|
CCDC144A
Coiled-coil domain-containing protein 144A is a protein that in humans is encoded by the ''CCDC144A'' gene. An alias of this gene is called KIAA0565. There are four members of the CCDC family: CCDC 144A, 144B, 144C and putative CCDC 144 N-terminal like proteins. Gene This gene has a nucleotide sequence that is 5140 bp long, and it encodes 641 amino acids. It is found on the short arm, plus (forward) strand of chromosome 17 at p11.2. The mRNA for the CCDC144A gene has 3 alternative splicing isoforms named A2RUR9-1, A2RUR9-2, AND A2RUR9-3, but there is no experimental confirmation available yet. Protein This protein for this gene is also known as coiled coil domain containing 144A (CCDC144A) protein. It consists of 641 amino acids. This protein weighs 75.8 kDa and has an isoelectric point of 6.357. This protein localizes near the nucleus, and is a soluble protein with a hydrophobicity of -1.021842. This protein is also non-secretory and has 10 potential serine and 3 potent ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metabolic reactions, DNA replication, Cell signaling, responding to stimuli, providing Cytoskeleton, structure to cells and Fibrous protein, organisms, and Intracellular transport, transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the Nucleic acid sequence, nucleotide sequence of their genes, and which usually results in protein folding into a specific Protein structure, 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called pep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Domain Of Unknown Function
A domain of unknown function (DUF) is a protein domain that has no characterised function. These families have been collected together in the Pfam database using the prefix DUF followed by a number, with examples being DUF2992 and DUF1220. As of 2019, there are almost 4,000 DUF families within the Pfam database representing over 22% of known families. Some DUFs are not named using the nomenclature due to popular usage but are nevertheless DUFs. The DUF designation is tentative, and such families tend to be renamed to a more specific name (or merged to an existing domain) after a function is identified. History The DUF naming scheme was introduced by Chris Ponting, through the addition of DUF1 and DUF2 to the SMART database. These two domains were found to be widely distributed in bacterial signaling proteins. Subsequently, the functions of these domains were identified and they have since been renamed as the GGDEF domain and EAL domain respectively. Characterisation Structural gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Random Coil
In polymer chemistry, a random coil is a conformation of polymers where the monomer subunits are oriented randomly while still being bonded to adjacent units. It is not one specific shape, but a statistical distribution of shapes for all the chains in a population of macromolecules. The conformation's name is derived from the idea that, in the absence of specific, stabilizing interactions, a polymer backbone will "sample" all possible conformations randomly. Many unbranched, linear homopolymers — in solution, or above their melting temperatures — assume (approximate) random coils. Random walk model: The Gaussian chain There are an enormous number of different ways in which a chain can be curled around in a relatively compact shape, like an unraveling ball of twine with much open space, and comparatively few ways it can be more or less stretched out. So, if each conformation has an equal probability or statistical weight, chains are much more likely to be ball-like th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helices
An alpha helix (or α-helix) is a sequence of amino acids in a protein that are twisted into a coil (a helix). The alpha helix is the most common structural arrangement in the secondary structure of proteins. It is also the most extreme type of local structure, and it is the local structure that is most easily predicted from a sequence of amino acids. The alpha helix has a right-handed helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid that is four residues earlier in the protein sequence. Other names The alpha helix is also commonly called a: * Pauling–Corey–Branson α-helix (from the names of three scientists who described its structure) * 3.613-helix because there are 3.6 amino acids in one ring, with 13 atoms being involved in the ring formed by the hydrogen bond (starting with amidic hydrogen and ending with carbonyl oxygen) Discovery In the early 1930s, William Astbury showed that there were dras ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Leucine Zipper
A leucine zipper (or leucine scissors) is a common three-dimensional structural motif in proteins. They were first described by Landschulz and collaborators in 1988 when they found that an enhancer binding protein had a very characteristic 30-amino acid segment and the display of these amino acid sequences on an idealized alpha helix revealed a periodic repetition of leucine residues at every seventh position over a distance covering eight helical turns. The polypeptide segments containing these periodic arrays of leucine residues were proposed to exist in an alpha-helical conformation and the leucine side chains from one alpha helix interdigitate with those from the alpha helix of a second polypeptide, facilitating dimerization. Leucine zippers are a dimerization motif of the BZIP domain, bZIP (Basic-region leucine zipper) class of eukaryotic transcription factors. The bZIP domain is 60 to 80 amino acids in length with a highly conserved DNA binding basic region and a more divers ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stem-loop Structure
Stem-loops are nucleic acid secondary structural elements which form via intramolecular base pairing in single-stranded DNA or RNA. They are also referred to as hairpins or hairpin loops. A stem-loop occurs when two regions of the same nucleic acid strand, usually complementary in nucleotide sequence, base-pair to form a double helix that ends in a loop of unpaired nucleotides. Stem-loops are most commonly found in RNA, and are a key building block of many RNA secondary structures. Stem-loops can direct RNA folding, protect structural stability for messenger RNA (mRNA), provide recognition sites for RNA binding proteins, and serve as a substrate for enzymatic reactions. Formation and stability The formation of a stem-loop is dependent on the stability of the helix and loop regions. The first prerequisite is the presence of a sequence that can fold back on itself to form a paired double helix. The stability of this helix is determined by its length, the number of mismatches ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
3' Untranslated Region
In molecular genetics, the three prime untranslated region (3′-UTR) is the section of messenger RNA (mRNA) that immediately follows the translation termination codon. The 3′-UTR often contains regulatory regions that post-transcriptionally influence gene expression. During gene expression, an mRNA molecule is transcribed from the DNA sequence and is later translated into a protein. Several regions of the mRNA molecule are not translated into a protein including the 5' cap, 5' untranslated region, 3′ untranslated region and poly(A) tail. Regulatory regions within the 3′-untranslated region can influence polyadenylation, translation efficiency, localization, and stability of the mRNA. The 3′-UTR contains binding sites for both regulatory proteins and microRNAs (miRNAs). By binding to specific sites within the 3′-UTR, miRNAs can decrease gene expression of various mRNAs by either inhibiting translation or directly causing degradation of the transcript. The 3′-U ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
5' UTR
The 5′ untranslated region (also known as 5′ UTR, leader sequence, transcript leader, or leader RNA) is the region of a messenger RNA (mRNA) that is directly Upstream and downstream (DNA), upstream from the initiation codon. This region is important for the regulation of translation (biology), translation of a transcript by differing mechanisms in viruses, prokaryotes and eukaryotes. Despite its name, the 5′ UTR, or a portion of it is sometimes translated into a protein product. This product may involve in regulation of Transcription (biology), transcription, and translation of the main coding sequence of the mRNA, such as the Drosophila melanogaster#Sex determination, sex-lethal gene in ''Drosophila''. Regulatory elements within 5′ UTRs have also been linked to mRNA export. In many organisms, however, the 5′ UTR is completely untranslated, instead forming a complex Nucleic acid secondary structure, secondary structure to regulate translation. General structure Length ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Heptad Repeat
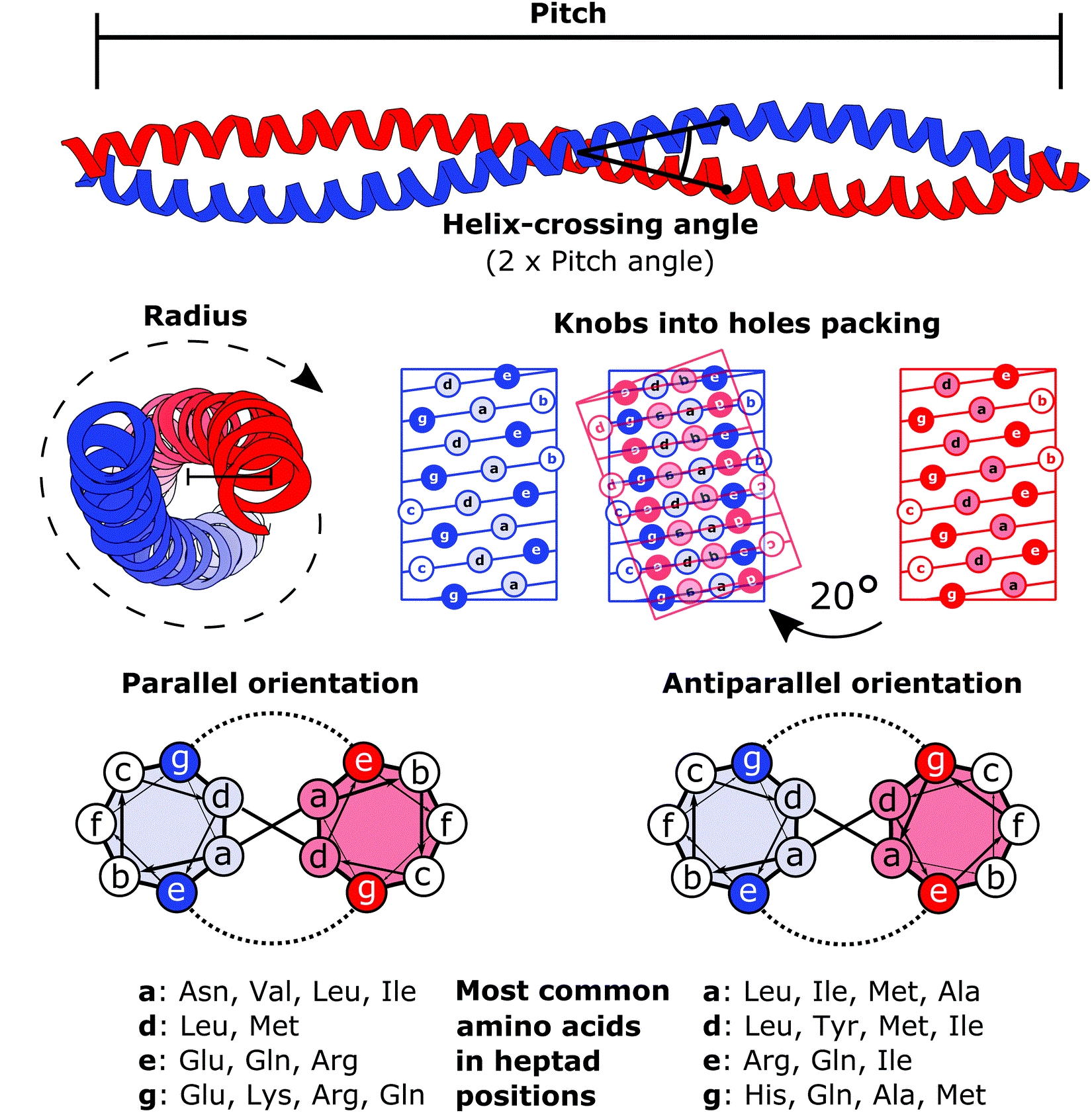
The heptad repeat is an example of a structural motif that consists of a repeating pattern of seven amino acids: ''a b c d e f g'' H P P H C P C where H represents hydrophobic residues, C represents, typically, charged residues, and P represents polar (and, therefore, hydrophilic) residues. The positions of the heptad repeat are commonly denoted by the lowercase letters ''a'' through ''g''. These motifs are the basis for most coiled coils and, in particular, leucine zippers, which have predominantly leucine in the ''d'' position of the heptad repeat. A conformational change in a heptad repeat in the SARS-CoV-2 spike protein In virology, a spike protein or peplomer protein is a protein that forms a large structure known as a spike or peplomer projecting from the surface of an viral envelope, enveloped virus. as cited in The proteins are usually glycoproteins that ... facilitates entry of the virus into the host cell membrane. References {{DEFAULTSORT:Heptad Rep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Coiled Coil
A coiled coil is a structural motif in proteins in which two to seven alpha-helices are coiled together like the strands of a rope. ( Dimers and trimers are the most common types.) They have been found in roughly 5-10% of proteins and have a variety of functions. They are one of the most widespread motifs found in protein-protein interactions. To aid protein study, several tools have been developed to predict coiled-coils in protein structures. Many coiled coil-type proteins are involved in important biological functions, such as the regulation of gene expression — e.g., transcription factors. Notable examples are the oncoproteins c-Fos and c-Jun, as well as the muscle protein tropomyosin. Discovery The possibility of coiled coils for α-keratin was initially somewhat controversial. Linus Pauling and Francis Crick independently came to the conclusion that this was possible at about the same time. In the summer of 1952, Pauling visited the laboratory in England where C ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Eukaryote
The eukaryotes ( ) constitute the Domain (biology), domain of Eukaryota or Eukarya, organisms whose Cell (biology), cells have a membrane-bound cell nucleus, nucleus. All animals, plants, Fungus, fungi, seaweeds, and many unicellular organisms are eukaryotes. They constitute a major group of Outline of life forms, life forms alongside the two groups of prokaryotes: the Bacteria and the Archaea. Eukaryotes represent a small minority of the number of organisms, but given their generally much larger size, their collective global biomass is much larger than that of prokaryotes. The eukaryotes emerged within the archaeal Kingdom (biology), kingdom Asgard (Archaea), Promethearchaeati and its sole phylum Promethearchaeota. This implies that there are only Two-domain system, two domains of life, Bacteria and Archaea, with eukaryotes incorporated among the Archaea. Eukaryotes first emerged during the Paleoproterozoic, likely as Flagellated cell, flagellated cells. The leading evolutiona ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Signal Peptide
A signal peptide (sometimes referred to as signal sequence, targeting signal, localization signal, localization sequence, transit peptide, leader sequence or leader peptide) is a short peptide (usually 16–30 amino acids long) present at the N-terminus (or occasionally nonclassically at the C-terminus or internally) of most newly synthesized proteins that are destined toward the secretory pathway. These proteins include those that reside either inside certain organelles (the endoplasmic reticulum, Golgi or endosomes), secreted from the cell, or inserted into most cellular membranes. Although most type I membrane-bound proteins have signal peptides, most type II and multi-spanning membrane-bound proteins are targeted to the secretory pathway by their first transmembrane domain, which biochemically resembles a signal sequence except that it is not cleaved. They are a kind of target peptide. Function (translocation) Signal peptides function to prompt a cell to transloc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |