|
Master Mix (PCR)
A master mix is a mixture containing precursors and enzymes used as an ingredient in RT-PCR techniques in molecular biology. Such mixtures contain a mixture dNTPs (required as a substrate for the building of new DNA strands), MgCl2, Taq polymerase (an enzyme required to building new DNA strands), a pH buffer and come mixed in nuclease-free water. Master mixes for real-time PCR include a fluorescent compound (frequently SYBR green), and the choice of mix also influence test sensitivity and consistency. Differences in the choice of master mixes can sometimes explain difference in experimental results, a particular case being the measurement of telomere A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature mos ... length. References DNA sequencing {{Molecular-biology-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RT-PCR
Reverse transcription polymerase chain reaction (RT-PCR) is a laboratory technique combining reverse transcription of RNA into DNA (in this context called complementary DNA or cDNA) and amplification of specific DNA targets using polymerase chain reaction (PCR). It is primarily used to measure the amount of a specific RNA. This is achieved by monitoring the amplification reaction using fluorescence, a technique called real-time PCR or quantitative PCR (qPCR). Combined RT-PCR and qPCR are routinely used for analysis of gene expression and quantification of viral RNA in research and clinical settings. The close association between RT-PCR and qPCR has led to metonymic use of the term qPCR to mean RT-PCR. Such use may be confusing, as RT-PCR can be used without qPCR, for example to enable molecular cloning, sequencing or simple detection of RNA. Conversely, qPCR may be used without RT-PCR, for example to quantify the copy number of a specific piece of DNA. Nomenclature The combine ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNTP
A nucleoside triphosphate is a nucleoside containing a nitrogenous base bound to a 5-carbon sugar (either ribose or deoxyribose), with three phosphate groups bound to the sugar. They are the molecular precursors of both DNA and RNA, which are chains of nucleotides made through the processes of DNA replication and transcription. Nucleoside triphosphates also serve as a source of energy for cellular reactions and are involved in signalling pathways. Nucleoside triphosphates cannot be absorbed well, so they are typically synthesized within the cell. Synthesis pathways differ depending on the specific nucleoside triphosphate being made, but given the many important roles of nucleoside triphosphates, synthesis is tightly regulated in all cases. Nucleoside analogues may also be used to treat viral infections. For example, azidothymidine (AZT) is a nucleoside analogue used to prevent and treat HIV/AIDS. Naming The term nucleoside refers to a nitrogenous base linked to a 5-carbon sugar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Taq Polymerase
''Taq'' polymerase is a thermostable DNA polymerase I named after the thermophilic eubacterial microorganism ''Thermus aquaticus,'' from which it was originally isolated by Chien et al. in 1976. Its name is often abbreviated to ''Taq'' or ''Taq'' pol. It is frequently used in the polymerase chain reaction (PCR), a method for greatly amplifying the quantity of short segments of DNA. ''T. aquaticus'' is a bacterium that lives in hot springs and hydrothermal vents, and ''Taq'' polymerase was identified as an enzyme able to withstand the protein-denaturing conditions (high temperature) required during PCR. Therefore, it replaced the DNA polymerase from '' E. coli'' originally used in PCR. Enzymatic properties ''Taqs optimum temperature for activity is 75–80 °C, with a half-life of greater than 2 hours at 92.5 °C, 40 minutes at 95 °C and 9 minutes at 97.5 °C, and can replicate a 1000 base pair strand of DNA in less than 10 seconds at 72 °C. At ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
PH Buffer
A buffer solution (more precisely, pH buffer or hydrogen ion buffer) is an aqueous solution consisting of a mixture of a weak acid and its conjugate base, or vice versa. Its pH changes very little when a small amount of strong acid or base is added to it. Buffer solutions are used as a means of keeping pH at a nearly constant value in a wide variety of chemical applications. In nature, there are many living systems that use buffering for pH regulation. For example, the bicarbonate buffering system is used to regulate the pH of blood, and bicarbonate also acts as a buffer in the ocean. Principles of buffering Buffer solutions resist pH change because of a chemical equilibrium between the weak acid HA and its conjugate base A−: When some strong acid is added to an equilibrium mixture of the weak acid and its conjugate base, hydrogen ions (H+) are added, and the equilibrium is shifted to the left, in accordance with Le Chatelier's principle. Because of this, the hydrogen i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nuclease
A nuclease (also archaically known as nucleodepolymerase or polynucleotidase) is an enzyme capable of cleaving the phosphodiester bonds between nucleotides of nucleic acids. Nucleases variously effect single and double stranded breaks in their target molecules. In living organisms, they are essential machinery for many aspects of DNA repair. Defects in certain nucleases can cause genetic instability or immunodeficiency. Nucleases are also extensively used in molecular cloning. There are two primary classifications based on the locus of activity. Exonucleases digest nucleic acids from the ends. Endonucleases act on regions in the ''middle'' of target molecules. They are further subcategorized as deoxyribonucleases and ribonucleases. The former acts on DNA, the latter on RNA. History In the late 1960s, scientists Stuart Linn and Werner Arber isolated examples of the two types of enzymes responsible for phage growth restriction in Escherichia coli ( E. coli) bacteria. One of the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
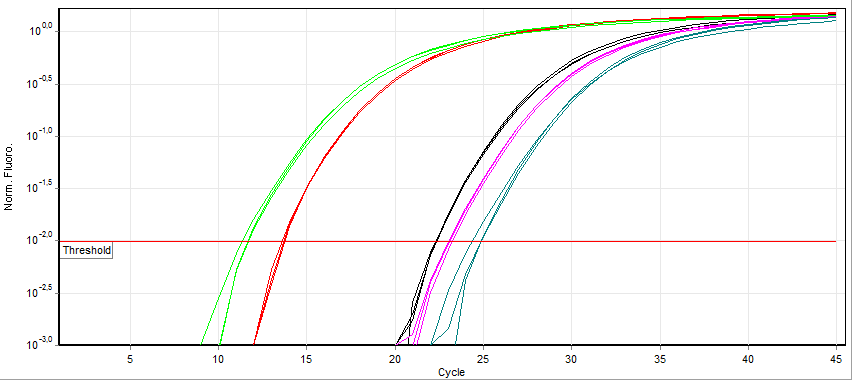
Real-time PCR
A real-time polymerase chain reaction (real-time PCR, or qPCR) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real time), not at its end, as in conventional PCR. Real-time PCR can be used quantitatively (quantitative real-time PCR) and semi-quantitatively (i.e., above/below a certain amount of DNA molecules) (semi-quantitative real-time PCR). Two common methods for the detection of PCR products in real-time PCR are (1) non-specific fluorescent dyes that intercalate with any double-stranded DNA and (2) sequence-specific DNA probes consisting of oligonucleotides that are labelled with a fluorescent reporter, which permits detection only after hybridization of the probe with its complementary sequence. The Minimum Information for Publication of Quantitative Real-Time PCR Experiments ( MIQE) guidelines propose that the abbreviation ''qPCR'' be used for q ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SYBR Green
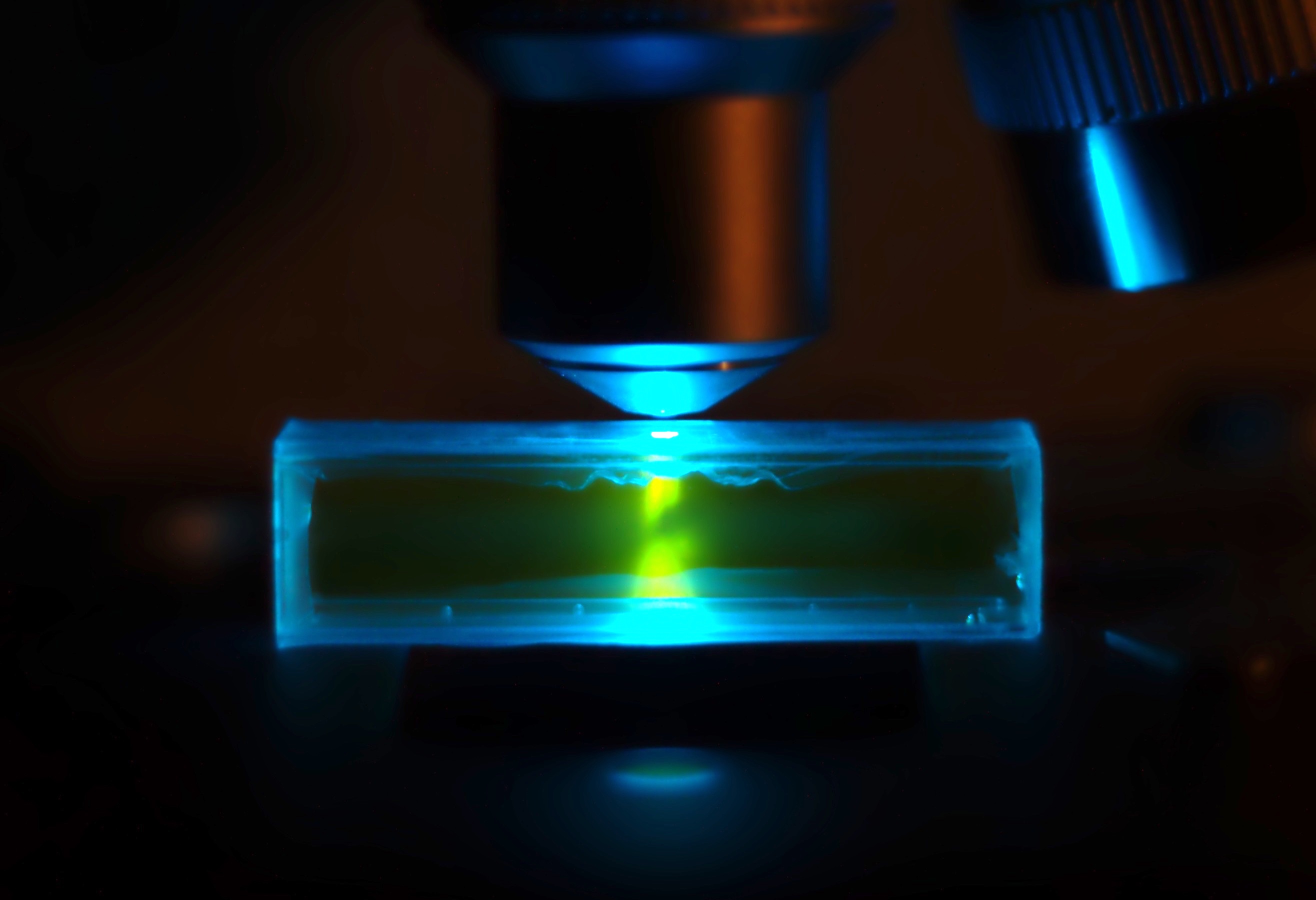
SYBR Green I (SG) is an asymmetrical cyanine dye used as a nucleic acid stain in molecular biology. The SYBR family of dyes is produced by Molecular Probes Inc., now owned by Thermo Fisher Scientific. SYBR Green I binds to DNA. The resulting DNA-dye-complex best absorbs 497 nanometer blue light (λmax = 497 nm) and emits green light (λmax = 520 nm). The stain preferentially binds to double-stranded DNA, but will stain single-stranded (ss) DNA with lower performance. SYBR Green can also stain RNA with a lower performance than ssDNA. Uses SYBR Green finds usage in several areas of biochemistry and molecular biology. It is used as a dye for the quantification of double stranded DNA in some methods of quantitative PCR. It is also used to visualise DNA in gel electrophoresis. Higher concentrations of SYBR Green can be used to stain agarose gels in order to visualise the DNA present. In addition to labelling pure nucleic acids, SYBR Green can also be used for labelling o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
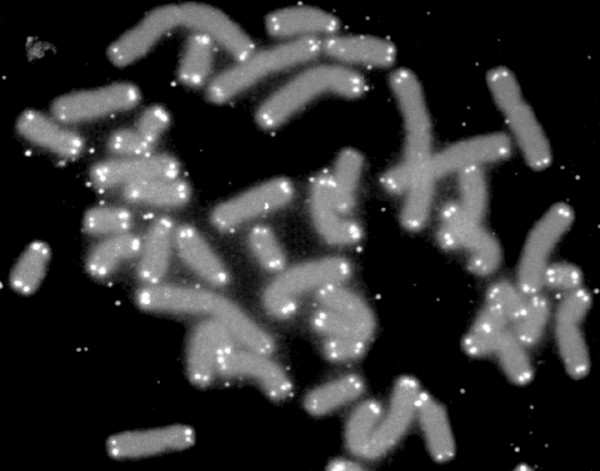
Telomere
A telomere (; ) is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Although there are different architectures, telomeres, in a broad sense, are a widespread genetic feature most commonly found in eukaryotes. In most, if not all species possessing them, they protect the terminal regions of chromosomal DNA from progressive degradation and ensure the integrity of linear chromosomes by preventing DNA repair systems from mistaking the very ends of the DNA strand for a double-strand break. Discovery In the early 1970s, Soviet theorist Alexei Olovnikov first recognized that chromosomes could not completely replicate their ends; this is known as the "end replication problem". Building on this, and accommodating Leonard Hayflick's idea of limited somatic cell division, Olovnikov suggested that DNA sequences are lost every time a cell replicates until the loss reaches a critical level, at which point cell division end ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |