|
Luteovirus
''Luteovirus'' is a genus of viruses, in the family ''Tombusviridae''. There are 13 species in this genus. Plants serve as natural hosts. The geographical distribution of Luteoviruses is widespread, with the virus primarily infecting plants via transmission by aphid vectors. The virus only replicates within the host cell and not within the vecto The name 'luteovirus' arises from the Latin ''luteus'', which is translated as 'yellow'. ''Luteovirus'' was given this name due to the symptomatic yellowing of the plant that occurs as a result of infection. Taxonomy The genus contains the following species: *'' Apple associated luteovirus'' *'' Apple luteovirus 1'' *'' Barley yellow dwarf virus kerII'' *'' Barley yellow dwarf virus kerIII'' *'' Barley yellow dwarf virus MAV'' *'' Barley yellow dwarf virus PAS'' *'' Barley yellow dwarf virus PAV'' *''Bean leafroll virus'' *'' Cherry associated luteovirus'' *'' Nectarine stem pitting associated virus'' *'' Red clover associated luteovirus' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bean Leafroll Virus
Bean leafroll virus (BLRV) is a plant pathogenic virus of the genus '' Luteovirus''. External linksICTVdB - The Universal Virus Database: Bean leaf roll virus Viral plant pathogens and diseases {{Virus-plant-disease-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beet Yellow Net Virus
Beet yellow net virus (BYNV) is a plant pathogenic virus of the family Luteoviridae ''Luteoviridae'' was a family of viruses. The family was abolished in 2020 based on evidence that its three genera and seven species unassigned to a genus belonged to two other, existing families. * The genus '' Enamovirus'' was reassigned to the .... External linksICTVdB - The Universal Virus Database: Beet yellow net virus Viral plant pathogens and diseases {{Virus-plant-disease-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tombusviridae
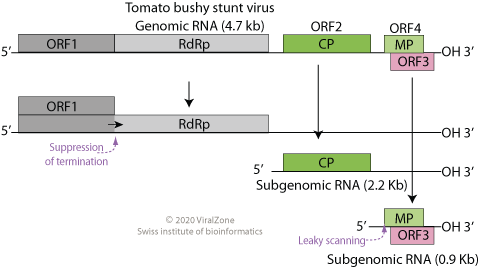
''Tombusviridae'' is a family of single-stranded positive sense RNA plant viruses. There are three subfamilies, 17 genera, and 95 species in this family. The name is derived from ''Tomato bushy stunt virus'' (TBSV). Genome All viruses in the family have a non-segmented (monopartite) linear genome, with the exception of Dianthoviruses, whose genome is bipartite. The genome is approximately 4.6–4.8kb in length, lacks a 5' cap and a poly(A) tail, and it encodes 4–6 ORFs. The polymerase encodes an amber stop codon which is the site of a readthrough event within ORF1, producing two products necessary for replication. There is no helicase encoded by the virus. ICTVFamily - Tombusviridae in: Virus Taxonomy. Ninth Report of the International Committee on Taxonomy of Viruses 2012, pp 1111-1138, 23 November 2011, doi:10.1016/B978-0-12-384684-6.00096-3 Structure The RNA is encapsulated in an icosahedral (T=3) capsid, composed of 180 units of a single coat protein 27–42K in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Soybean Dwarf Virus
Soybean dwarf virus (SbDV) is a pathogenic plant virus which infects soybeans. See also * List of soybean diseases Soybean plants (''Glycine max'') are subject to a variety of diseases and pests. Bacterial diseases Fungal diseases Nematodes, parasitic Viral diseases See also * Soybean management practices References Common Names of Diseases, T ... References External linksICTVdB - The Universal Virus Database: Soybean dwarf virus Viral plant pathogens and diseases Soybean diseases {{Virus-plant-disease-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polyadenylation
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature mRNA for translation. In many bacteria, the poly(A) tail promotes degradation of the mRNA. It, therefore, forms part of the larger process of gene expression. The process of polyadenylation begins as the transcription of a gene terminates. The 3′-most segment of the newly made pre-mRNA is first cleaved off by a set of proteins; these proteins then synthesize the poly(A) tail at the RNA's 3′ end. In some genes these proteins add a poly(A) tail at one of several possible sites. Therefore, polyadenylation can produce more than one transcript from a single gene (alternative polyadenylation), similar to alternative splicing. The poly(A) tail is important for the nuclea ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Upstream And Downstream (DNA)
In molecular biology and genetics, upstream and downstream both refer to relative positions of genetic code in DNA or RNA. Each strand of DNA or RNA has a 5' end and a 3' end, so named for the carbon position on the deoxyribose (or ribose) ring. By convention, upstream and downstream relate to the 5' to 3' direction respectively in which RNA transcription takes place. Upstream is toward the 5' end of the RNA molecule and downstream is toward the 3' end. When considering double-stranded DNA, upstream is toward the 5' end of the coding strand for the gene in question and downstream is toward the 3' end. Due to the anti-parallel nature of DNA, this means the 3' end of the template strand is upstream of the gene and the 5' end is downstream. Some genes on the same DNA molecule may be transcribed in opposite directions. This means the upstream and downstream areas of the molecule may change depending on which gene is used as the reference. See also *Upstream and downstream (transdu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Open Reading Frame
In molecular biology, open reading frames (ORFs) are defined as spans of DNA sequence between the start and stop codons. Usually, this is considered within a studied region of a prokaryotic DNA sequence, where only one of the six possible reading frames will be "open" (the "reading", however, refers to the RNA produced by transcription of the DNA and its subsequent interaction with the ribosome in translation). Such an ORF may contain a start codon (usually AUG in terms of RNA) and by definition cannot extend beyond a stop codon (usually UAA, UAG or UGA in RNA). That start codon (not necessarily the first) indicates where translation may start. The transcription termination site is located after the ORF, beyond the translation stop codon. If transcription were to cease before the stop codon, an incomplete protein would be made during translation. In eukaryotic genes with multiple exons, introns are removed and exons are then joined together after transcription to yield the final ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Directionality (molecular Biology)
Directionality, in molecular biology and biochemistry, is the end-to-end chemical orientation of a single strand of nucleic acid. In a single strand of DNA or RNA, the chemical convention of naming carbon atoms in the nucleotide pentose-sugar-ring means that there will be a 5′ end (usually pronounced "five-prime end"), which frequently contains a phosphate group attached to the 5′ carbon of the ribose ring, and a 3′ end (usually pronounced "three-prime end"), which typically is unmodified from the ribose -OH substituent. In a DNA double helix, the strands run in opposite directions to permit base pairing between them, which is essential for replication or transcription of the encoded information. Nucleic acids can only be synthesized in vivo in the 5′-to-3′ direction, as the polymerases that assemble various types of new strands generally rely on the energy produced by breaking nucleoside triphosphate bonds to attach new nucleoside monophosphates to the 3′- ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver. Nucleotides are composed of three subunit molecules: a nucleobase, a five-carbon sugar (ribose or deoxyribose), and a phosphate group consisting of one to three phosphates. The four nucleobases in DNA are guanine, adenine, cytosine and thymine; in RNA, uracil is used in place of thymine. Nucleotides also play a central role in metabolism at a fundamental, cellular level. They provide chemical energy—in the form of the nucleoside triphosphates, adenosine triphosphate (ATP), guanosine triphosphate (GTP), cytidine triphosphate (CTP) and uridine triphosphate (UTP)—throughout the cell for the many cellular func ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sense (molecular Biology)
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript. DNA sense Because of the complementary nature of base-pairing between nucleic acid polymers, a double-stranded DNA molecule will be composed of two strands with sequences that are reverse complements of each other. To help molecular biologists specifically identify each strand individually, the two strands are usually differentiated as the "sense" strand and the "antisense" strand. An individual strand of DNA is referred to as positive-sense (also positive (+) or simply sense) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Baltimore Classification
Baltimore classification is a system used to classify viruses based on their manner of messenger RNA (mRNA) synthesis. By organizing viruses based on their manner of mRNA production, it is possible to study viruses that behave similarly as a distinct group. Seven Baltimore groups are described that take into consideration whether the viral genome is made of deoxyribonucleic acid (DNA) or ribonucleic acid (RNA), whether the genome is single- or double-stranded, and whether the sense of a single-stranded RNA genome is positive or negative. Baltimore classification also closely corresponds to the manner of replicating the genome, so Baltimore classification is useful for grouping viruses together for both transcription and replication. Certain subjects pertaining to viruses are associated with multiple, specific Baltimore groups, such as specific forms of translation of mRNA and the host range of different types of viruses. Structural characteristics such as the shape of the vir ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleocapsid
A capsid is the protein shell of a virus, enclosing its genetic material. It consists of several oligomeric (repeating) structural subunits made of protein called protomers. The observable 3-dimensional morphological subunits, which may or may not correspond to individual proteins, are called capsomeres. The proteins making up the capsid are called capsid proteins or viral coat proteins (VCP). The capsid and inner genome is called the nucleocapsid. Capsids are broadly classified according to their structure. The majority of the viruses have capsids with either helical or icosahedral structure. Some viruses, such as bacteriophages, have developed more complicated structures due to constraints of elasticity and electrostatics. The icosahedral shape, which has 20 equilateral triangular faces, approximates a sphere, while the helical shape resembles the shape of a spring, taking the space of a cylinder but not being a cylinder itself. The capsid faces may consist of one or more ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |


.gif)