|
Functional Genomics
Functional genomics is a field of molecular biology that attempts to describe gene (and protein) functions and interactions. Functional genomics make use of the vast data generated by genomic and transcriptomic projects (such as genome sequencing projects and RNA sequencing). Functional genomics focuses on the dynamic aspects such as gene transcription, translation, regulation of gene expression and protein–protein interactions, as opposed to the static aspects of the genomic information such as DNA sequence or structures. A key characteristic of functional genomics studies is their genome-wide approach to these questions, generally involving high-throughput methods rather than a more traditional "gene-by-gene" approach. Definition and goals of functional genomics In order to understand functional genomics it is important to first define function. In their paper Graur et al. define function in two possible ways. These are "selected effect" and "causal role". The "selected ef ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology. Molecular biology was first described as an approach focused on the underpinnings of biological phenomena - uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology. In 1945 the term molecular biology was used by physicist William Astbury. In 1953 Francis Crick, James Watson, Rosalind Franklin, and colleagues, working at Medical Research Council unit, Cavendish laboratory, Cambridge (now the MRC Laboratory of Molecular Biology), made a double helix model of DNA which changed the entire research scenario. They proposed the DNA structure based on previous research done by Ro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Omics
The branches of science known informally as omics are various disciplines in biology whose names end in the suffix '' -omics'', such as genomics, proteomics, metabolomics, metagenomics, phenomics and transcriptomics. Omics aims at the collective characterization and quantification of pools of biological molecules that translate into the structure, function, and dynamics of an organism or organisms . The related suffix -ome is used to address the objects of study of such fields, such as the genome, proteome or metabolome respectively. The suffix ''-ome'' as used in molecular biology refers to a ''totality'' of some sort; it is an example of a "neo-suffix" formed by abstraction from various Greek terms in , a sequence that does not form an identifiable suffix in Greek. Functional genomics aims at identifying the functions of as many genes as possible of a given organism. It combines different -omics techniques such as transcriptomics and proteomics with saturated mutant collec ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNase-Seq
DNase-seq (DNase I hypersensitive sites sequencing) is a method in molecular biology used to identify the location of regulatory regions, based on the genome-wide sequencing of regions sensitive to cleavage by DNase I. FAIRE-Seq is a successor of DNase-seq for the genome-wide identification of accessible DNA regions in the genome. Both the protocols for identifying open chromatin regions have biases depending on underlying nucleosome structure. For example, FAIRE-seq provides higher tag counts at non-promoter regions. On the other hand, DNase-seq signal is higher at promoter regions, and DNase-seq has been shown to have better sensitivity than FAIRE-seq even at non-promoter regions. DNase-seq Footprinting DNase-seq requires some downstream bioinformatics analyses in order to provide genome-wide DNA footprints. The computational tools proposed can be categorized in two classes: segmentation-based and site-centric approaches. Segmentation-based methods are based on the application ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ATAC-seq
ATAC-seq (Assay for Transposase-Accessible Chromatin using sequencing) is a technique used in molecular biology to assess genome-wide chromatin accessibility. In 2013, the technique was first described as an alternative advanced method for MNase-seq, FAIRE-Seq and DNase-Seq. ATAC-seq is a faster and more sensitive analysis of the epigenome than DNase-seq or MNase-seq. Description ATAC-seq identifies accessible DNA regions by probing open chromatin with hyperactive mutant Tn5 Transposase that inserts sequencing adapters into open regions of the genome. While naturally occurring transposases have a low level of activity, ATAC-seq employs the mutated hyperactive transposase. In a process called "tagmentation", Tn5 transposase cleaves and tags double-stranded DNA with sequencing adaptors. The tagged DNA fragments are then purified, PCR-amplified, and sequenced using next-generation sequencing. Sequencing reads can then be used to infer regions of increased accessibility as well as to ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CUT&RUN Sequencing
CUT&RUN sequencing, also known as cleavage under targets and release using nuclease, is a method used to analyze protein interactions with DNA. CUT&RUN sequencing combines antibody-targeted controlled cleavage by micrococcal nuclease with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins. It can be used to map global DNA binding sites precisely for any protein of interest. Currently, ChIP-Seq is the most common technique utilized to study protein–DNA relations, however, it suffers from a number of practical and economical limitations that CUT&RUN sequencing does not. Uses CUT&RUN sequencing can be used to examine gene regulation or to analyze transcription factor and other chromatin-associated protein binding. Protein-DNA interactions regulate gene expression and are responsible for many biological processes and disease states. This epigenetic information is complementary to genotype and expression analysis. CUT&RUN is an alternative to th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ChIP-sequencing
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins. It can be used to map global binding sites precisely for any protein of interest. Previously, ChIP-on-chip was the most common technique utilized to study these protein–DNA relations. Uses ChIP-seq is primarily used to determine how transcription factors and other chromatin-associated proteins influence phenotype-affecting mechanisms. Determining how proteins interact with DNA to regulate gene expression is essential for fully understanding many biological processes and disease states. This epigenetic information is complementary to genotype and expression analysis. ChIP-seq technology is currently seen primarily as an alternative to ChIP-chip which requires a hybridization array. This introduces some bias, as an array is restrict ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Epistasis
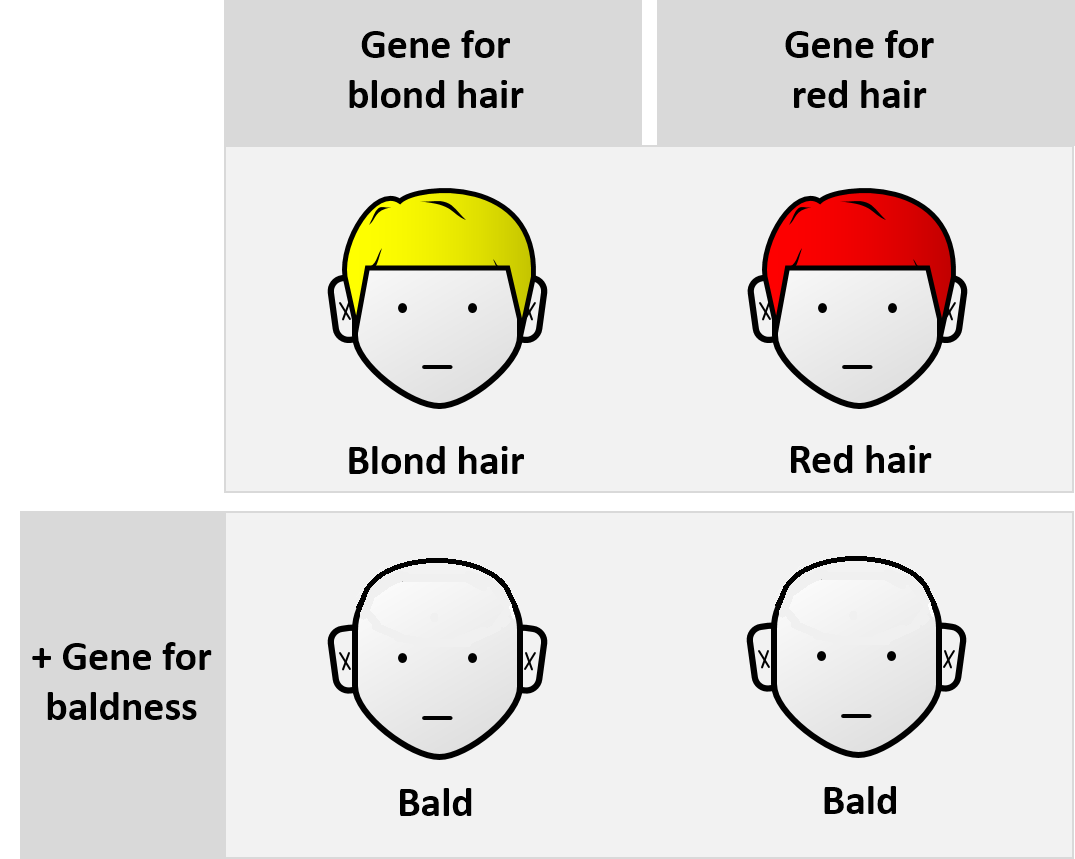
Epistasis is a phenomenon in genetics in which the effect of a gene mutation is dependent on the presence or absence of mutations in one or more other genes, respectively termed modifier genes. In other words, the effect of the mutation is dependent on the genetic background in which it appears. Epistatic mutations therefore have different effects on their own than when they occur together. Originally, the term ''epistasis'' specifically meant that the effect of a gene variant is masked by that of a different gene. The concept of ''epistasis'' originated in genetics in 1907 but is now used in biochemistry, computational biology and evolutionary biology. The phenomenon arises due to interactions, either between genes (such as mutations also being needed in regulators of gene expression) or within them (multiple mutations being needed before the gene loses function), leading to non-linear effects. Epistasis has a great influence on the shape of evolutionary landscapes, which l ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biological Specimen
A biological specimen (also called a biospecimen) is a biological laboratory specimen held by a biorepository for research. Such a specimen would be taken by sampling so as to be representative of any other specimen taken from the source of the specimen. When biological specimens are stored, ideally they remain equivalent to freshly-collected specimens for the purposes of research. Human biological specimens are stored in a type of biorepository called a biobank, and the science of preserving biological specimens is most active in the field of biobanking. Quality control Setting broad standards for quality of biological specimens was initially an underdeveloped aspect of biobank growth. There is currently discussion on what standards should be in place and who should manage those standards. Since many organizations set their own standards and since biobanks are necessarily used by multiple organizations and typically are driven towards expansion, the harmonization of standard ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genetic inf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Multiplex (assay)
In the biological sciences, a multiplex assay is a type of immunoassay that uses magnetic beads to simultaneously measure multiple analytes in a single experiment. A multiplex assay is a derivative of an ELISA using beads for binding the capture antibody. Multiplex assays are still more common in research than in clinical settings. In a multiplex assay, microspheres of designated colors are coated with antibodies of defined binding specificities. The results can be read by flow cytometry Flow cytometry (FC) is a technique used to detect and measure physical and chemical characteristics of a population of cells or particles. In this process, a sample containing cells or particles is suspended in a fluid and injected into the flo ... because the beads are distinguishable by fluorescent signature. The number of analytes measured is determined by the number of different bead colors. Multiplex assays within a given application area or class of technology can be further stratified ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Metabolomics
Metabolomics is the scientific study of chemical processes involving metabolites, the small molecule substrates, intermediates, and products of cell metabolism. Specifically, metabolomics is the "systematic study of the unique chemical fingerprints that specific cellular processes leave behind", the study of their small-molecule metabolite profiles. The metabolome represents the complete set of metabolites in a biological cell, tissue, organ, or organism, which are the end products of cellular processes. Messenger RNA (mRNA), gene expression data, and proteomics, proteomic analyses reveal the set of gene products being produced in the cell, data that represents one aspect of cellular function. Conversely, metabolic profiling can give an instantaneous snapshot of the physiology of that cell, and thus, metabolomics provides a direct "functional readout of the physiological state" of an organism. There are indeed quantifiable correlations between the metabolome and the other cellular ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Production
Protein production is the biotechnological process of generating a specific protein. It is typically achieved by the manipulation of gene expression in an organism such that it expresses large amounts of a recombinant gene. This includes the transcription of the recombinant DNA to messenger RNA (mRNA), the translation of mRNA into polypeptide chains, which are ultimately folded into functional proteins and may be targeted to specific subcellular or extracellular locations. Protein production systems (also known as expression systems) are used in the life sciences, biotechnology, and medicine. Molecular biology research uses numerous proteins and enzymes, many of which are from expression systems; particularly DNA polymerase for PCR, reverse transcriptase for RNA analysis, restriction endonucleases for cloning, and to make proteins that are screened in drug discovery as biological targets or as potential drugs themselves. There are also significant applications for expressi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |