|
Atg1
AuTophaGy related 1 (Atg1) is a 101.7kDa serine/threonine kinase in ''S.cerevisiae'', encoded by the gene ATG1. It is essential for the initial building of the autophagosome and Cvt vesicles. In a non-kinase role it is - through complex formation with Atg13 and Atg17 - directly controlled by the TOR kinase, a sensor for nutrient availability. Introduction Atg1 can associate with a number of other proteins of the Atg family to form a complex that functions in autophagosome or Cvt vesicle formation. The initiation of autophagy involves the building of the pre-autophagosomal structure (PAS). Most Atg proteins accumulate at the PAS and generate either Cvt vesicles under normal growing conditions or autophagosomes under starvation. To date, there are 31 ATG genes, which can be classified into several different groups according to their functions at the different steps of the pathway. 17 of these genes only work in the Cvt pathway. Structure The Atg1 gene lies on chromosome VII of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Atg1 Interaction Partners
AuTophaGy related 1 (Atg1) is a 101.7kDa serine/threonine kinase in ''S.cerevisiae'', encoded by the gene ATG1. It is essential for the initial building of the autophagosome and Cvt vesicles. In a non-kinase role it is - through complex formation with Atg13 and Atg17 - directly controlled by the TOR kinase, a sensor for nutrient availability. Introduction Atg1 can associate with a number of other proteins of the Atg family to form a complex that functions in autophagosome or Cvt vesicle formation. The initiation of autophagy involves the building of the pre-autophagosomal structure (PAS). Most Atg proteins accumulate at the PAS and generate either Cvt vesicles under normal growing conditions or autophagosomes under starvation. To date, there are 31 ATG genes, which can be classified into several different groups according to their functions at the different steps of the pathway. 17 of these genes only work in the Cvt pathway. Structure The Atg1 gene lies on chromosome VII of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Autophagosome
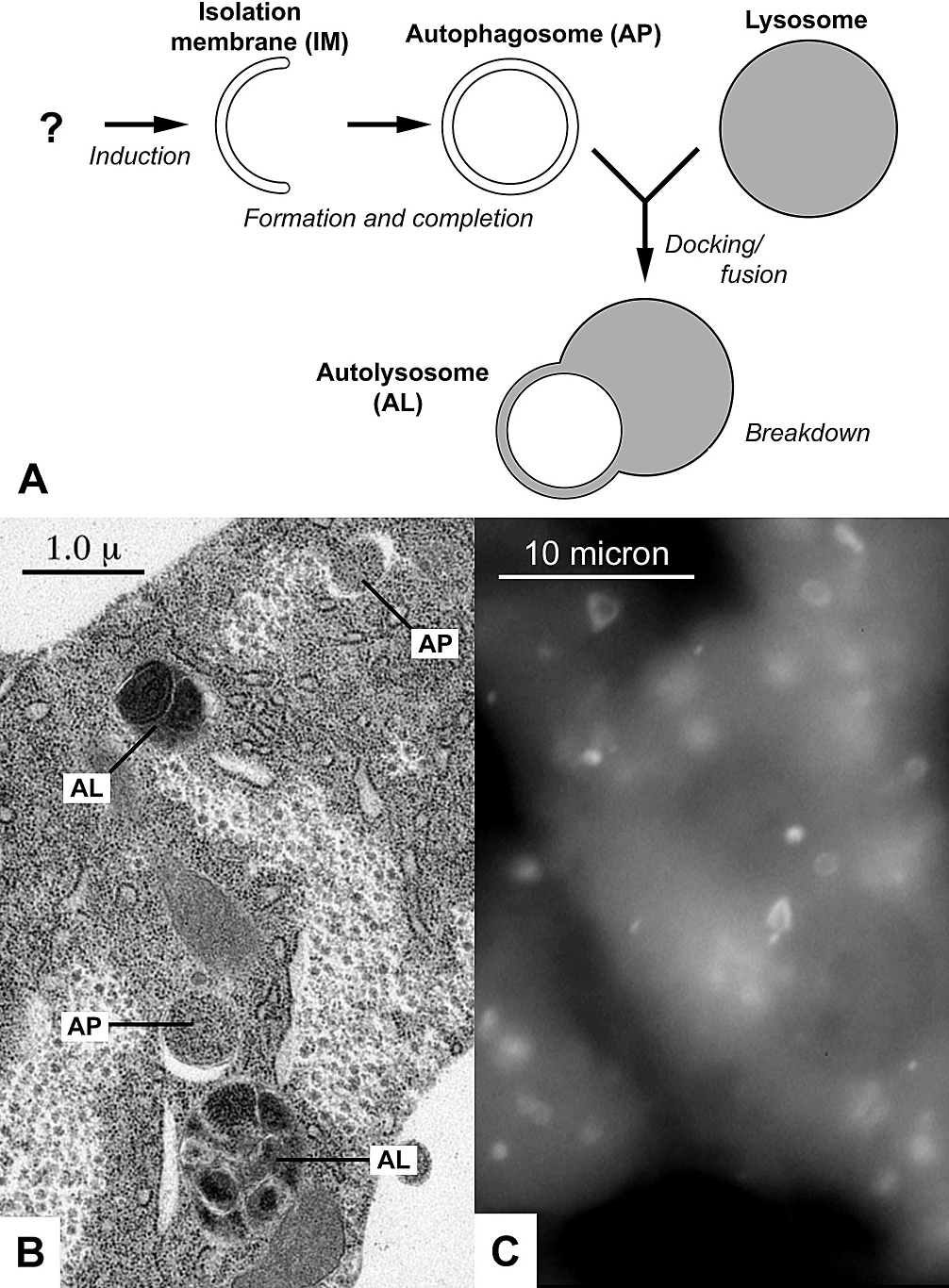
An autophagosome is a spherical structure with double layer membranes. It is the key structure in macroautophagy, the intracellular degradation system for cytoplasmic contents (e.g., abnormal intracellular proteins, excess or damaged organelles, invading microorganisms). After formation, autophagosomes deliver cytoplasmic components to the lysosomes. The outer membrane of an autophagosome fuses with a lysosome to form an autolysosome. The lysosome's hydrolases degrade the autophagosome-delivered contents and its inner membrane. The formation of autophagosomes is regulated by genes that are well-conserved from yeast to higher eukaryotes. The nomenclature of these genes has differed from paper to paper, but it has been simplified in recent years. The gene families formerly known as APG, AUT, CVT, GSA, PAZ, and PDD are now unified as the ATG (AuTophaGy related) family. The size of autophagosomes vary between mammals and yeast. Yeast autophagosomes are about 500-900 nm, while mam ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Autophagy
Autophagy (or autophagocytosis; from the Ancient Greek , , meaning "self-devouring" and , , meaning "hollow") is the natural, conserved degradation of the cell that removes unnecessary or dysfunctional components through a lysosome-dependent regulated mechanism. It allows the orderly degradation and recycling of cellular components. Although initially characterized as a primordial degradation pathway induced to protect against starvation, it has become increasingly clear that autophagy also plays a major role in the homeostasis of non-starved cells. Defects in autophagy have been linked to various human diseases, including neurodegeneration and cancer, and interest in modulating autophagy as a potential treatment for these diseases has grown rapidly. Four forms of autophagy have been identified: macroautophagy, microautophagy, chaperone-mediated autophagy (CMA), and crinophagy. In macroautophagy (the most thoroughly researched form of autophagy), cytoplasmic components (like mit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TOR (gene)
The mammalian target of sirolimus, rapamycin (mTOR), also referred to as the mechanistic target of rapamycin, and sometimes called FK506-binding protein 12-rapamycin-associated protein 1 (FRAP1), is a kinase that in humans is encoded by the ''MTOR'' gene. mTOR is a member of the phosphatidylinositol 3-kinase-related kinase family of protein kinases. mTOR links with other proteins and serves as a core component of two distinct protein complexes, mTORC1, mTOR complex 1 and mTORC2, mTOR complex 2, which regulate different cellular processes. In particular, as a core component of both complexes, mTOR functions as a serine/threonine protein kinase that regulates cell growth, cell proliferation, cell motility, cell survival, protein synthesis, autophagy, and Transcription (genetics), transcription. As a core component of mTORC2, mTOR also functions as a tyrosine protein kinase that promotes the activation of insulin receptors and insulin-like growth factor 1 receptors. mTORC2 has also ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ULK1
ULK1 is an enzyme that in humans is encoded by the ''ULK1'' gene. Unc-51 like autophagy activating kinase (ULK1/2) are two similar isoforms of an enzyme that in humans are encoded by the ''ULK1/2'' genes. .html" ;"title="/sup>">/sup> .html" ;"title="/sup>">/sup> It is specifically a kinase that is involved with autophagy, particularly in response to amino acid withdrawal. Not many studies have been done comparing the two isoforms, but some differences have been recorded. Function Ulk1/2 is an important protein in autophagy for mammalian cells, and is homologous to ATG1 in yeast. It is part of the ULK1-complex, which is needed in early steps of autophagosome biogenesis. The ULK1 complex also consists of the FAK family kinase interacting protein of 200 kDa (FIP200 or RB1CC1) and the HORMA (Hop/Rev7/Mad2) domain-containing proteins ATG13 and ATG101. ULK1, specifically, appears to be the most essential for autophagy and is activated under conditions of nutrient deprivation by sev ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell Death
Cell death is the event of a biological cell ceasing to carry out its functions. This may be the result of the natural process of old cells dying and being replaced by new ones, as in programmed cell death, or may result from factors such as diseases, localized injury, or the death of the organism of which the cells are part. Apoptosis or Type I cell-death, and autophagy or Type II cell-death are both forms of programmed cell death, while necrosis is a non-physiological process that occurs as a result of infection or injury. Programmed cell death Programmed cell death (PCD) is cell death mediated by an intracellular program. PCD is carried out in a regulated process, which usually confers advantage during an organism's life-cycle. For example, the differentiation of fingers and toes in a developing human embryo occurs because cells between the fingers apoptose; the result is that the digits separate. PCD serves fundamental functions during both plant and metazoa (multicellula ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homology (biology)
In biology, homology is similarity due to shared ancestry between a pair of structures or genes in different taxa. A common example of homologous structures is the forelimbs of vertebrates, where the wings of bats and birds, the arms of primates, the front flippers of whales and the forelegs of four-legged vertebrates like dogs and crocodiles are all derived from the same ancestral tetrapod structure. Evolutionary biology explains homologous structures adapted to different purposes as the result of descent with modification from a common ancestor. The term was first applied to biology in a non-evolutionary context by the anatomist Richard Owen in 1843. Homology was later explained by Charles Darwin's theory of evolution in 1859, but had been observed before this, from Aristotle onwards, and it was explicitly analysed by Pierre Belon in 1555. In developmental biology, organs that developed in the embryo in the same manner and from similar origins, such as from matching p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Kinase A
In cell biology, protein kinase A (PKA) is a family of enzymes whose activity is dependent on cellular levels of cyclic AMP (cAMP). PKA is also known as cAMP-dependent protein kinase (). PKA has several functions in the cell, including regulation of glycogen, sugar, and lipid metabolism. It should not be confused with 5'-AMP-activated protein kinase (AMP-activated protein kinase). History Protein kinase A, more precisely known as adenosine 3',5'-monophosphate (cyclic AMP)-dependent protein kinase, abbreviated to PKA, was discovered by chemists Edmond H. Fischer and Edwin G. Krebs in 1968. They won the Nobel Prize in Physiology or Medicine in 1992 for their work on phosphorylation and dephosphorylation and how it relates to PKA activity. PKA is one of the most widely researched protein kinases, in part because of its uniqueness; out of 540 different protein kinase genes that make up the human kinome, only one other protein kinase, casein kinase 2, is known to exist in a physio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|