|
Transcriptomic Technologies
Transcriptomics technologies are the techniques used to study an organism's transcriptome, the sum of all of its RNA transcripts. The information content of an organism is recorded in the DNA of its genome and expressed through transcription. Here, mRNA serves as a transient intermediary molecule in the information network, whilst non-coding RNAs perform additional diverse functions. A transcriptome captures a snapshot in time of the total transcripts present in a cell. Transcriptomics technologies provide a broad account of which cellular processes are active and which are dormant. A major challenge in molecular biology is to understand how a single genome gives rise to a variety of cells. Another is how gene expression is regulated. The first attempts to study whole transcriptomes began in the early 1990s. Subsequent technological advances since the late 1990s have repeatedly transformed the field and made transcriptomics a widespread discipline in biological sciences. There a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcriptome
The transcriptome is the set of all RNA transcripts, including coding and non-coding, in an individual or a population of cells. The term can also sometimes be used to refer to all RNAs, or just mRNA, depending on the particular experiment. The term ''transcriptome'' is a portmanteau of the words ''transcript'' and ''genome''; it is associated with the process of transcript production during the biological process of transcription. The early stages of transcriptome annotations began with cDNA libraries published in the 1980s. Subsequently, the advent of high-throughput technology led to faster and more efficient ways of obtaining data about the transcriptome. Two biological techniques are used to study the transcriptome, namely DNA microarray, a hybridization-based technique and RNA-seq, a sequence-based approach. RNA-seq is the preferred method and has been the dominant transcriptomics technique since the 2010s. Single-cell transcriptomics allows tracking of transcript change ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Assay
An assay is an investigative (analytic) procedure in laboratory medicine, mining, pharmacology, environmental biology and molecular biology for qualitatively assessing or quantitatively measuring the presence, amount, or functional activity of a target entity. The measured entity is often called the analyte, the measurand, or the target of the assay. The analyte can be a drug, biochemical substance, chemical element or compound, or cell in an organism or organic sample. An assay usually aims to measure an analyte's intensive property and express it in the relevant measurement unit (e.g. molarity, density, functional activity in enzyme international units, degree of effect in comparison to a standard, etc.). If the assay involves exogenous reactants (the reagents), then their quantities are kept fixed (or in excess) so that the quantity and quality of the target are the only limiting factors. The difference in the assay outcome is used to deduce the unknown quality ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequencing By Synthesis
Illumina dye sequencing is a technique used to determine the series of base pairs in DNA, also known as DNA sequencing. The reversible terminated chemistry concept was invented by Bruno Canard and Simon Sarfati at the Pasteur Institute in Paris. It was developed by Shankar Balasubramanian and David Klenerman of Cambridge University, who subsequently founded Solexa, a company later acquired by Illumina. This sequencing method is based on reversible dye-terminators that enable the identification of single nucleotides as they are washed over DNA strands. It can also be used for whole-genome and region sequencing, transcriptome analysis, metagenomics, small RNA discovery, methylation profiling, and genome-wide protein-nucleic acid interaction analysis. Overview This works in three basic steps: amplify, sequence, and analyze. The process begins with purified DNA. The DNA is fragmented and adapters are added that contain segments that act as reference points during amplification, seque ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Expressed Sequence Tag
In genetics, an expressed sequence tag (EST) is a short sub-sequence of a cDNA sequence. ESTs may be used to identify gene transcripts, and were instrumental in gene discovery and in gene-sequence determination. The identification of ESTs has proceeded rapidly, with approximately 74.2 million ESTs now available in public databases (e.g. GenBank 1 January 2013, all species). EST approaches have largely been superseded by whole genome and transcriptome sequencing and metagenome sequencing. An EST results from one-shot sequencing of a cloned cDNA. The cDNAs used for EST generation are typically individual clones from a cDNA library. The resulting sequence is a relatively low-quality fragment whose length is limited by current technology to approximately 500 to 800 nucleotides. Because these clones consist of DNA that is complementary to mRNA, the ESTs represent portions of expressed genes. They may be represented in databases as either cDNA/mRNA sequence or as the reverse compleme ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sanger Sequencing
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Frederick Sanger and colleagues in 1977, it became the most widely used sequencing method for approximately 40 years. An automated instrument using slab gel electrophoresis and fluorescent labels was first commercialized by Applied Biosystems in March 1987. Later, automated slab gels were replaced with automated capillary array electrophoresis. Recently, higher volume Sanger sequencing has been replaced by next generation sequencing methods, especially for large-scale, automated genome analyses. However, the Sanger method remains in wide use for smaller-scale projects and for validation of deep sequencing results. It still has the advantage over short-read sequencing technologies (like Illumina) in that it can produce DNA sequence reads of > ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
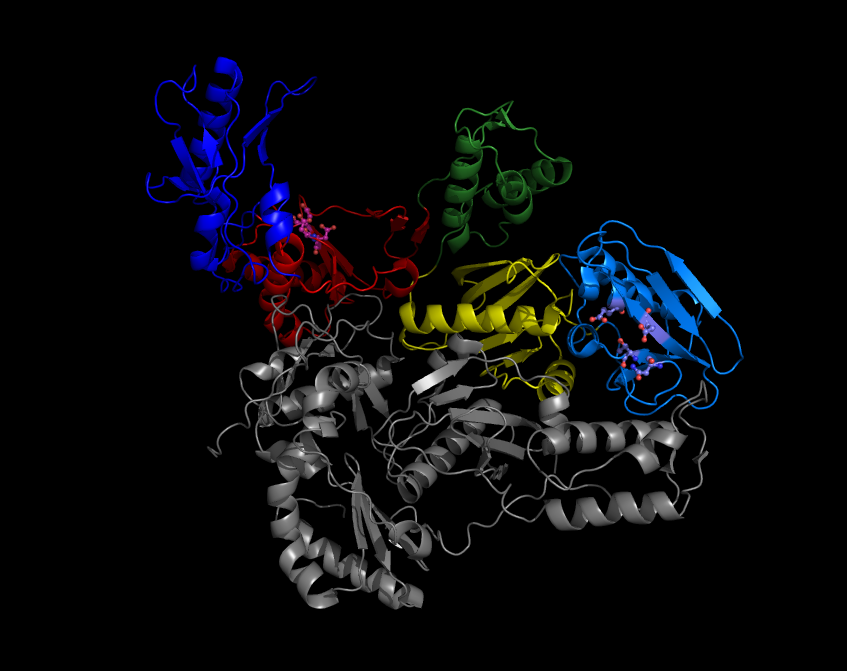
Reverse Transcriptase
A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. The process does not violate the flows of genetic information as described by the classical central dogma, but rather expands it to include transfers of information from RNA to DNA. Retroviral RT has three sequential biochemical activities: RNA-dependent DNA polymerase activity, ribonuclease H (RNase H), and DNA-dependent DNA polymerase activity. Collectively, these activities enable the enzyme to convert single-stranded RNA into double-stranded cDNA. In retroviruses and retrotransposons, this cDNA can then integrate into the host genome, from which new RNA copies can be made via host-cell ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Complementary DNA
In genetics, complementary DNA (cDNA) is DNA that was reverse transcribed (via reverse transcriptase) from an RNA (e.g., messenger RNA or microRNA). cDNA exists in both single-stranded and double-stranded forms and in both natural and engineered forms. In engineered forms, it often is a copy (replicate) of the naturally occurring DNA from any particular organism's natural genome; the organism's own mRNA was naturally transcribed from its DNA, and the cDNA is reverse transcribed from the mRNA, yielding a duplicate of the original DNA. Engineered cDNA is often used to express a specific protein in a cell that does not normally express that protein (i.e., heterologous expression), or to sequence or quantify mRNA molecules using DNA based methods (qPCR, RNA-seq). cDNA that codes for a specific protein can be transferred to a recipient cell for expression as part of recombinant DNA, often bacterial or yeast expression systems. cDNA is also generated to analyze transcriptomic pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Antheraea Polyphemus
''Antheraea polyphemus'', the Polyphemus moth, is a North American member of the family Saturniidae, the giant silk moths. It is a tan-colored moth, with an average wingspan of 15 cm (6 in). The most notable feature of the moth is its large, purplish eyespots on its two hindwings. The eyespots give it its name – from the Greek myth of the cyclops Polyphemus. The species was first described by Pieter Cramer in 1776. The species is widespread in continental North America, with local populations found throughout subarctic Canada and the United States. The caterpillar can eat 86,000 times its weight at emergence in a little less than two months. Polyphemus moths are considered to be very polyphagous, meaning they eat from a wide variety of plants. Life cycle The life cycle of the moth is much like that of any other Saturniidae species. It lays flat, light-brown eggs on the leaves of a number of host trees, preferring ''Ulmus americana'' (American elm), ''Betula'' (birc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
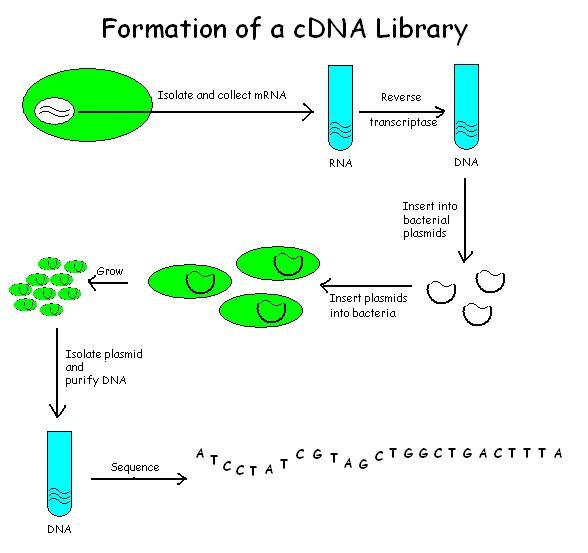
CDNA Library
A cDNA library is a combination of cloned cDNA (complementary DNA) fragments inserted into a collection of host cells, which constitute some portion of the transcriptome of the organism and are stored as a "library". cDNA is produced from fully transcribed mRNA found in the nucleus and therefore contains only the expressed genes of an organism. Similarly, tissue-specific cDNA libraries can be produced. In eukaryotic cells the mature mRNA is already spliced, hence the cDNA produced lacks introns and can be readily expressed in a bacterial cell. While information in cDNA libraries is a powerful and useful tool since gene products are easily identified, the libraries lack information about enhancers, introns, and other regulatory elements found in a genomic DNA library. cDNA Library Construction cDNA is created from a mature mRNA from a eukaryotic cell with the use of reverse transcriptase. In eukaryotes, a poly-(A) tail (consisting of a long sequence of adenine nucleotides) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Primary Transcript
A primary transcript is the single-stranded ribonucleic acid (RNA) product synthesized by transcription of DNA, and processed to yield various mature RNA products such as mRNAs, tRNAs, and rRNAs. The primary transcripts designated to be mRNAs are modified in preparation for translation. For example, a precursor mRNA (pre-mRNA) is a type of primary transcript that becomes a messenger RNA (mRNA) after processing. Pre-mRNA is synthesized from a DNA template in the cell nucleus by transcription. Pre-mRNA comprises the bulk of heterogeneous nuclear RNA (hnRNA). Once pre-mRNA has been completely processed, it is termed " mature messenger RNA", or simply "messenger RNA". The term hnRNA is often used as a synonym for pre-mRNA, although, in the strict sense, hnRNA may include nuclear RNA transcripts that do not end up as cytoplasmic mRNA. There are several steps contributing to the production of primary transcripts. All these steps involve a series of interactions to initiate and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell (biology)
The cell is the basic structural and functional unit of all life, forms of life. Every cell consists of cytoplasm enclosed within a Cell membrane, membrane; many cells contain organelles, each with a specific function. The term comes from the Latin word meaning 'small room'. Most cells are only visible under a light microscope, microscope. Cells Abiogenesis, emerged on Earth about 4 billion years ago. All cells are capable of Self-replication, replication, protein synthesis, and cell motility, motility. Cells are broadly categorized into two types: eukaryotic cells, which possess a Cell nucleus, nucleus, and prokaryotic, prokaryotic cells, which lack a nucleus but have a nucleoid region. Prokaryotes are single-celled organisms such as bacteria, whereas eukaryotes can be either single-celled, such as amoebae, or multicellular organism, multicellular, such as some algae, plants, animals, and fungi. Eukaryotic cells contain organelles including Mitochondrion, mitochondria, which ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tissue (biology)
In biology, tissue is an assembly of similar cells and their extracellular matrix from the same embryonic origin that together carry out a specific function. Tissues occupy a Biological organisation#Levels, biological organizational level between cell (biology), cells and a complete organ (biology), organ. Accordingly, organs are formed by the functional grouping together of multiple tissues. The English word "tissue" Morphological derivation, derives from the French word "", the past participle of the verb tisser, "to weave". The study of tissues is known as histology or, in connection with disease, as histopathology. Xavier Bichat is considered as the "Father of Histology". Plant histology is Studied Space Shuttle designs, studied in both plant anatomy and Plant physiology, physiology. The classical tools for studying tissues are the Microtome#Applications, paraffin block in which tissue is embedded and then sectioned, the staining, histological stain, and the Microscope, o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |