|
Human Proteome Resource
Proteomics is the large-scale study of proteins. Proteins are vital macromolecules of all living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replication of DNA. In addition, other kinds of proteins include antibodies that protect an organism from infection, and hormones that send important signals throughout the body. The proteome is the entire set of proteins produced or modified by an organism or system. Proteomics enables the identification of ever-increasing numbers of proteins. This varies with time and distinct requirements, or stresses, that a cell or organism undergoes. Proteomics is an interdisciplinary domain that has benefited greatly from the genetic information of various genome projects, including the Human Genome Project. It covers the exploration of proteomes from the overall level of protein composition, structure, and activity, and is an important component of function ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Pattern Analyzer
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residues ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process (including its Regulation of gene expression, regulation) by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, proteins or non-coding RNA, and ultimately affect a phenotype. These products are often proteins, but in non-protein-coding genes such as Transfer RNA, transfer RNA (tRNA) and Small nuclear RNA, small nuclear RNA (snRNA), the product is a functional List of RNAs, non-coding RNA. The process of gene expression is used by all known life—eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and viruses—to generate the macromolecule, macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenotype, ''i.e.'' observable trait. The genetic information stored in DNA represents the genotype, whereas the phenotype results from the "interpretation" of that informati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Methylation
Methylation, in the chemistry, chemical sciences, is the addition of a methyl group on a substrate (chemistry), substrate, or the substitution of an atom (or group) by a methyl group. Methylation is a form of alkylation, with a methyl group replacing a hydrogen#Compounds, hydrogen atom. These terms are commonly used in chemistry, biochemistry, soil science, and biology. In biological systems, methylation is Catalysis, catalyzed by enzymes; such methylation can be involved in modification of heavy metals, regulation of gene expression, regulation of Protein#Functions, protein function, and RNA processing. ''In vitro'' methylation of tissue samples is also a way to reduce some histology#Histological Artifacts, histological staining artifacts. The reverse of methylation is demethylation. In biology In biological systems, methylation is accomplished by enzymes. Methylation can modify heavy metals and can regulate gene expression, RNA processing, and protein function. It is a key pro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitination
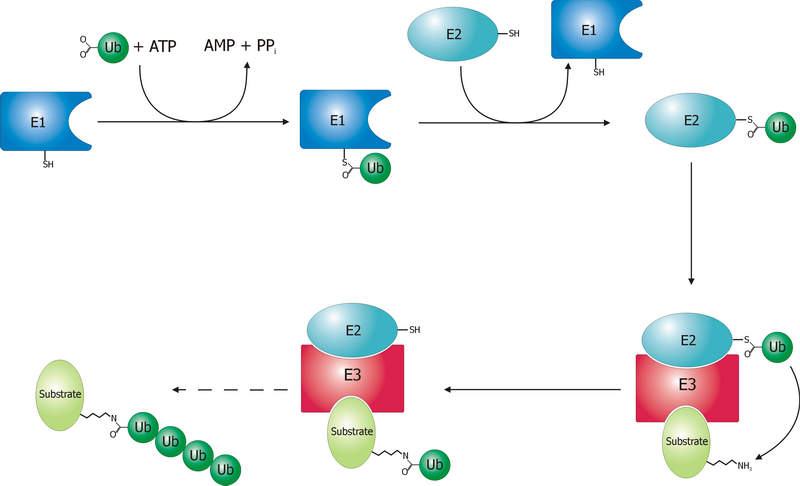
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A. The addition of ubiquitin to a substrate protein is called ubiquitylation (or ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for degradation via the 26S proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitylation involves three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade is to bind ubiquitin to lysine residues on the protein substrate via an isopeptide bond, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
E3 Ubiquitin Ligase
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another protein (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A. The addition of ubiquitin to a substrate protein is called ubiquitylation (or ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for degradation via the 26S proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitylation involves three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade is to bind ubiquitin to lysine residues on the protein substrate via an isopeptide bond, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Kinases
In biochemistry, a kinase () is an enzyme that catalysis, catalyzes the transfer of phosphate groups from High-energy phosphate, high-energy, phosphate-donating molecules to specific Substrate (biochemistry), substrates. This process is known as phosphorylation, where the high-energy adenosine triphosphate, ATP molecule donates a phosphate group to the substrate (biology), substrate molecule. As a result, kinase produces a phosphorylated substrate and Adenosine diphosphate, ADP. Conversely, it is referred to as dephosphorylation when the phosphorylated substrate donates a phosphate group and ADP gains a phosphate group (producing a dephosphorylated substrate and the high energy molecule of ATP). These two processes, phosphorylation and dephosphorylation, occur four times during glycolysis. Kinases are part of the larger family of phosphotransferases. Kinases should not be confused with phosphorylases, which catalyze the addition of inorganic phosphate groups to an acceptor, nor w ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tyrosine
-Tyrosine or tyrosine (symbol Tyr or Y) or 4-hydroxyphenylalanine is one of the 20 standard amino acids that are used by cells to synthesize proteins. It is a conditionally essential amino acid with a polar side group. The word "tyrosine" is from the Greek ''tyrós'', meaning ''cheese'', as it was first discovered in 1846 by German chemist Justus von Liebig in the protein casein from cheese. It is called tyrosyl when referred to as a functional group or side chain. While tyrosine is generally classified as a hydrophobic amino acid, it is more hydrophilic than phenylalanine. It is encoded by the codons UAC and UAU in messenger RNA. The one-letter symbol Y was assigned to tyrosine for being alphabetically nearest of those letters available. Note that T was assigned to the structurally simpler threonine, U was avoided for its similarity with V for valine, W was assigned to tryptophan, while X was reserved for undetermined or atypical amino acids. The mnemonic t''Y''rosine was ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Kinase
In biochemistry, a kinase () is an enzyme that catalyzes the transfer of phosphate groups from high-energy, phosphate-donating molecules to specific substrates. This process is known as phosphorylation, where the high-energy ATP molecule donates a phosphate group to the substrate molecule. As a result, kinase produces a phosphorylated substrate and ADP. Conversely, it is referred to as dephosphorylation when the phosphorylated substrate donates a phosphate group and ADP gains a phosphate group (producing a dephosphorylated substrate and the high energy molecule of ATP). These two processes, phosphorylation and dephosphorylation, occur four times during glycolysis. Kinases are part of the larger family of phosphotransferases. Kinases should not be confused with phosphorylases, which catalyze the addition of inorganic phosphate groups to an acceptor, nor with phosphatases, which remove phosphate groups (dephosphorylation). The phosphorylation state of a molecule, whether it ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Threonine
Threonine (symbol Thr or T) is an amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated −NH form when dissolved in water), a carboxyl group (which is in the deprotonated −COO− form when dissolved in water), and a side chain containing a hydroxyl group, making it a polar, uncharged amino acid. It is essential in humans, meaning the body cannot synthesize it: it must be obtained from the diet. Threonine is synthesized from aspartate in bacteria such as ''E. coli''. It is encoded by all the codons starting AC (ACU, ACC, ACA, and ACG). Threonine sidechains are often hydrogen bonded; the most common small motifs formed are based on interactions with serine: ST turns, ST motifs (often at the beginning of alpha helices) and ST staples (usually at the middle of alpha helices). Modifications The threonine residue is susceptible to numerous posttranslational modifications. The hydroxyl side-chain can und ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine
Serine (symbol Ser or S) is an α-amino acid that is used in the biosynthesis of proteins. It contains an α- amino group (which is in the protonated − form under biological conditions), a carboxyl group (which is in the deprotonated − form under biological conditions), and a side chain consisting of a hydroxymethyl group, classifying it as a polar amino acid. It can be synthesized in the human body under normal physiological circumstances, making it a nonessential amino acid. It is encoded by the codons UCU, UCC, UCA, UCG, AGU and AGC. Occurrence This compound is one of the proteinogenic amino acids. Only the L- stereoisomer appears naturally in proteins. It is not essential to the human diet, since it is synthesized in the body from other metabolites, including glycine. Serine was first obtained from silk protein, a particularly rich source, in 1865 by Emil Cramer. Its name is derived from the Latin for silk, '' sericum''. Serine's structure was established in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell Signaling
In biology, cell signaling (cell signalling in British English) is the Biological process, process by which a Cell (biology), cell interacts with itself, other cells, and the environment. Cell signaling is a fundamental property of all Cell (biology), cellular life in both prokaryotes and eukaryotes. Typically, the signaling process involves three components: the signal, the receptor, and the effector. In biology, signals are mostly chemical in nature, but can also be physical cues such as pressure, Membrane potential, voltage, temperature, or light. Chemical signals are molecules with the ability to bind and activate a specific Receptor (biochemistry), receptor. These molecules, also referred to as Ligand (biochemistry), ligands, are chemically diverse, including ions (e.g. Na+, K+, Ca2+, etc.), lipids (e.g. steroid, prostaglandin), peptides (e.g. insulin, ACTH), carbohydrates, glycosylated proteins (proteoglycans), nucleic acids, etc. Peptide and lipid ligands are particularly ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |