|
Chromosome Segregation
Chromosome segregation is the process in eukaryotes by which two sister chromatids formed as a consequence of DNA replication, or paired homologous chromosomes, separate from each other and migrate to opposite poles of the nucleus. This segregation process occurs during both mitosis and meiosis. Chromosome segregation also occurs in prokaryotes. However, in contrast to eukaryotic chromosome segregation, replication and segregation are not temporally separated. Instead segregation occurs progressively following replication. Mitotic chromatid segregation During mitosis chromosome segregation occurs routinely as a step in cell division (see mitosis diagram). As indicated in the mitosis diagram, mitosis is preceded by a round of DNA replication, so that each chromosome forms two copies called chromatids. These chromatids separate to opposite poles, a process facilitated by a protein complex referred to as cohesin. Upon proper segregation, a complete set of chromatids ends ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Eukaryote
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bacteria and Archaea (both prokaryotes) make up the other two domains. The eukaryotes are usually now regarded as having emerged in the Archaea or as a sister of the Asgard archaea. This implies that there are only two domains of life, Bacteria and Archaea, with eukaryotes incorporated among archaea. Eukaryotes represent a small minority of the number of organisms, but, due to their generally much larger size, their collective global biomass is estimated to be about equal to that of prokaryotes. Eukaryotes emerged approximately 2.3–1.8 billion years ago, during the Proterozoic eon, likely as flagellated phagotrophs. Their name comes from the Greek εὖ (''eu'', "well" or "good") and κάρυον (''karyon'', "nut" or "kernel"). E ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Aneuploidy
Aneuploidy is the presence of an abnormal number of chromosomes in a cell, for example a human cell having 45 or 47 chromosomes instead of the usual 46. It does not include a difference of one or more complete sets of chromosomes. A cell with any number of complete chromosome sets is called a ''euploid'' cell. An extra or missing chromosome is a common cause of some genetic disorders. Some cancer cells also have abnormal numbers of chromosomes. About 68% of human solid tumors are aneuploid. Aneuploidy originates during cell division when the chromosomes do not separate properly between the two cells ( nondisjunction). Most cases of aneuploidy in the autosomes result in miscarriage, and the most common extra autosomal chromosomes among live births are 21, 18 and 13. Chromosome abnormalities are detected in 1 of 160 live human births. Autosomal aneuploidy is more dangerous than sex chromosome aneuploidy, as autosomal aneuploidy is almost always lethal to embryos that cease dev ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Dimer
In biochemistry, a protein dimer is a macromolecular complex formed by two protein monomers, or single proteins, which are usually non-covalently bound. Many macromolecules, such as proteins or nucleic acids, form dimers. The word ''dimer'' has roots meaning "two parts", '' di-'' + '' -mer''. A protein dimer is a type of protein quaternary structure. A protein homodimer is formed by two identical proteins. A protein heterodimer is formed by two different proteins. Most protein dimers in biochemistry are not connected by covalent bonds. An example of a non-covalent heterodimer is the enzyme reverse transcriptase, which is composed of two different amino acid chains. An exception is dimers that are linked by disulfide bridges such as the homodimeric protein NEMO. Some proteins contain specialized domains to ensure dimerization (dimerization domains) and specificity. The G protein-coupled cannabinoid receptors have the ability to form both homo- and heterodimers with several ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MLH3
DNA mismatch repair protein Mlh3 is a protein that in humans is encoded by the ''MLH3'' gene. Function This gene is a member of the MutL-homolog (MLH) family of DNA mismatch repair (MMR) genes. MLH genes are implicated in maintaining genomic integrity during DNA replication and after meiotic recombination. The protein encoded by this gene functions as a heterodimer with other family members. Somatic mutations in this gene frequently occur in tumors exhibiting microsatellite instability, and germline mutations have been linked to hereditary nonpolyposis colorectal cancer type 7 (HNPCC7). Several alternatively spliced transcript variants have been identified, but the full-length nature of only two transcript variants has been determined. Orthologs of human MLH3 have also been studied in other organisms including mouse and the budding yeast ''Saccharomyces cerevisiae''. Meiosis In addition to its role in DNA mismatch repair, MLH3 protein is also involved in meiotic crossing over. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MLH1
DNA mismatch repair protein Mlh1 or MutL protein homolog 1 is a protein that in humans is encoded by the MLH1 gene located on chromosome 3. It is a gene commonly associated with hereditary nonpolyposis colorectal cancer. Orthologs of human MLH1 have also been studied in other organisms including mouse and the budding yeast ''Saccharomyces cerevisiae''. Function This gene was identified as a locus frequently mutated in hereditary nonpolyposis colon cancer. It is a human homolog of the ''E. coli'' DNA mismatch repair gene, mutL, which mediates protein-protein interactions during mismatch recognition, strand discrimination, and strand removal. Defects in MLH1 are associated with the microsatellite instability observed in hereditary nonpolyposis colon cancer. Alternatively spliced transcript variants encoding different isoforms have been described, but their full-length natures have not been determined. Role in DNA mismatch repair MLH1 protein is one component of a system of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Saccharomyces Cerevisiae
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have been originally isolated from the skin of grapes. It is one of the most intensively studied eukaryotic model organisms in molecular and cell biology, much like '' Escherichia coli'' as the model bacterium. It is the microorganism behind the most common type of fermentation. ''S. cerevisiae'' cells are round to ovoid, 5–10 μm in diameter. It reproduces by budding. Many proteins important in human biology were first discovered by studying their homologs in yeast; these proteins include cell cycle proteins, signaling proteins, and protein-processing enzymes. ''S. cerevisiae'' is currently the only yeast cell known to have Berkeley bodies present, which are involved in particular secretory pathways. Antibodies agains ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Topoisomerase
DNA topoisomerases (or topoisomerases) are enzymes that catalyze changes in the topological state of DNA, interconverting relaxed and supercoiled forms, linked (catenated) and unlinked species, and knotted and unknotted DNA. Topological issues in DNA arise due to the intertwined nature of its double-helical structure, which, for example, can lead to overwinding of the DNA duplex during DNA replication and transcription. If left unchanged, this torsion would eventually stop the DNA or RNA polymerases involved in these processes from continuing along the DNA helix. A second topological challenge results from the linking or tangling of DNA during replication. Left unresolved, links between replicated DNA will impede cell division. The DNA topoisomerases prevent and correct these types of topological problems. They do this by binding to DNA and cutting the sugar-phosphate backbone of either one (type I topoisomerases) or both (type II topoisomerases) of the DNA strands. This transie ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Repair
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages (interstrand crosslinks or ICLs). This can eventually lead to malig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Holliday Junction
A Holliday junction is a branched nucleic acid structure that contains four double-stranded arms joined. These arms may adopt one of several conformations depending on buffer salt concentrations and the sequence of nucleobases closest to the junction. The structure is named after Robin Holliday, the molecular biologist who proposed its existence in 1964. In biology, Holliday junctions are a key intermediate in many types of genetic recombination, as well as in double-strand break repair. These junctions usually have a symmetrical sequence and are thus mobile, meaning that the four individual arms may slide through the junction in a specific pattern that largely preserves base pairing. Additionally, four-arm junctions similar to Holliday junctions appear in some functional RNA molecules. Immobile Holliday junctions, with asymmetrical sequences that lock the strands in a specific position, were artificially created by scientists to study their structure as a model for natura ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cohesin
Cohesin is a protein complex that mediates sister chromatid cohesion, homologous recombination, and DNA looping. Cohesin is formed of SMC3, SMC1, SCC1 and SCC3 ( SA1 or SA2 in humans). Cohesin holds sister chromatids together after DNA replication until anaphase when removal of cohesin leads to separation of sister chromatids. The complex forms a ring-like structure and it is believed that sister chromatids are held together by entrapment inside the cohesin ring. Cohesin is a member of the SMC family of protein complexes which includes Condensin, MukBEF and SMC-ScpAB. Cohesin was separately discovered in budding yeast by Douglas Koshland and Kim Nasmyth. Structure Cohesin is a multi-subunit protein complex, made up of SMC1, SMC3, RAD21 and SCC3 (SA1 or SA2). SMC1 and SMC3 are members of the Structural Maintenance of Chromosomes (SMC) family. SMC proteins have two main structural characteristics: an ATP-binding cassette-like 'head' domain with ATPase activity (form ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
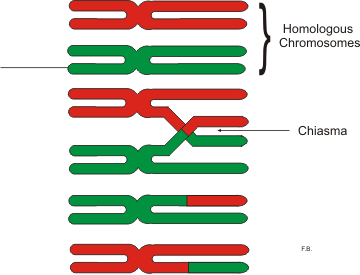
Chiasma (genetics)
In genetics, a chiasma (pl. chiasmata) is the point of contact, the physical link, between two (non-sister) chromatids belonging to homologous chromosomes. At a given chiasma, an exchange of genetic material can occur between both chromatids, what is called a chromosomal crossover, but this is much more frequent during meiosis than mitosis. In meiosis, absence of a chiasma generally results in improper chromosomal segregation and aneuploidy. Points of crossing over become visible as chiasma after the synaptonemal complex dissembles and the homologous chromosomes slightly apart from each other. The phenomenon of genetic chiasmata (''chiasmatypie'') was discovered and described in 1909 by Frans Alfons Janssens, a Professor at the University of Leuven in Belgium. When each tetrad, which is composed of two pairs of sister chromatids, begins to split, the only points of contact are at the chiasmata. The chiasmata become visible during the diplotene stage of prophase I of meiosi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromosomal Crossover
Chromosomal crossover, or crossing over, is the exchange of genetic material during sexual reproduction between two homologous chromosomes' non-sister chromatids that results in recombinant chromosomes. It is one of the final phases of genetic recombination, which occurs in the ''pachytene'' stage of prophase I of meiosis during a process called synapsis. Synapsis begins before the synaptonemal complex develops and is not completed until near the end of prophase I. Crossover usually occurs when matching regions on matching chromosomes break and then reconnect to the other chromosome. Crossing over was described, in theory, by Thomas Hunt Morgan. He relied on the discovery of Frans Alfons Janssens who described the phenomenon in 1909 and had called it "chiasmatypie". The term '' chiasma'' is linked, if not identical, to chromosomal crossover. Morgan immediately saw the great importance of Janssens' cytological interpretation of chiasmata to the experimental results ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |