to recreate lost or damaged cells, tissues, organs and limbs
* Regeneration (ecology), the ability of ecosystems to regenerate biomass, using photosynthesis
...
and carcinoma
Carcinoma is a malignancy that develops from epithelial cells. Specifically, a carcinoma is a cancer that begins in a tissue that lines the inner or outer surfaces of the body, and that arises from cells originating in the endodermal, mesodermal ...
s of the liver
The liver is a major Organ (anatomy), organ only found in vertebrates which performs many essential biological functions such as detoxification of the organism, and the Protein biosynthesis, synthesis of proteins and biochemicals necessary for ...
.
References
{{Authority control Genetic engineering Genome editing