|
Regulator Gene
In genetics, a regulator gene, regulator, or regulatory gene is a gene involved in controlling the expression of one or more other genes. Regulatory sequences, which encode regulatory genes, are often at the five prime end (5') to the start site of transcription of the gene they regulate. In addition, these sequences can also be found at the three prime end (3') to the transcription start site. In both cases, whether the regulatory sequence occurs before (5') or after (3') the gene it regulates, the sequence is often many kilobases away from the transcription start site. A regulator gene may encode a protein, or it may work at the level of RNA, as in the case of genes encoding microRNAs. An example of a regulator gene is a gene that codes for a repressor protein that inhibits the activity of an operator (a gene which binds repressor proteins thus inhibiting the translation of RNA to protein via RNA polymerase). In prokaryotes, regulator genes often code for repressor pro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Regulatory Network
A gene (or genetic) regulatory network (GRN) is a collection of molecular regulators that interact with each other and with other substances in the cell to govern the gene expression levels of mRNA and proteins which, in turn, determine the function of the cell. GRN also play a central role in morphogenesis, the creation of body structures, which in turn is central to evolutionary developmental biology (evo-devo). The regulator can be DNA, RNA, protein or any combination of two or more of these three that form a complex, such as a specific sequence of DNA and a transcription factor to activate that sequence. The interaction can be direct or indirect (through transcribed RNA or translated protein). In general, each mRNA molecule goes on to make a specific protein (or set of proteins). In some cases this protein will be structural, and will accumulate at the cell membrane or within the cell to give it particular structural properties. In other cases the protein will be an enz ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Promoter (biology)
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription (genetics), transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, Upstream and downstream (DNA), upstream on the DNA (towards the Directionality (molecular biology)#5′-end, 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism. Overview For transcription to take place, the enzyme that synthesizes RNA, known as RNA polymerase, must attach to the DNA near a gene. Promoters contain specific DNA sequences such as response elements that provide a secure initial binding site for RNA polymerase and for ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Silencer (DNA)
In genetics, a silencer is a DNA sequence capable of binding transcription factors, transcription regulation factors, called repressors. DNA contains genes and provides the template to produce messenger RNA (mRNA). That mRNA is then Translation (biology), translated into proteins. When a repressor protein binds to the silencer region of DNA, RNA polymerase is prevented from Transcription (genetics), transcribing the DNA sequence into RNA. With transcription blocked, the translation of RNA into proteins is impossible. Thus, silencers prevent genes from being Gene expression, expressed as proteins. RNA polymerase, a DNA-dependent enzyme, transcribes the DNA sequences, called nucleotides, in the directionality (molecular biology), 3' to 5' direction while the complementary RNA is synthesized in the 5' to 3' direction. RNA is similar to DNA, except that RNA contains uracil, instead of thymine, which forms a base pair with adenine. An important region for the activity of gene repress ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enhancer (genetics)
In genetics, an enhancer is a short (50–1500 bp) region of DNA that can be bound by proteins ( activators) to increase the likelihood that transcription of a particular gene will occur. These proteins are usually referred to as transcription factors. Enhancers are ''cis''-acting. They can be located up to 1 Mbp (1,000,000 bp) away from the gene, upstream or downstream from the start site. There are hundreds of thousands of enhancers in the human genome. They are found in both prokaryotes and eukaryotes. Active enhancers typically get transcribed as enhancer or regulatory non-coding RNA, whose expression levels correlate with mRNA levels of target genes. The first discovery of a eukaryotic enhancer was in the immunoglobulin heavy chain gene in 1983. This enhancer, located in the large intron, provided an explanation for the transcriptional activation of rearranged Vh gene promoters while unrearranged Vh promoters remained inactive. Lately, enhancers have been shown to be in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcription (genetics)
Transcription is the process of copying a segment of DNA into RNA for the purpose of gene expression. Some segments of DNA are transcribed into RNA molecules that can encode proteins, called messenger RNA (mRNA). Other segments of DNA are transcribed into RNA molecules called non-coding RNAs (ncRNAs). Both DNA and RNA are nucleic acids, which use base pairs of nucleotides as a complementary language. During transcription, a DNA sequence is read by an RNA polymerase, which produces a complementary, antiparallel RNA strand called a primary transcript. In virology, the term transcription is used when referring to mRNA synthesis from a viral RNA molecule. The genome of many RNA viruses is composed of negative-sense RNA which acts as a template for positive sense viral messenger RNA - a necessary step in the synthesis of viral proteins needed for viral replication. This process is catalyzed by a viral RNA dependent RNA polymerase. Background A DNA transcription unit encoding ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Promoter (genetics)
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism. Overview For transcription to take place, the enzyme that synthesizes RNA, known as RNA polymerase, must attach to the DNA near a gene. Promoters contain specific DNA sequences such as response elements that provide a secure initial binding site for RNA polymerase and for proteins called transcription factors that recruit RNA polymerase. These transcription factor ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Evolutionary Developmental Biology
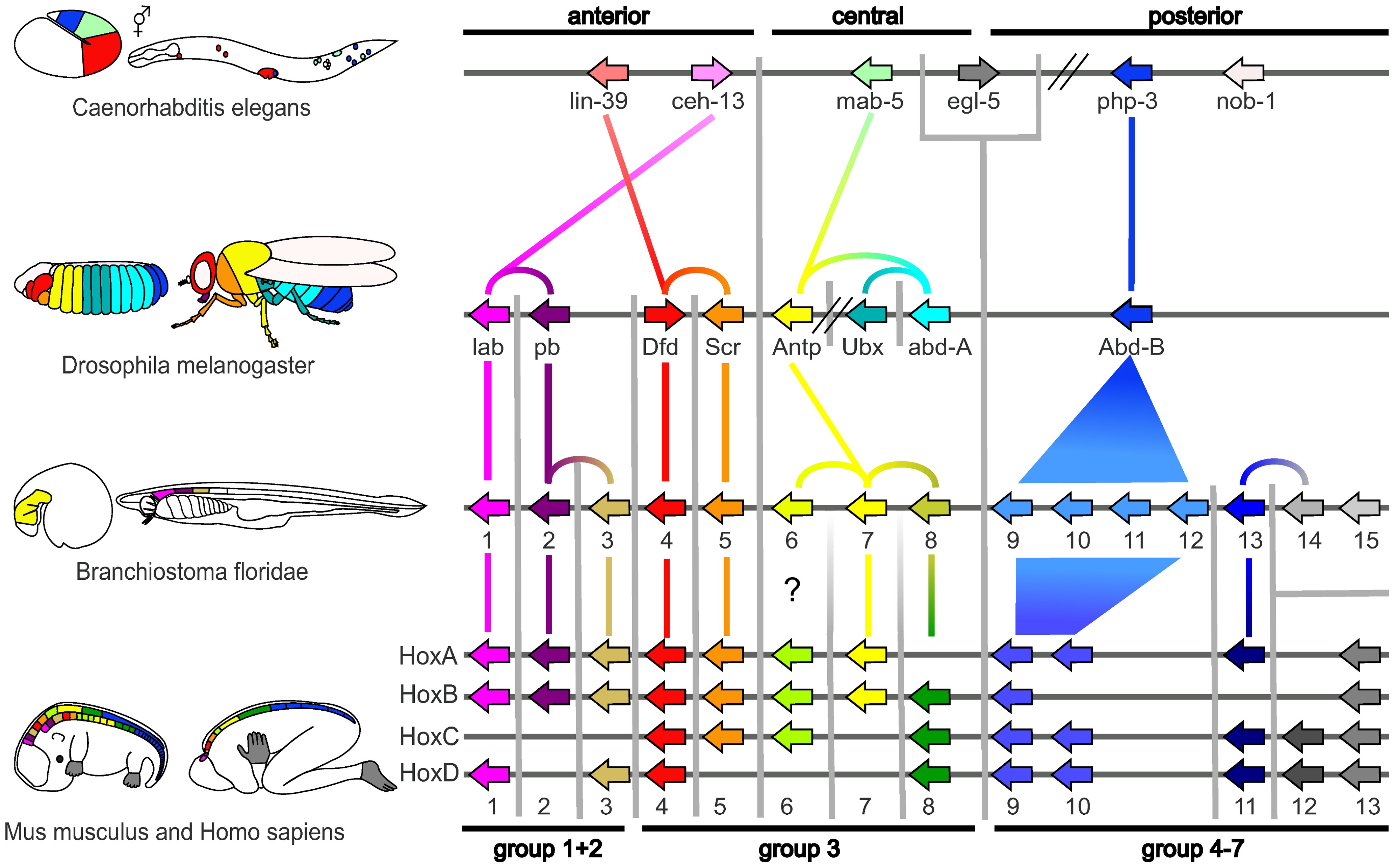
Evolutionary developmental biology, informally known as evo-devo, is a field of biological research that compares the developmental biology, developmental processes of different organisms to infer how developmental processes evolution, evolved. The field grew from 19th-century beginnings, where embryology faced a mystery: zoology, zoologists did not know how embryogenesis, embryonic development was controlled at the molecular level. Charles Darwin noted that having similar embryos implied common ancestry, but little progress was made until the 1970s. Then, recombinant DNA technology at last brought embryology together with molecular genetics. A key early discovery was that of homeotic genes that regulate development in a wide range of eukaryotes. The field is composed of multiple core evolutionary concepts. One is deep homology, the finding that dissimilar organs such as the eyes of insects, vertebrates and cephalopod molluscs, long thought to have evolved separately, are contr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Regulation Of Gene Expression
Regulation of gene expression, or gene regulation, includes a wide range of mechanisms that are used by cells to increase or decrease the production of specific gene products (protein or RNA). Sophisticated programs of gene expression are widely observed in biology, for example to trigger developmental pathways, respond to environmental stimuli, or adapt to new food sources. Virtually any step of gene expression can be modulated, from Transcriptional regulation, transcriptional initiation, to RNA processing, and to the post-translational modification of a protein. Often, one gene regulator controls another, and so on, in a gene regulatory network. Gene regulation is essential for viruses, prokaryotes and eukaryotes as it increases the versatility and adaptability of an organism by allowing the cell to express protein when needed. Although as early as 1951, Barbara McClintock showed interaction between two genetic loci, Activator (''Ac'') and Dissociator (''Ds''), in the color f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Catabolite Activator Protein
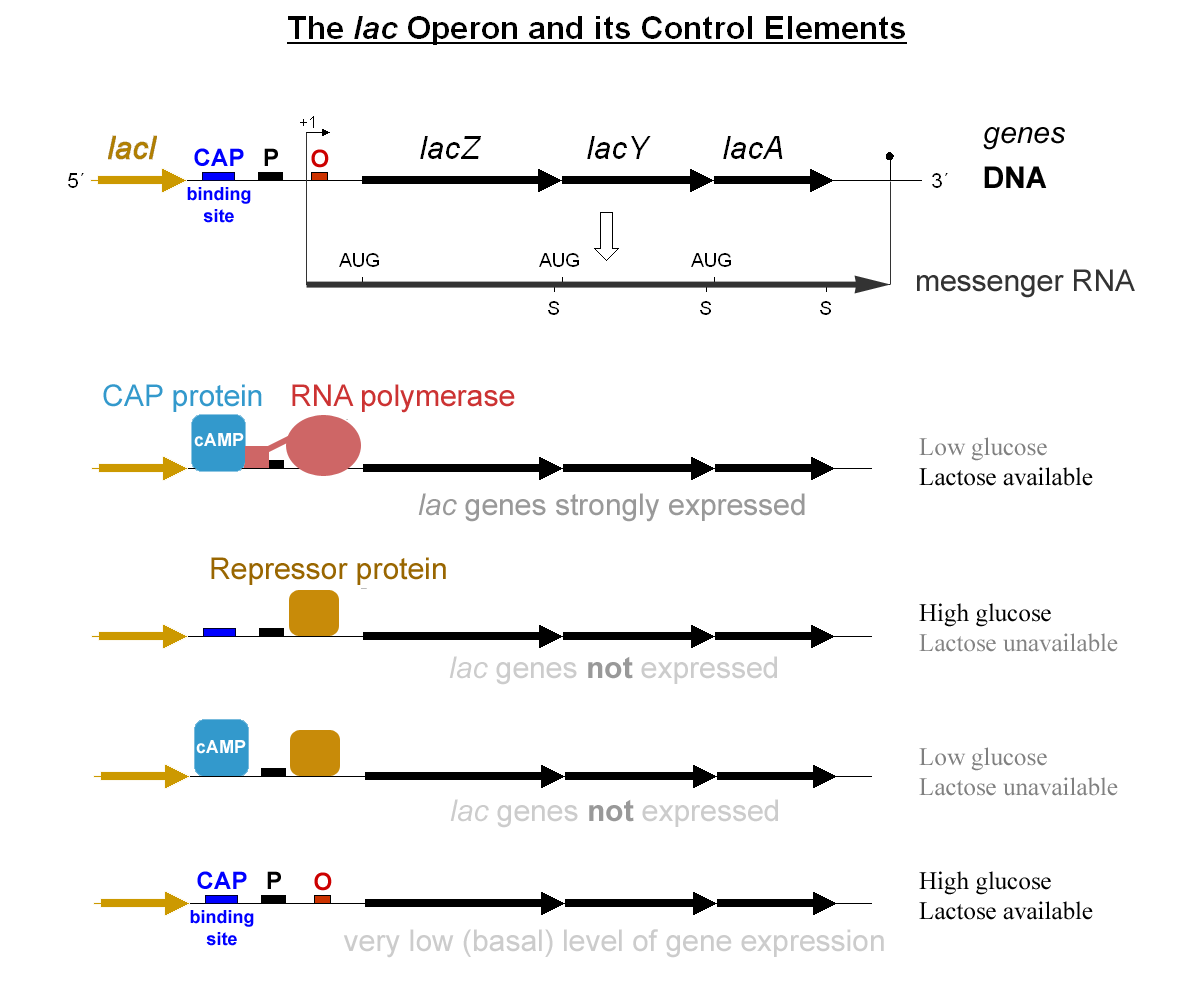
In cell biology, catabolite activator protein (CAP), which is also known as cAMP receptor protein (CRP), is a trans-acting transcriptional activator in bacteria that effectively catalyzes the initiation of DNA transcription by interacting with RNA polymerase in a way that causes the DNA to bend. CAP's name reflects the protein's ability to affect transcription of genes involved in many catabolic pathways. For example, when the amount of glucose transported into a cell is low, a cascade of events results in the increase of cAMP levels in the cell's cytosol, and this increase in cAMP levels is sensed by CAP, which goes on to activate the transcription of many other catabolic genes. CAP exists as a homodimer in solution, and it is bound to by two cyclic AMP (cAMP) ligand molecules with negative cooperativity. By increasing CAP's affinity for DNA, cyclic AMP functions as an allosteric effector. With its cyclic-AMP ligand, CAP binds a DNA region upstream from the site at whic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Activator Protein
A transcriptional activator is a protein (transcription factor) that increases transcription of a gene or set of genes. Activators are considered to have ''positive'' control over gene expression, as they function to promote gene transcription and, in some cases, are required for the transcription of genes to occur. Most activators are DNA-binding proteins that bind to enhancers or promoter-proximal elements. The DNA site bound by the activator is referred to as an "activator-binding site". The part of the activator that makes protein–protein interactions with the general transcription machinery is referred to as an "activating region" or "activation domain". Most activators function by binding sequence-specifically to a regulatory DNA site located near a promoter and making protein–protein interactions with the general transcription machinery (RNA polymerase and general transcription factors), thereby facilitating the binding of the general transcription machinery to the p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |