|
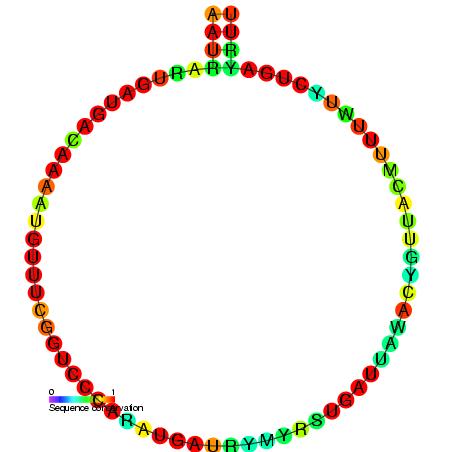
Small Nucleolar RNA SNORD75
In molecular biology, Small Nucleolar RNA SNORD75 (also known as U75) is a non-coding RNA (ncRNA) molecule which functions in the biogenesis (modification) of other small nuclear RNAs (snRNAs). This type of modifying RNA is located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA. U75 (SNORD75) belongs to the C/D box class of snoRNAs which contain the C (UGAUGA) and D (CUGA) box motifs. Most of the members of the box C/D family function in directing site-specific 2′-O-methylation of substrate RNAs. The mouse snoRNA Z19 is orthologous to human U75. U75 is predicted to guide the 2′-O-ribose methylation of 28S ribosomal RNA Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional form of ''local segments'' of proteins. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein folds into its three dimensional tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the amino hydrogen and carboxyl oxygen atoms in the peptide backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik Linderstrøm-Lang at Stanford in 1952. Other types of biopolymers such as nucleic acids also possess characteristic secondary structures. Types The most common secondary st ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Guide RNA
A guide RNA (gRNA) is a piece of RNA that functions as a guide for RNA- or DNA-targeting enzymes, with which it forms complexes. Very often these enzymes will delete, insert or otherwise alter the targeted RNA or DNA. They occur naturally, serving important functions, but can also be designed to be used for targeted editing, such as with CRISPR-Cas9 and CRISPR-Cas12. History RNA-editing Guide RNA was discovered in 1990 by B. Blum, N. Bakalara, and L. Simpson in the mitochondria of protists called Leishmania tarentolae. The guide RNA there is encoded in maxicircle DNA and contains sequences matching those within the edited regions of the mRNA. They enable the cleavage, insertion and deletion of bases. Guide RNA in Protists Trypanosomatid protists and other kinetoplastids have a novel post-transcriptional mitochondrial RNA modification process known as "RNA editing". They have a large segment of highly organized DNA segments in their mitochondria. This mitochondrial DNA is cir ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nucleolar RNA SNORD79
In molecular biology, snoRNA U79 (also known as SNORD79 or Z22) is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA. U79 belongs to the C/D box class of snoRNAs which contain the conserved sequence motifs known as the C box (UGAUGA) and the D box (CUGA). Most of the members of the box C/D family function in directing site-specific 2'-O-methylation of substrate RNAs. snoRNA U79 is found in intron 7 of the GAS5 gene in humans and is also present in mice. U79 is predicted to guide the 2'O-ribose methylation of 28S ribosomal RNA Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synt ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nucleolar RNA SNORD78
In molecular biology, SNORD78 (also known as U78) belongs to the C/D family of snoRNAs. It is predicted to guide 2'O-ribose methylation of the large 28S rRNA Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from riboso ... subunit at position G4593. The snoRNAs U44, U47, U74, U75, U76, U77, U79, U80 and U81, also of the C/D family, share the same host gene with U78 (protein non-coding). References External links * Small nuclear RNA {{molecular-cell-biology-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nucleolar RNA SNORD77
In molecular biology, SNORD77 (also known as U77) belongs to the C/D family of snoRNAs. It is predicted to guide 2'O-ribose methylation of large 28S rRNA Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from riboso ... subunit at position A1521. The C/D snoRNAs U44, U47, U74, U75, U76, U78, U79, U80 and U81 share the same host gene as U77 (non-coding). References External links * Small nuclear RNA {{molecular-cell-biology-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nucleolar RNA SNORD47
In molecular biology, SNORD47 (also known as U47) is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA. snoRNA U57 belongs to the C/D box class of snoRNAs which contain the conserved sequence motifs known as the C box (UGAUGA) and the D box (CUGA). Most of the members of the box C/D family function in directing site-specific 2'-O-methylation of substrate RNAs. This snoRNA was originally cloned from HeLa cells and expression verified by northern blotting. It is predicted to guide 2'O-ribose methylation of ribosomal RNA Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nucleolar RNA SNORD44
In molecular biology, snoRNA U44 (also known as SNORD44) is a non-coding RNA (ncRNA) molecule which functions in the modification of other small nuclear RNAs (snRNAs). This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. It is known as a small nucleolar RNA (snoRNA) and also often referred to as a guide RNA. snoRNA U44 belongs to the C/D box class of snoRNAs which contain the conserved sequence motifs known as the C box (UGAUGA) and the D box (CUGA). Most of the members of the box C/D family function in directing site-specific 2'-O-methylation of substrate RNAs. U44 was originally cloned from HeLa cells and expression verified by northern blotting. It is predicted to guide 2'O-ribose methylation of 18S ribosomal RNA Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal DNA (rDNA) and then bound to ribosomal proteins to form small and large ribosome subunits. rRNA is the physical and mechanical factor of the ribosome that forces transfer RNA (tRNA) and messenger RNA (mRNA) to process and translate the latter into proteins. Ribosomal RNA is the predominant form of RNA found in most cells; it makes up about 80% of cellular RNA despite never being translated into proteins itself. Ribosomes are composed of approximately 60% rRNA and 40% ribosomal proteins by mass. Structure Although the primary structure of rRNA sequences can vary across organisms, base-pairing within these sequences commonly forms stem-loop configurations. The length and position of these rRNA stem-loops allow them to create three-dimensio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Methylation
In the chemical sciences, methylation denotes the addition of a methyl group on a substrate, or the substitution of an atom (or group) by a methyl group. Methylation is a form of alkylation, with a methyl group replacing a hydrogen atom. These terms are commonly used in chemistry, biochemistry, soil science, and the biological sciences. In biological systems, methylation is catalyzed by enzymes; such methylation can be involved in modification of heavy metals, regulation of gene expression, regulation of protein function, and RNA processing. In vitro methylation of tissue samples is also one method for reducing certain histological staining artifacts. The reverse of methylation is demethylation. In biology In biological systems, methylation is accomplished by enzymes. Methylation can modify heavy metals, regulate gene expression, RNA processing and protein function. It has been recognized as a key process underlying epigenetics. Methanogenesis Methanogenesis, the proce ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SnoRNA
In molecular biology, Small nucleolar RNAs (snoRNAs) are a class of small RNA molecules that primarily guide chemical modifications of other RNAs, mainly ribosomal RNAs, transfer RNAs and small nuclear RNAs. There are two main classes of snoRNA, the C/D box snoRNAs, which are associated with methylation, and the H/ACA box snoRNAs, which are associated with pseudouridylation. SnoRNAs are commonly referred to as guide RNAs but should not be confused with the guide RNAs that direct RNA editing in trypanosomes. snoRNA guided modifications After transcription, nascent rRNA molecules (termed pre-rRNA) undergo a series of processing steps to generate the mature rRNA molecule. Prior to cleavage by exo- and endonucleases, the pre-rRNA undergoes a complex pattern of nucleoside modifications. These include methylations and pseudouridylations, guided by snoRNAs. *Methylation is the attachment or substitution of a methyl group onto various substrates. The rRNA of humans contain approxima ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Conservation
In evolutionary biology, conserved sequences are identical or similar Sequence (biology), sequences in nucleic acids (DNA sequence, DNA and RNA) or peptide sequence, proteins across species (homology (biology)#Orthology, orthologous sequences), or within a genome (homology (biology)#Paralogy, paralogous sequences), or between donor and receptor taxa (Sequence homology#Xenology, xenologous sequences). Conservation indicates that a sequence has been maintained by natural selection. A highly conserved sequence is one that has remained relatively unchanged far back up the phylogenetic tree, and hence far back in geological time. Examples of highly conserved sequences include the Ribosomal RNA, RNA components of ribosomes present in all domain (biology), domains of life, the homeobox sequences widespread amongst Eukaryotes, and the tmRNA in Bacteria. The study of sequence conservation overlaps with the fields of genomics, proteomics, evolutionary biology, phylogenetics, bioinformatics ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |