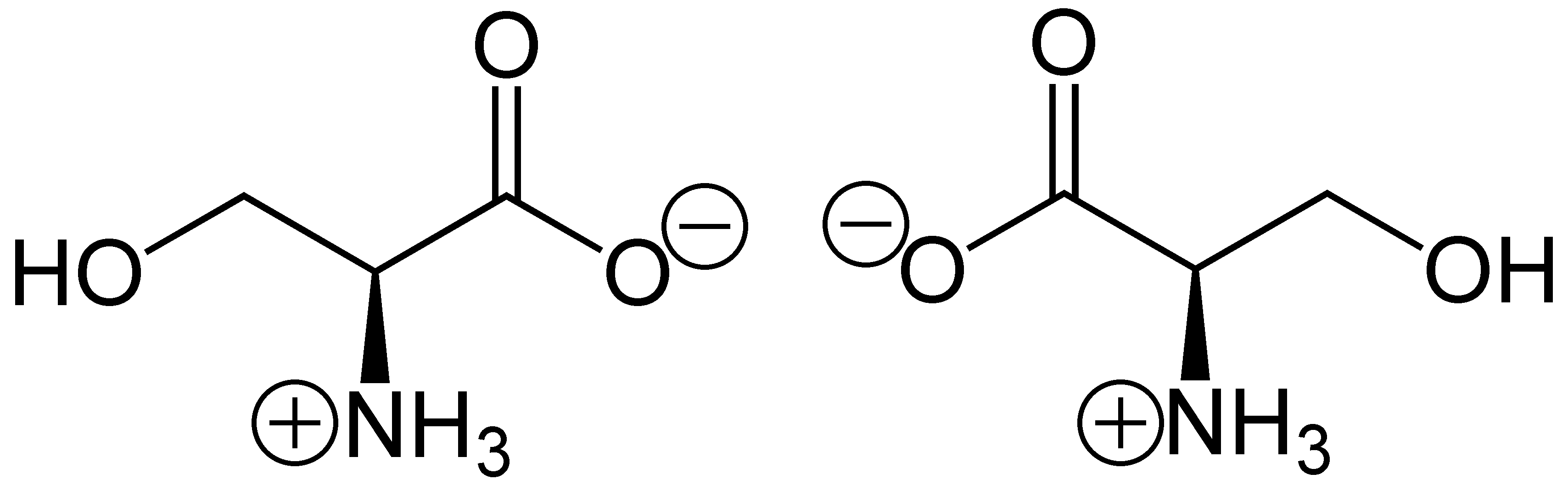|
Myophosphorylase
Myophosphorylase or glycogen phosphorylase, muscle associated (PYGM) is the muscle isoform of the enzyme ''glycogen phosphorylase'' and is encoded by the ''PYGM'' gene. This enzyme helps break down glycogen (a form of stored carbohydrate) into glucose-1-phosphate (not glucose), so it can be used within the muscle cell. Mutations in this gene are associated with McArdle disease (GSD-V, myophosphorylase deficiency), a glycogen storage disease of muscle. Myophosphorylase comes in two forms: form 'a' is phosphorylated by phosphorylase kinase, form 'b' is not phosphorylated. Form 'a' is de-phosphorylated into form 'b' by the enzyme phosphoprotein phosphatase, which is activated by elevated insulin. Both forms 'a' and 'b' of myophosphorylase have two conformational states: active (R or relaxed) and inactive (T or tense). When either form 'a' or 'b' are in the active state, then the enzyme converts glycogen into glucose-1-phosphate. Myophosphorylase-b is allosterically activated by ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
McArdle Disease
Glycogen storage disease type V (GSD5, GSD-V), also known as McArdle's disease, is a metabolic disorder, one of the metabolic myopathies, more specifically a muscle glycogen storage disease, caused by a deficiency of myophosphorylase. Its incidence is reported as one in 100,000, roughly the same as glycogen storage disease type I. The disease was first reported in 1951 by Dr. Brian McArdle of Guy's Hospital, London. Signs and symptoms The onset of this disease is usually noticed in childhood, but often not diagnosed until the third or fourth decade of life. Symptoms include exercise intolerance with muscle pain, early fatigue, painful cramps, inappropriate rapid heart rate response to exercise, and may include myoglobin in the urine (often provoked by a bout of exercise).Lucia A, Martinuzzi A, Nogales-Gadea G, Quinlivan R, Reason S; International Association for Muscle Glycogen Storage Disease study group. Clinical practice guidelines for glycogen storage disease V & VII (McA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Isoform
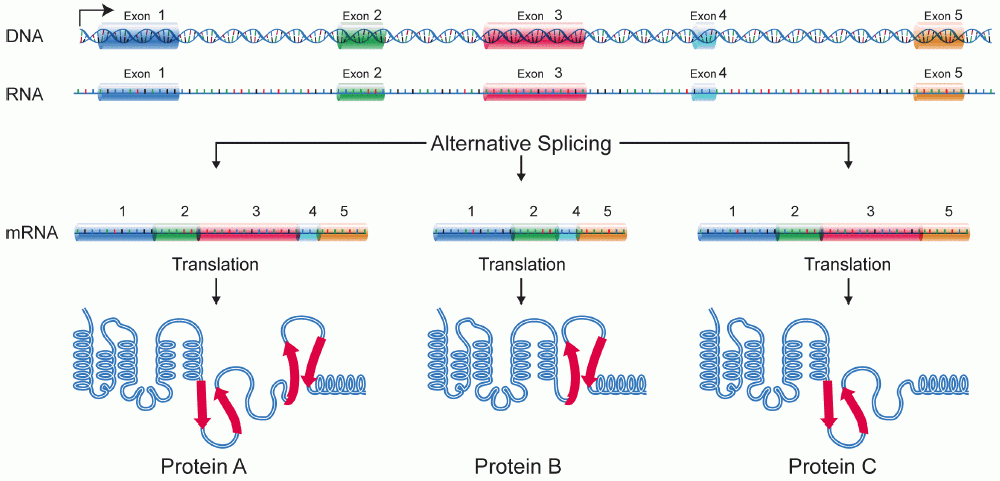
A protein isoform, or "protein variant", is a member of a set of highly similar proteins that originate from a single gene or gene family and are the result of genetic differences. While many perform the same or similar biological roles, some isoforms have unique functions. A set of protein isoforms may be formed from alternative splicings, variable promoter usage, or other post-transcriptional modifications of a single gene; post-translational modifications are generally not considered. (For that, see Proteoforms.) Through RNA splicing mechanisms, mRNA has the ability to select different protein-coding segments ( exons) of a gene, or even different parts of exons from RNA to form different mRNA sequences. Each unique sequence produces a specific form of a protein. The discovery of isoforms could explain the discrepancy between the small number of protein coding regions genes revealed by the human genome project and the large diversity of proteins seen in an organism: different ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exon
An exon is any part of a gene that will form a part of the final mature RNA produced by that gene after introns have been removed by RNA splicing. The term ''exon'' refers to both the DNA sequence within a gene and to the corresponding sequence in RNA transcripts. In RNA splicing, introns are removed and exons are covalently joined to one another as part of generating the mature RNA. Just as the entire set of genes for a species constitutes the genome, the entire set of exons constitutes the exome. History The term ''exon'' derives from the expressed region and was coined by American biochemist Walter Gilbert in 1978: "The notion of the cistron… must be replaced by that of a transcription unit containing regions which will be lost from the mature messengerwhich I suggest we call introns (for intragenic regions)alternating with regions which will be expressedexons." This definition was originally made for protein-coding transcripts that are spliced before being translated. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allosteric Enzyme
Allosteric enzymes are enzymes that change their conformational ensemble upon binding of an effector (allosteric modulator) which results in an apparent change in binding affinity at a different ligand binding site. This "action at a distance" through binding of one ligand affecting the binding of another at a distinctly different site, is the essence of the allosteric concept. Allostery plays a crucial role in many fundamental biological processes, including but not limited to cell signaling and the regulation of metabolism. Allosteric enzymes need not be oligomers as previously thought, and in fact many systems have demonstrated allostery within single enzymes. In biochemistry, allosteric regulation (or allosteric control) is the regulation of a protein by binding an effector molecule at a site other than the enzyme's active site. The site to which the effector binds is termed the ''allosteric site''. Allosteric sites allow effectors to bind to the protein, often resulting in a c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphorylase
In biochemistry, phosphorylases are enzymes that catalyze the addition of a phosphate group from an inorganic phosphate (phosphate+hydrogen) to an acceptor. :A-B + P A + P-B They include allosteric enzymes that catalyze the production of glucose-1-phosphate from a glucan such as glycogen, starch or maltodextrin. Phosphorylase is also a common name used for glycogen phosphorylase in honor of Earl W. Sutherland Jr., who in the late 1930s discovered it as the first phosphorylase. Function Phosphorylases should not be confused with phosphatases, which remove phosphate groups. In more general terms, phosphorylases are enzymes that catalyze the addition of a phosphate group from an inorganic phosphate (phosphate + hydrogen) to an acceptor, not to be confused with a phosphatase (a hydrolase) or a kinase (a phosphotransferase). A phosphatase removes a phosphate group from a donor using water, whereas a kinase transfers a phosphate group from a donor (usually ATP) to an acceptor. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine
Serine (symbol Ser or S) is an α-amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated − form under biological conditions), a carboxyl group (which is in the deprotonated − form under biological conditions), and a side chain consisting of a hydroxymethyl group, classifying it as a polar amino acid. It can be synthesized in the human body under normal physiological circumstances, making it a nonessential amino acid. It is encoded by the codons UCU, UCC, UCA, UCG, AGU and AGC. Occurrence This compound is one of the naturally occurring proteinogenic amino acids. Only the L-stereoisomer appears naturally in proteins. It is not essential to the human diet, since it is synthesized in the body from other metabolites, including glycine. Serine was first obtained from silk protein, a particularly rich source, in 1865 by Emil Cramer. Its name is derived from the Latin for silk, ''sericum''. Serine's structure was estab ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphorylation
In chemistry, phosphorylation is the attachment of a phosphate group to a molecule or an ion. This process and its inverse, dephosphorylation, are common in biology and could be driven by natural selection. Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. Protein phosphorylation often activates (or deactivates) many enzymes. Glucose Phosphorylation of sugars is often the first stage in their catabolism. Phosphorylation allows cells to accumulate sugars because the phosphate group prevents the molecules from diffusing back across their transporter. Phosphorylation of glucose is a key reaction in sugar metabolism. The chemical equation for the conversion of D-glucose to D-glucose-6-phosphate in the first step of glycolysis is given by :D-glucose + ATP → D-glucose-6-phosphate + ADP : ΔG° = −16.7 kJ/mol (° indicates measurement at standard condition) Hepatic cells are freely permeable to glucose, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Post-translational Modification
Post-translational modification (PTM) is the covalent and generally enzymatic modification of proteins following protein biosynthesis. This process occurs in the endoplasmic reticulum and the golgi apparatus. Proteins are synthesized by ribosomes translating mRNA into polypeptide chains, which may then undergo PTM to form the mature protein product. PTMs are important components in cell signaling, as for example when prohormones are converted to hormones. Post-translational modifications can occur on the amino acid side chains or at the protein's C- or N- termini. They can extend the chemical repertoire of the 20 standard amino acids by modifying an existing functional group or introducing a new one such as phosphate. Phosphorylation is a highly effective mechanism for regulating the activity of enzymes and is the most common post-translational modification. Many eukaryotic and prokaryotic proteins also have carbohydrate molecules attached to them in a process called glycosyla ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lysine
Lysine (symbol Lys or K) is an α-amino acid that is a precursor to many proteins. It contains an α-amino group (which is in the protonated form under biological conditions), an α-carboxylic acid group (which is in the deprotonated −COO− form under biological conditions), and a side chain lysyl ((CH2)4NH2), classifying it as a basic, charged (at physiological pH), aliphatic amino acid. It is encoded by the codons AAA and AAG. Like almost all other amino acids, the α-carbon is chiral and lysine may refer to either enantiomer or a racemic mixture of both. For the purpose of this article, lysine will refer to the biologically active enantiomer L-lysine, where the α-carbon is in the ''S'' configuration. The human body cannot synthesize lysine. It is essential in humans and must therefore be obtained from the diet. In organisms that synthesise lysine, two main biosynthetic pathways exist, the diaminopimelate and α-aminoadipate pathways, which employ distinct e ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphoserine
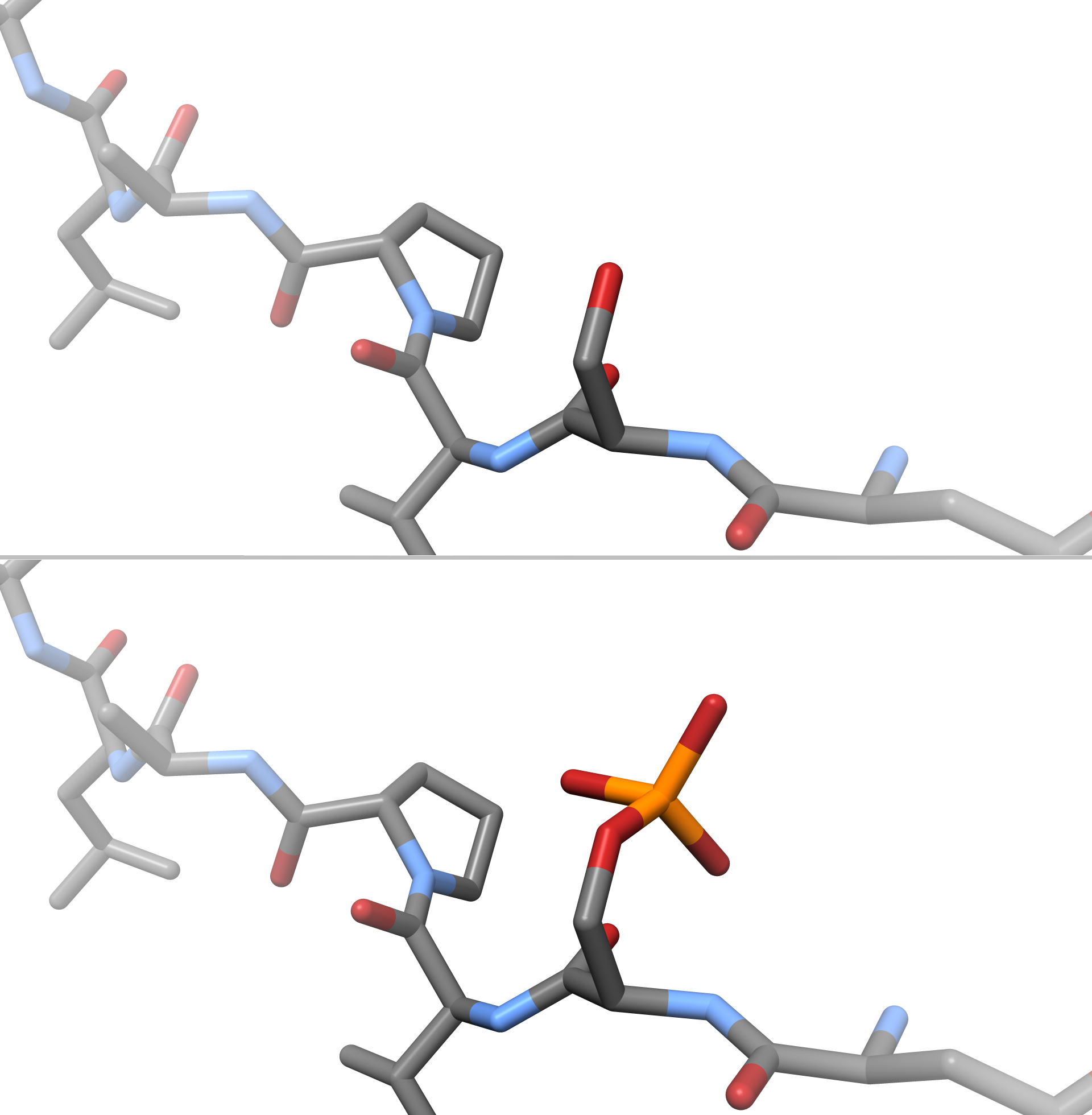
Phosphoserine (abbreviated as SEP or J) is an ester of serine and phosphoric acid. Phosphoserine is a component of many proteins as the result of posttranslational modifications. The phosphorylation of the alcohol functional group in serine to produce phosphoserine is catalyzed by various types of kinases. Through the use of technologies that utilize an expanded genetic code, phosphoserine can also be incorporated into proteins during translation. It is a normal metabolite In biochemistry, a metabolite is an intermediate or end product of metabolism. The term is usually used for small molecules. Metabolites have various functions, including fuel, structure, signaling, stimulatory and inhibitory effects on enzymes, c ... found in human biofluids. Phosphoserine has three potential coordination sites (carboxyl, amine and phosphate group) Determination of the mode of coordination between phosphorylated ligands and metal ions occurring in an organism is a first step to explain the fu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helix
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence. The alpha helix is also called a classic Pauling–Corey–Branson α-helix. The name 3.613-helix is also used for this type of helix, denoting the average number of residues per helical turn, with 13 atoms being involved in the ring formed by the hydrogen bond. Among types of local structure in proteins, the α-helix is the most extreme and the most predictable from sequence, as well as the most prevalent. Discovery In the early 1930s, William Astbury showed that there were drastic changes in the X-ray fiber diffraction of moist wool or hair fibers upon significant stretching. The data suggested that the unstretched fibers had a coiled molecular structure with a characteristic repeat of ≈. Astb ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta Sheet
The beta sheet, (β-sheet) (also β-pleated sheet) is a common motif of the regular protein secondary structure. Beta sheets consist of beta strands (β-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. A β-strand is a stretch of polypeptide chain typically 3 to 10 amino acids long with backbone in an extended conformation. The supramolecular association of β-sheets has been implicated in the formation of the fibrils and protein aggregates observed in amyloidosis, notably Alzheimer's disease. History The first β-sheet structure was proposed by William Astbury in the 1930s. He proposed the idea of hydrogen bonding between the peptide bonds of parallel or antiparallel extended β-strands. However, Astbury did not have the necessary data on the bond geometry of the amino acids in order to build accurate models, especially since he did not then know that the peptide bond was planar. A refined versi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |