|
Replication Protein A
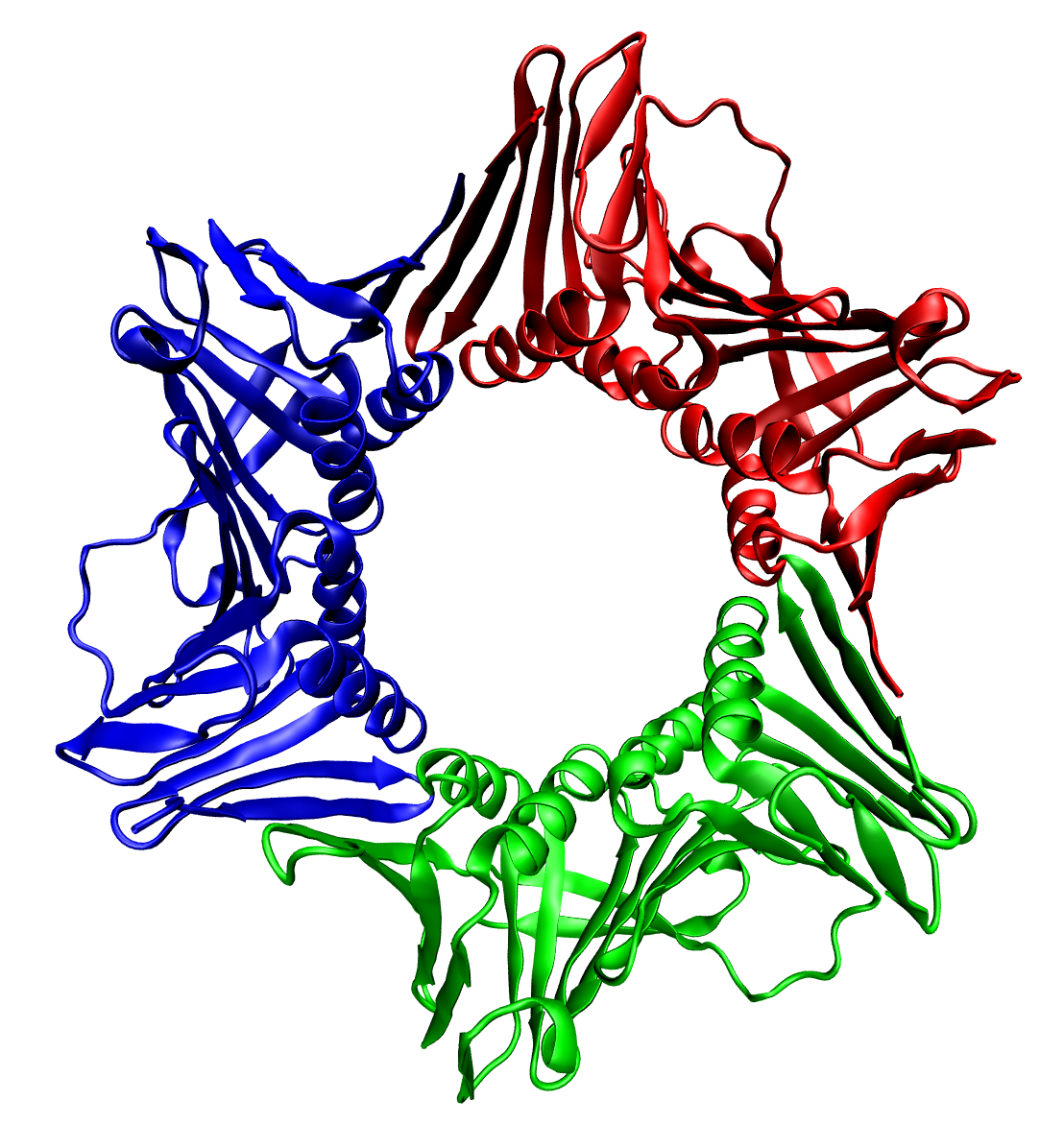
Replication protein A (RPA) is the major protein that binds to single-stranded DNA (ssDNA) in eukaryotic cells. In vitro, RPA shows a much higher affinity for ssDNA than RNA or double-stranded DNA. RPA is required in replication, recombination and repair processes such as nucleotide excision repair and homologous recombination. It also plays roles in responding to damaged DNA. Structure RPA is a heterotrimer, composed of the subunits RPA1 (RPA70) (70kDa subunit), RPA2 (RPA32) (32kDa subunit) and RPA3 (RPA14) (14kDa subunit). The three RPA subunits contain six OB-folds (oligonucleotide/oligosaccharide binding),with DNA-binding domains (DBD) designated DBDs A-F, that bind RPA to single-stranded DNA. DBDs A, B, C and F are located on RPA1, DBD D is located on RPA2, and DBD E is located on RPA3. DBDs C, D, and E make up the trimerization core of the protein with flexible linker regions connecting them all together. Due to these flexible linker regions RPA is considere ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Replication Protein A1
Replication protein A 70 kDa DNA-binding subunit is a protein that in humans is encoded by the ''RPA1'' gene. Interactions Replication protein A1 has been shown to interact with: * BRCA2, * BLM, * MCM2, * MCM4, * MCM6, * MCM7, * MUTYH, * ORC2L, * ORC6L, * P53, * RPA2, * RPA3, * TIPIN, * TP53BP1, and * XPA. See also * Replication protein A * Replication protein A2 * Replication protein A3 * Single-stranded binding protein Single-stranded binding proteins (SSBs) are a class of proteins that have been identified in both viruses and organisms from bacteria to humans. Viral SSB Although the overall picture of ''human cytomegalovirus'' (HHV-5) DNA synthesis appears ... References Further reading * * * * * * * * * * * * * * * * * * {{DNA replication ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Trimer
In biochemistry, a protein trimer is a macromolecular complex formed by three, usually non-covalently bound, macromolecules like proteins or nucleic acids. A homotrimer would be formed by three identical molecules. A heterotrimer would be formed by three different macromolecules. Type II Collagen is an example of homotrimeric protein. Porins usually arrange themselves in membranes as trimers. Bacteriophage T4 tail fiber Multiple copies of a polypeptide encoded by a gene often can form an aggregate referred to as a multimer. When a multimer is formed from polypeptides produced by two different mutant alleles of a particular gene, the mixed multimer may exhibit greater functional activity than the unmixed multimers formed by each of the mutants alone. When a mixed multimer displays increased functionality relative to the unmixed multimers, the phenomenon is referred to as intragenic complementation. The distal portion of each of the bacteriophage T4 tail fibers is encoded by ge ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Meiosis
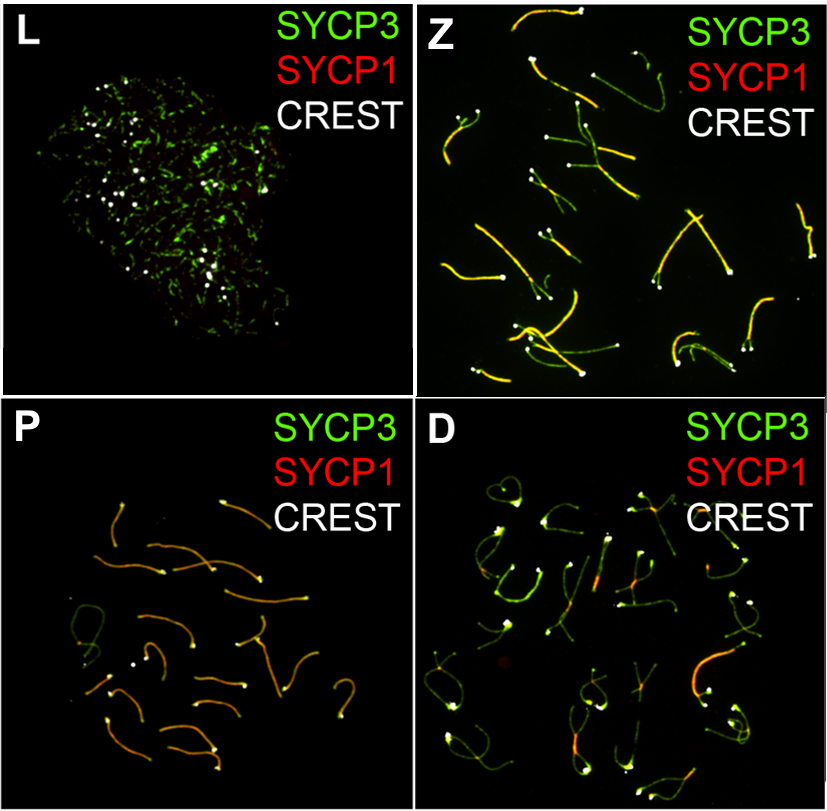
Meiosis (; , since it is a reductional division) is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately result in four cells with only one copy of each chromosome ( haploid). Additionally, prior to the division, genetic material from the paternal and maternal copies of each chromosome is crossed over, creating new combinations of code on each chromosome. Later on, during fertilisation, the haploid cells produced by meiosis from a male and female will fuse to create a cell with two copies of each chromosome again, the zygote. Errors in meiosis resulting in aneuploidy (an abnormal number of chromosomes) are the leading known cause of miscarriage and the most frequent genetic cause of developmental disabilities. In meiosis, DNA replication is followed by two rounds of cell division to produce four daughter cells, each with half the number of chr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Meiosis
Meiosis (; , since it is a reductional division) is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately result in four cells with only one copy of each chromosome ( haploid). Additionally, prior to the division, genetic material from the paternal and maternal copies of each chromosome is crossed over, creating new combinations of code on each chromosome. Later on, during fertilisation, the haploid cells produced by meiosis from a male and female will fuse to create a cell with two copies of each chromosome again, the zygote. Errors in meiosis resulting in aneuploidy (an abnormal number of chromosomes) are the leading known cause of miscarriage and the most frequent genetic cause of developmental disabilities. In meiosis, DNA replication is followed by two rounds of cell division to produce four daughter cells, each with half the number of chr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Repair
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages (interstrand crosslinks or ICLs). This can eventually lead to malignant ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase
A polymerase is an enzyme ( EC 2.7.7.6/7/19/48/49) that synthesizes long chains of polymers or nucleic acids. DNA polymerase and RNA polymerase are used to assemble DNA and RNA molecules, respectively, by copying a DNA template strand using base-pairing interactions or RNA by half ladder replication. A DNA polymerase from the thermophilic bacterium, ''Thermus aquaticus'' (''Taq'') ( PDBbr>1BGX EC 2.7.7.7) is used in the polymerase chain reaction, an important technique of molecular biology. A polymerase may be template dependent or template independent. Poly-A-polymerase is an example of template independent polymerase. Terminal deoxynucleotidyl transferase also known to have template independent and template dependent activities. Types By function *DNA polymerase (DNA-directed DNA polymerase, DdDP) **Family A: DNA polymerase I; Pol γ, θ, ν **Family B: DNA polymerase II; Pol α, δ, ε, ζ **Family C: DNA polymerase III holoenzyme **Family X: Pol β, λ, μ * ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Endonuclease
Endonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain. Some, such as deoxyribonuclease I, cut DNA relatively nonspecifically (without regard to sequence), while many, typically called restriction endonucleases or restriction enzymes, cleave only at very specific nucleotide sequences. Endonucleases differ from exonucleases, which cleave the ends of recognition sequences instead of the middle (endo) portion. Some enzymes known as "exo-endonucleases", however, are not limited to either nuclease function, displaying qualities that are both endo- and exo-like. Evidence suggests that endonuclease activity experiences a lag compared to exonuclease activity. Restriction enzymes are endonucleases from eubacteria and archaea that recognize a specific DNA sequence. The nucleotide sequence recognized for cleavage by a restriction enzyme is called the restriction site. Typically, a restriction site will be a palindromic sequence about four to six nucleotides ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stoichiometry
Stoichiometry refers to the relationship between the quantities of reactants and products before, during, and following chemical reactions. Stoichiometry is founded on the law of conservation of mass where the total mass of the reactants equals the total mass of the products, leading to the insight that the relations among quantities of reactants and products typically form a ratio of positive integers. This means that if the amounts of the separate reactants are known, then the amount of the product can be calculated. Conversely, if one reactant has a known quantity and the quantity of the products can be empirically determined, then the amount of the other reactants can also be calculated. This is illustrated in the image here, where the balanced equation is: : Here, one molecule of methane reacts with two molecules of oxygen gas to yield one molecule of carbon dioxide and two molecules of water. This particular chemical equation is an example of complete combustion. St ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CST Complex
The CST complex is a cellular multiprotein complex involved in telomere maintenance. In budding yeast (''Saccharomyces cerevisiae''), it is composed of the proteins Cdc13, Stn1, and Ten1; in mammals, it consists of the proteins CTC1, STN1, and TEN1. It is related to the Replication protein A complex. Structure For budding yeast as well as for mammals, CST is a protein heterotrimer, consisting of three distinct proteins. Yeast Stn1 and Ten1 are orthologous proteins to mammalian STN1 and TEN1. But yeast Cdc13 and mammalian CTC1 are very different in amino acid sequence, length, and to some extent in function. Function For both budding yeast and mammals, the CST complex contributes to telomere maintenance, but this function is more crucial for budding yeast, where the CST complex performs the functions that shelterin performs in vertebrates. At least four factors contribute to telomere maintenance: telomerase, shelterin, TERRA and the CST Complex. CST protection of telomeres for ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphorylation
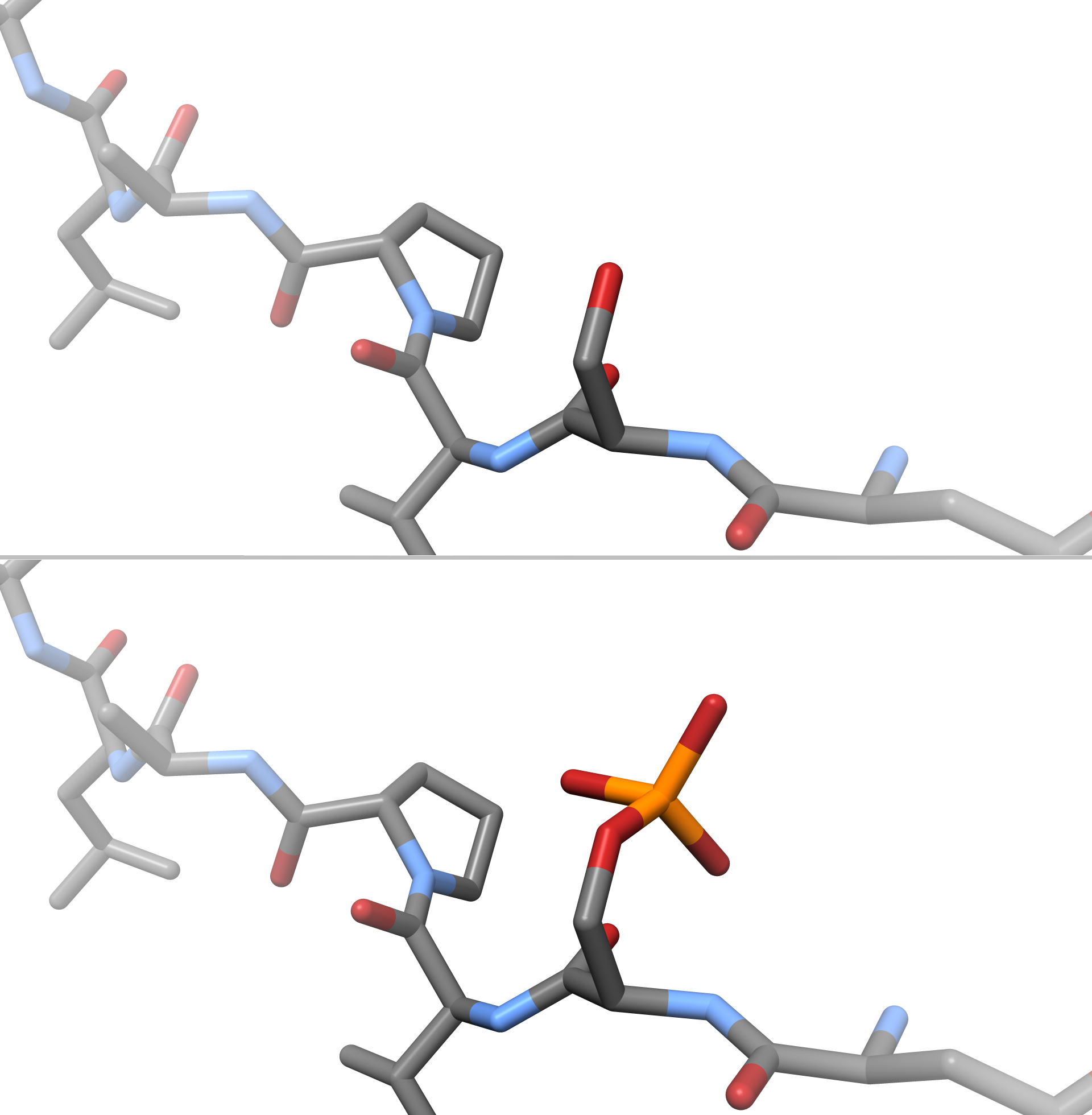
In chemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology and could be driven by natural selection. Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. Protein phosphorylation often activates (or deactivates) many enzymes. Glucose Phosphorylation of sugars is often the first stage in their catabolism. Phosphorylation allows cells to accumulate sugars because the phosphate group prevents the molecules from diffusing back across their transporter. Phosphorylation of glucose is a key reaction in sugar metabolism. The chemical equation for the conversion of D-glucose to D-glucose-6-phosphate in the first step of glycolysis is given by :D-glucose + ATP → D-glucose-6-phosphate + ADP : ΔG° = −16.7 kJ/mol (° indicates measurement at standard condition) Hepatic cells are freely permeable to glucose, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C-terminus
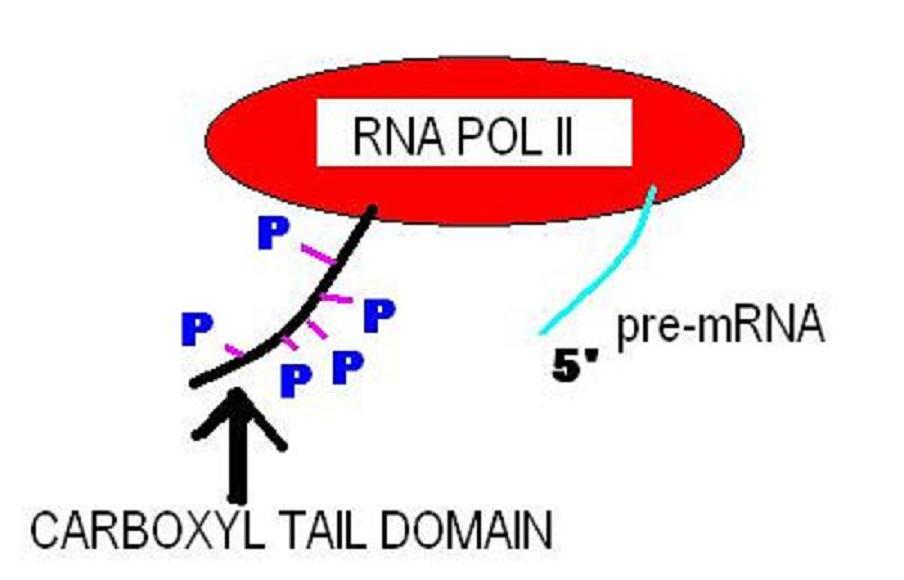
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is translated from messenger RNA, it is created from N-terminus to C-terminus. The convention for writing peptide sequences is to put the C-terminal end on the right and write the sequence from N- to C-terminus. Chemistry Each amino acid has a carboxyl group and an amine group. Amino acids link to one another to form a chain by a dehydration reaction which joins the amine group of one amino acid to the carboxyl group of the next. Thus polypeptide chains have an end with an unbound carboxyl group, the C-terminus, and an end with an unbound amine group, the N-terminus. Proteins are naturally synthesized starting from the N-terminus and ending at the C-terminus. Function C-terminal retention signals While the N-terminus of a protein often c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
N-terminus
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein. Chemistry Each amino acid has an amine group and a carboxylic group. Amino acids link to one another by peptide bonds which form through a dehydration reaction that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |