|
GeneMark
GeneMark is a generic name for a family of ab initio gene prediction programs developed at the Georgia Institute of Technology in Atlanta. Developed in 1993, original GeneMark was used in 1995 as a primary gene prediction tool for annotation of the first completely sequenced bacterial genome of ''Haemophilus influenzae'', and in 1996 for the first archaeal genome of '' Methanococcus jannaschii''. The algorithm introduced inhomogeneous three-periodic Markov chain models of protein-coding DNA sequence that became standard in gene prediction as well as Bayesian approach to gene prediction in two DNA strands simultaneously. Species specific parameters of the models were estimated from training sets of sequences of known type (protein-coding and non-coding). The major step of the algorithm computes for a given DNA fragment posterior probabilities of either being "protein-coding" (carrying genetic code) in each of six possible reading frames (including three frames in complementary D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Prediction
In computational biology, gene prediction or gene finding refers to the process of identifying the regions of genomic DNA that encode genes. This includes protein-coding genes as well as RNA genes, but may also include prediction of other functional elements such as regulatory regions. Gene finding is one of the first and most important steps in understanding the genome of a species once it has been sequenced. In its earliest days, "gene finding" was based on painstaking experimentation on living cells and organisms. Statistical analysis of the rates of homologous recombination of several different genes could determine their order on a certain chromosome, and information from many such experiments could be combined to create a genetic map specifying the rough location of known genes relative to each other. Today, with comprehensive genome sequence and powerful computational resources at the disposal of the research community, gene finding has been redefined as a largely computatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Prediction
In computational biology, gene prediction or gene finding refers to the process of identifying the regions of genomic DNA that encode genes. This includes protein-coding genes as well as RNA genes, but may also include prediction of other functional elements such as regulatory regions. Gene finding is one of the first and most important steps in understanding the genome of a species once it has been sequenced. In its earliest days, "gene finding" was based on painstaking experimentation on living cells and organisms. Statistical analysis of the rates of homologous recombination of several different genes could determine their order on a certain chromosome, and information from many such experiments could be combined to create a genetic map specifying the rough location of known genes relative to each other. Today, with comprehensive genome sequence and powerful computational resources at the disposal of the research community, gene finding has been redefined as a largely computatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
List Of Gene Prediction Software
This is a list of software tools and web portals used for gene prediction. See also * Gene prediction * List of RNA structure prediction software * Comparison of software for molecular mechanics modeling References {{Reflist, 2 Prediction A prediction (Latin ''præ-'', "before," and ''dicere'', "to say"), or forecast, is a statement about a future event or data. They are often, but not always, based upon experience or knowledge. There is no universal agreement about the exact ... gene prediction software ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Metagenomics Software
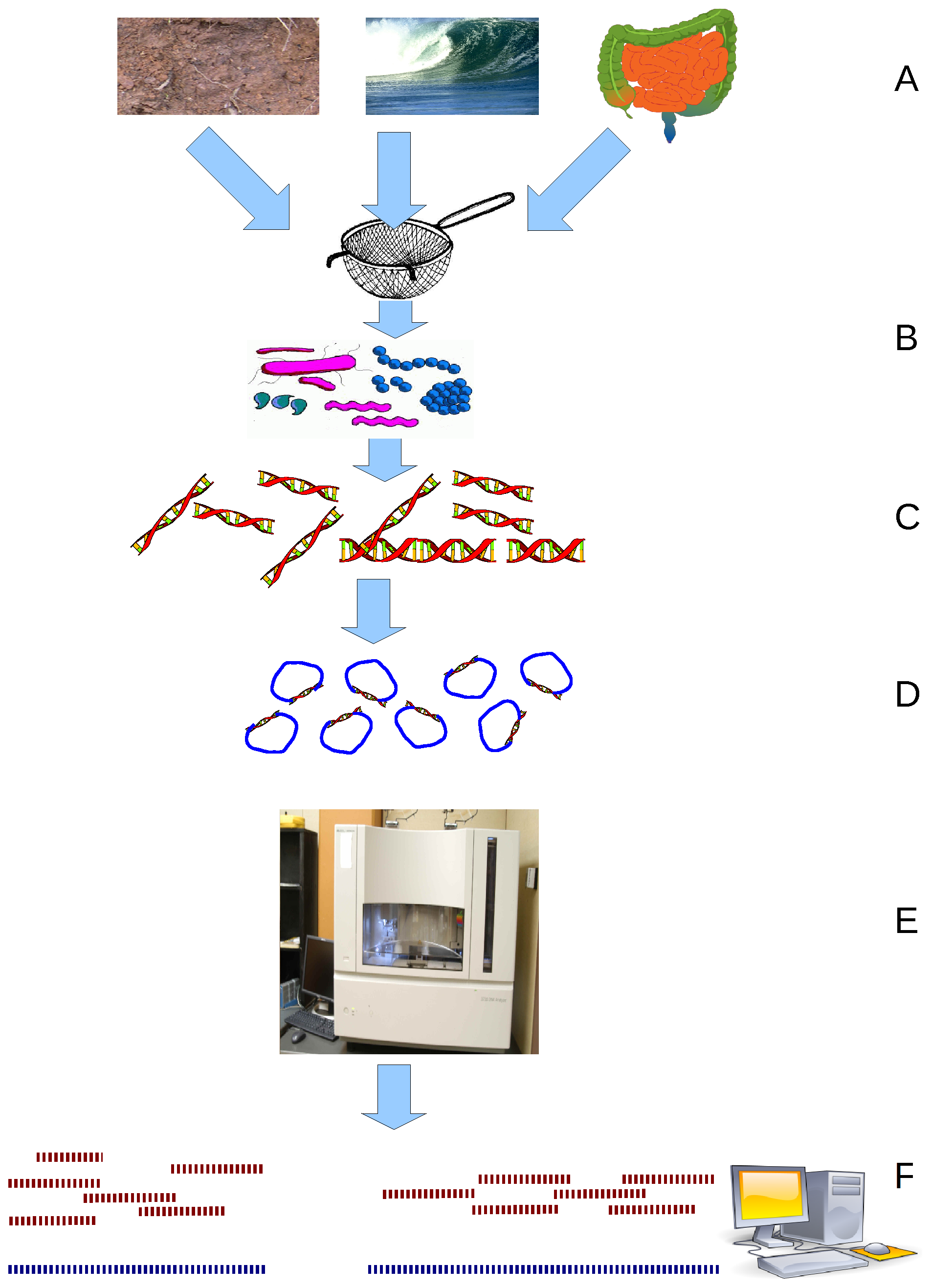
Metagenomics is the study of genetic material recovered directly from environmental or clinical samples by a method called sequencing. The broad field may also be referred to as environmental genomics, ecogenomics, community genomics or microbiomics. While traditional microbiology and microbial genome sequencing and genomics rely upon cultivated clonal cultures, early environmental gene sequencing cloned specific genes (often the 16S rRNA gene) to produce a profile of diversity in a natural sample. Such work revealed that the vast majority of microbial biodiversity had been missed by cultivation-based methods. Because of its ability to reveal the previously hidden diversity of microscopic life, metagenomics offers a powerful lens for viewing the microbial world that has the potential to revolutionize understanding of the entire living world. As the price of DNA sequencing continues to fall, metagenomics now allows microbial ecology to be investigated at a much greater scale ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Georgia Institute Of Technology
The Georgia Institute of Technology, commonly referred to as Georgia Tech or, in the state of Georgia, as Tech or The Institute, is a public research university and institute of technology in Atlanta, Georgia. Established in 1885, it is part of the University System of Georgia and has satellite campuses in Savannah, Georgia; Metz, France; Shenzhen, China; and Singapore. The school was founded as the Georgia School of Technology as part of Reconstruction plans to build an industrial economy in the post-Civil War Southern United States. Initially, it offered only a degree in mechanical engineering. By 1901, its curriculum had expanded to include electrical, civil, and chemical engineering. In 1948, the school changed its name to reflect its evolution from a trade school to a larger and more capable technical institute and research university. Today, Georgia Tech is organized into six colleges and contains about 31 departments/units, with emphasis on science and technology. I ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribosome
Ribosomes ( ) are macromolecular machines, found within all cells, that perform biological protein synthesis (mRNA translation). Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to form polypeptide chains. Ribosomes consist of two major components: the small and large ribosomal subunits. Each subunit consists of one or more ribosomal RNA (rRNA) molecules and many ribosomal proteins (RPs or r-proteins). The ribosomes and associated molecules are also known as the ''translational apparatus''. Overview The sequence of DNA that encodes the sequence of the amino acids in a protein is transcribed into a messenger RNA chain. Ribosomes bind to messenger RNAs and use their sequences for determining the correct sequence of amino acids to generate a given protein. Amino acids are selected and carried to the ribosome by transfer RNA (tRNA) molecules, which enter the ribosome and bind to the messenger RNA chain via an anti-c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genomics
Genomics is an interdisciplinary field of biology focusing on the structure, function, evolution, mapping, and editing of genomes. A genome is an organism's complete set of DNA, including all of its genes as well as its hierarchical, three-dimensional structural configuration. In contrast to genetics, which refers to the study of ''individual'' genes and their roles in inheritance, genomics aims at the collective characterization and quantification of ''all'' of an organism's genes, their interrelations and influence on the organism. Genes may direct the production of proteins with the assistance of enzymes and messenger molecules. In turn, proteins make up body structures such as organs and tissues as well as control chemical reactions and carry signals between cells. Genomics also involves the sequencing and analysis of genomes through uses of high throughput DNA sequencing and bioinformatics to assemble and analyze the function and structure of entire genomes. Advances in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mathematical And Theoretical Biology
Mathematical and theoretical biology, or biomathematics, is a branch of biology which employs theoretical analysis, mathematical models and abstractions of the living organisms to investigate the principles that govern the structure, development and behavior of the systems, as opposed to experimental biology which deals with the conduction of experiments to prove and validate the scientific theories. The field is sometimes called mathematical biology or biomathematics to stress the mathematical side, or theoretical biology to stress the biological side. Theoretical biology focuses more on the development of theoretical principles for biology while mathematical biology focuses on the use of mathematical tools to study biological systems, even though the two terms are sometimes interchanged. Mathematical biology aims at the mathematical representation and modeling of biological processes, using techniques and tools of applied mathematics. It can be useful in both theoretical and prac ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Intergenic Region
An intergenic region is a stretch of DNA sequences located between genes. Intergenic regions may contain functional elements and junk DNA. ''Inter''genic regions should not be confused with ''intra''genic regions (or introns), which are non-coding regions that are found ''within'' genes, especially within the genes of eukaryotic organisms. Properties and functions Intergenic regions may contain a number of functional DNA sequences such as promoters and regulatory elements, enhancers, spacers, and (in eukaryotes) centromeres. They may also contain origins of replication, scaffold attachment regions, and transposons and viruses. Non-functional DNA elements such as pseudogenes and repetitive DNA, both of which are types of junk DNA, can also be found in intergenic regions—although they may also be located within genes in introns. As all scientific knowledge is ultimately tentative—and in principle subject to revision given better evidence—it is possible s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Intron
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene... must be replaced by that of a transcription unit containing regions which will be lost from the mature messenger – which I suggest we call introns (for intragenic regions) – alternating with regions which will be expressed – exons." (Gilbert 1978) The term ''intron'' refers to both the DNA sequence within a gene and the corresponding RNA sequence in RNA transcripts. The non-intron sequences that become joined by this RNA processing to form the mature RNA are called exons. Introns are found in the genes of most organisms and many viruses and they can be located in both protein-coding genes and genes that function as RNA (noncoding genes). There are four main types of introns: tRNA introns, group I introns, group II introns, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exon
An exon is any part of a gene that will form a part of the final mature RNA produced by that gene after introns have been removed by RNA splicing. The term ''exon'' refers to both the DNA sequence within a gene and to the corresponding sequence in RNA transcripts. In RNA splicing, introns are removed and exons are covalently joined to one another as part of generating the mature RNA. Just as the entire set of genes for a species constitutes the genome, the entire set of exons constitutes the exome. History The term ''exon'' derives from the expressed region and was coined by American biochemist Walter Gilbert in 1978: "The notion of the cistron… must be replaced by that of a transcription unit containing regions which will be lost from the mature messengerwhich I suggest we call introns (for intragenic regions)alternating with regions which will be expressedexons." This definition was originally made for protein-coding transcripts that are spliced before being translated. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Binding Site
In biochemistry and molecular biology, a binding site is a region on a macromolecule such as a protein that binds to another molecule with specificity. The binding partner of the macromolecule is often referred to as a ligand. Ligands may include other proteins (resulting in a protein-protein interaction), enzyme substrates, second messengers, hormones, or allosteric modulators. The binding event is often, but not always, accompanied by a conformational change that alters the protein's function. Binding to protein binding sites is most often reversible (transient and non-covalent), but can also be covalent reversible or irreversible. Function Binding of a ligand to a binding site on protein often triggers a change in conformation in the protein and results in altered cellular function. Hence binding site on protein are critical parts of signal transduction pathways. Types of ligands include neurotransmitters, toxins, neuropeptides, and steroid hormones. Binding sites in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |