Phylogenetic tree on:
[Wikipedia]
[Google]
[Amazon]
A phylogenetic tree or phylogeny is a graphical representation which shows the
 A rooted phylogenetic
A rooted phylogenetic
 Unrooted trees illustrate the relatedness of the leaf nodes without making assumptions about ancestry. They do not require the ancestral root to be known or inferred. Rooted trees can be generated from unrooted ones by inserting a root. Inferring the root of an unrooted tree requires some means of identifying ancestry. This is normally done by including an outgroup in the input data so that the root is necessarily between the outgroup and the rest of the taxa in the tree, or by introducing additional assumptions about the relative rates of evolution on each branch, such as an application of the molecular clock
Unrooted trees illustrate the relatedness of the leaf nodes without making assumptions about ancestry. They do not require the ancestral root to be known or inferred. Rooted trees can be generated from unrooted ones by inserting a root. Inferring the root of an unrooted tree requires some means of identifying ancestry. This is normally done by including an outgroup in the input data so that the root is necessarily between the outgroup and the rest of the taxa in the tree, or by introducing additional assumptions about the relative rates of evolution on each branch, such as an application of the molecular clock
 The number of possible trees for a given number of leaf nodes depends on the specific type of tree, but there are always more labeled than unlabeled trees, more multifurcating than bifurcating trees, and more rooted than unrooted trees. The last distinction is the most biologically relevant; it arises because there are many places on an unrooted tree to put the root. For bifurcating labeled trees, the total number of rooted trees is:
: for , represents the number of leaf nodes.
For bifurcating labeled trees, the total number of unrooted trees is:
: for .
Among labeled bifurcating trees, the number of unrooted trees with leaves is equal to the number of rooted trees with leaves.
The number of rooted trees grows quickly as a function of the number of tips. For 10 tips, there are more than possible bifurcating trees, and the number of multifurcating trees rises faster, with ca. 7 times as many of the latter as of the former.
The number of possible trees for a given number of leaf nodes depends on the specific type of tree, but there are always more labeled than unlabeled trees, more multifurcating than bifurcating trees, and more rooted than unrooted trees. The last distinction is the most biologically relevant; it arises because there are many places on an unrooted tree to put the root. For bifurcating labeled trees, the total number of rooted trees is:
: for , represents the number of leaf nodes.
For bifurcating labeled trees, the total number of unrooted trees is:
: for .
Among labeled bifurcating trees, the number of unrooted trees with leaves is equal to the number of rooted trees with leaves.
The number of rooted trees grows quickly as a function of the number of tips. For 10 tips, there are more than possible bifurcating trees, and the number of multifurcating trees rises faster, with ca. 7 times as many of the latter as of the former.


 A spindle diagram, or bubble diagram, is often called a romerogram, after its popularisation by the American palaeontologist Alfred Romer.
It represents taxonomic diversity (horizontal width) against geological time (vertical axis) in order to reflect the variation of abundance of various taxa through time.
A spindle diagram is not an evolutionary tree: the taxonomic spindles obscure the actual relationships of the parent taxon to the daughter taxon and have the disadvantage of involving the
A spindle diagram, or bubble diagram, is often called a romerogram, after its popularisation by the American palaeontologist Alfred Romer.
It represents taxonomic diversity (horizontal width) against geological time (vertical axis) in order to reflect the variation of abundance of various taxa through time.
A spindle diagram is not an evolutionary tree: the taxonomic spindles obscure the actual relationships of the parent taxon to the daughter taxon and have the disadvantage of involving the
 Darwin also mentioned that the ''coral'' may be a more suitable metaphor than the ''tree''. Indeed, phylogenetic corals are useful for portraying past and present life, and they have some advantages over trees ( anastomoses allowed, etc.).
Darwin also mentioned that the ''coral'' may be a more suitable metaphor than the ''tree''. Indeed, phylogenetic corals are useful for portraying past and present life, and they have some advantages over trees ( anastomoses allowed, etc.).
Human Y-Chromosome 2002 Phylogenetic Tree
iTOL: Interactive Tree Of Life
Phylogenetic Tree of Artificial Organisms Evolved on Computers
Miyamoto and Goodman's Phylogram of Eutherian Mammals
OneZoom: Tree of Life – all living species as intuitive and zoomable fractal explorer (responsive design)
Discover Life
An interactive tree based on the U.S. National Science Foundation's Assembling the Tree of Life Project
* ttp://tolweb.org/tree Tree of Life Web Project
Phylogenetic inferring on the T-REX server
NCBI's Taxonomy Database
ETE: A Python Environment for Tree Exploration
This is a programming library to analyze, manipulate and visualize phylogenetic trees
Ref.
A daily-updated tree of (sequenced) life
{{DEFAULTSORT:Phylogenetic Tree Phylogenetics Trees (data structures)
evolution
Evolution is the change in the heritable Phenotypic trait, characteristics of biological populations over successive generations. It occurs when evolutionary processes such as natural selection and genetic drift act on genetic variation, re ...
ary history between a set of species
A species () is often defined as the largest group of organisms in which any two individuals of the appropriate sexes or mating types can produce fertile offspring, typically by sexual reproduction. It is the basic unit of Taxonomy (biology), ...
or taxa during a specific time.Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA. In other words, it is a branching diagram
A diagram is a symbolic Depiction, representation of information using Visualization (graphics), visualization techniques. Diagrams have been used since prehistoric times on Cave painting, walls of caves, but became more prevalent during the Age o ...
or a tree
In botany, a tree is a perennial plant with an elongated stem, or trunk, usually supporting branches and leaves. In some usages, the definition of a tree may be narrower, e.g., including only woody plants with secondary growth, only ...
showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics. In evolutionary biology, all life on Earth is theoretically part of a single phylogenetic tree, indicating common ancestry. Phylogenetics
In biology, phylogenetics () is the study of the evolutionary history of life using observable characteristics of organisms (or genes), which is known as phylogenetic inference. It infers the relationship among organisms based on empirical dat ...
is the study of phylogenetic trees. The main challenge is to find a phylogenetic tree representing optimal evolutionary ancestry between a set of species or taxa. Computational phylogenetics (also phylogeny inference) focuses on the algorithms involved in finding optimal phylogenetic tree in the phylogenetic landscape.
Phylogenetic trees may be rooted or unrooted. In a ''rooted'' phylogenetic tree, each node with descendants represents the inferred most recent common ancestor of those descendants, and the edge lengths in some trees may be interpreted as time estimates. Each node is called a taxonomic unit. Internal nodes are generally called hypothetical taxonomic units, as they cannot be directly observed. Trees are useful in fields of biology such as bioinformatics
Bioinformatics () is an interdisciplinary field of science that develops methods and Bioinformatics software, software tools for understanding biological data, especially when the data sets are large and complex. Bioinformatics uses biology, ...
, systematics
Systematics is the study of the diversification of living forms, both past and present, and the relationships among living things through time. Relationships are visualized as evolutionary trees (synonyms: phylogenetic trees, phylogenies). Phy ...
, and phylogenetics
In biology, phylogenetics () is the study of the evolutionary history of life using observable characteristics of organisms (or genes), which is known as phylogenetic inference. It infers the relationship among organisms based on empirical dat ...
. ''Unrooted'' trees illustrate only the relatedness of the leaf nodes and do not require the ancestral root to be known or inferred.
History
The idea of atree of life
The tree of life is a fundamental archetype in many of the world's mythology, mythological, religion, religious, and philosophy, philosophical traditions. It is closely related to the concept of the sacred tree.Giovino, Mariana (2007). ''The ...
arose from ancient notions of a ladder-like progression from lower into higher forms of life
Life, also known as biota, refers to matter that has biological processes, such as Cell signaling, signaling and self-sustaining processes. It is defined descriptively by the capacity for homeostasis, Structure#Biological, organisation, met ...
(such as in the Great Chain of Being). Early representations of "branching" phylogenetic trees include a "paleontological chart" showing the geological relationships among plants and animals in the book ''Elementary Geology'', by Edward Hitchcock (first edition: 1840).
Charles Darwin
Charles Robert Darwin ( ; 12 February 1809 – 19 April 1882) was an English Natural history#Before 1900, naturalist, geologist, and biologist, widely known for his contributions to evolutionary biology. His proposition that all speci ...
featured a diagrammatic evolutionary "tree" in his 1859 book '' On the Origin of Species''. Over a century later, evolutionary biologists still use tree diagrams to depict evolution
Evolution is the change in the heritable Phenotypic trait, characteristics of biological populations over successive generations. It occurs when evolutionary processes such as natural selection and genetic drift act on genetic variation, re ...
because such diagrams effectively convey the concept that speciation occurs through the adaptive and semirandom splitting of lineages.
The term ''phylogenetic'', or ''phylogeny'', derives from the two ancient greek
Ancient Greek (, ; ) includes the forms of the Greek language used in ancient Greece and the classical antiquity, ancient world from around 1500 BC to 300 BC. It is often roughly divided into the following periods: Mycenaean Greek (), Greek ...
words (), meaning "race, lineage", and (), meaning "origin, source".
Properties
Rooted tree
 A rooted phylogenetic
A rooted phylogenetic tree
In botany, a tree is a perennial plant with an elongated stem, or trunk, usually supporting branches and leaves. In some usages, the definition of a tree may be narrower, e.g., including only woody plants with secondary growth, only ...
(see two graphics at top) is a directed
Direct may refer to:
Mathematics
* Directed set, in order theory
* Direct limit of (pre), sheaves
* Direct sum of modules, a construction in abstract algebra which combines several vector spaces
Computing
* Direct access (disambiguation), a ...
tree with a unique node — the root — corresponding to the (usually imputed) most recent common ancestor of all the entities at the leaves
A leaf (: leaves) is a principal appendage of the stem of a vascular plant, usually borne laterally above ground and specialized for photosynthesis. Leaves are collectively called foliage, as in "autumn foliage", while the leaves, stem, ...
of the tree. The root node does not have a parent node, but serves as the parent of all other nodes in the tree. The root is therefore a node of degree 2, while other internal nodes have a minimum degree of 3 (where "degree" here refers to the total number of incoming and outgoing edges).
The most common method for rooting trees is the use of an uncontroversial outgroup—close enough to allow inference from trait data or molecular sequencing, but far enough to be a clear outgroup. Another method is midpoint rooting, or a tree can also be rooted by using a non-stationary substitution model.
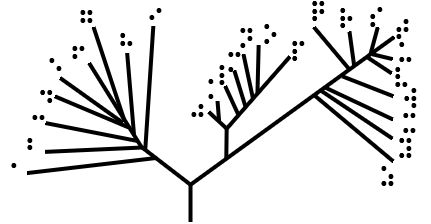
Unrooted tree
 Unrooted trees illustrate the relatedness of the leaf nodes without making assumptions about ancestry. They do not require the ancestral root to be known or inferred. Rooted trees can be generated from unrooted ones by inserting a root. Inferring the root of an unrooted tree requires some means of identifying ancestry. This is normally done by including an outgroup in the input data so that the root is necessarily between the outgroup and the rest of the taxa in the tree, or by introducing additional assumptions about the relative rates of evolution on each branch, such as an application of the molecular clock
Unrooted trees illustrate the relatedness of the leaf nodes without making assumptions about ancestry. They do not require the ancestral root to be known or inferred. Rooted trees can be generated from unrooted ones by inserting a root. Inferring the root of an unrooted tree requires some means of identifying ancestry. This is normally done by including an outgroup in the input data so that the root is necessarily between the outgroup and the rest of the taxa in the tree, or by introducing additional assumptions about the relative rates of evolution on each branch, such as an application of the molecular clock hypothesis
A hypothesis (: hypotheses) is a proposed explanation for a phenomenon. A scientific hypothesis must be based on observations and make a testable and reproducible prediction about reality, in a process beginning with an educated guess o ...
.
Bifurcating versus multifurcating
Both rooted and unrooted trees can be either bifurcating or multifurcating. A rooted bifurcating tree has exactly two descendants arising from each interior node (that is, it forms abinary tree
In computer science, a binary tree is a tree data structure in which each node has at most two children, referred to as the ''left child'' and the ''right child''. That is, it is a ''k''-ary tree with . A recursive definition using set theor ...
), and an unrooted bifurcating tree takes the form of an unrooted binary tree, a free tree with exactly three neighbors at each internal node. In contrast, a rooted multifurcating tree may have more than two children at some nodes and an unrooted multifurcating tree may have more than three neighbors at some nodes.
Labeled versus unlabeled
Both rooted and unrooted trees can be either labeled or unlabeled. A labeled tree has specific values assigned to its leaves, while an unlabeled tree, sometimes called a tree shape, defines a topology only. Some sequence-based trees built from a small genomic locus, such as Phylotree, feature internal nodes labeled with inferred ancestral haplotypes.Enumerating trees
Special tree types
Dendrogram
A dendrogram is a general name for a tree, whether phylogenetic or not, and hence also for the diagrammatic representation of a phylogenetic tree.
Cladogram
Acladogram
A cladogram (from Greek language, Greek ''clados'' "branch" and ''gramma'' "character") is a diagram used in cladistics to show relations among organisms. A cladogram is not, however, an Phylogenetic tree, evolutionary tree because it does not s ...
only represents a branching pattern; i.e., its branch lengths do not represent time or relative amount of character change, and its internal nodes do not represent ancestors.
Phylogram
A phylogram is a phylogenetic tree that has branch lengths proportional to the amount of character change.Chronogram
A chronogram is a phylogenetic tree that explicitly represents time through its branch lengths.Dahlgrenogram
A Dahlgrenogram is a diagram representing a cross section of a phylogenetic tree.Phylogenetic network
A phylogenetic network is not strictly speaking a tree, but rather a more general graph, or a directed acyclic graph in the case of rooted networks. They are used to overcome some of the limitations inherent to trees.Spindle diagram
 A spindle diagram, or bubble diagram, is often called a romerogram, after its popularisation by the American palaeontologist Alfred Romer.
It represents taxonomic diversity (horizontal width) against geological time (vertical axis) in order to reflect the variation of abundance of various taxa through time.
A spindle diagram is not an evolutionary tree: the taxonomic spindles obscure the actual relationships of the parent taxon to the daughter taxon and have the disadvantage of involving the
A spindle diagram, or bubble diagram, is often called a romerogram, after its popularisation by the American palaeontologist Alfred Romer.
It represents taxonomic diversity (horizontal width) against geological time (vertical axis) in order to reflect the variation of abundance of various taxa through time.
A spindle diagram is not an evolutionary tree: the taxonomic spindles obscure the actual relationships of the parent taxon to the daughter taxon and have the disadvantage of involving the paraphyly
Paraphyly is a taxonomic term describing a grouping that consists of the grouping's last common ancestor and some but not all of its descendant lineages. The grouping is said to be paraphyletic ''with respect to'' the excluded subgroups. In co ...
of the parental group.
This type of diagram is no longer used in the form originally proposed.
Coral of life
Construction
Phylogenetic trees composed with a nontrivial number of input sequences are constructed usingcomputational phylogenetics
Computational phylogenetics, phylogeny inference, or phylogenetic inference focuses on computational and optimization algorithms, Heuristic (computer science), heuristics, and approaches involved in Phylogenetics, phylogenetic analyses. The goal i ...
methods. Distance-matrix methods such as neighbor-joining or UPGMA, which calculate genetic distance
Genetic distance is a measure of the genetics, genetic divergence between species or between population#Genetics, populations within a species, whether the distance measures time from common ancestor or degree of differentiation. Populations with ...
from multiple sequence alignments, are simplest to implement, but do not invoke an evolutionary model. Many sequence alignment methods such as ClustalW also create trees by using the simpler algorithms (i.e. those based on distance) of tree construction. Maximum parsimony is another simple method of estimating phylogenetic trees, but implies an implicit model of evolution (i.e. parsimony). More advanced methods use the optimality criterion of maximum likelihood, often within a Bayesian framework, and apply an explicit model of evolution to phylogenetic tree estimation. Identifying the optimal tree using many of these techniques is NP-hard, so heuristic search and optimization
Mathematical optimization (alternatively spelled ''optimisation'') or mathematical programming is the selection of a best element, with regard to some criteria, from some set of available alternatives. It is generally divided into two subfiel ...
methods are used in combination with tree-scoring functions to identify a reasonably good tree that fits the data.
Tree-building methods can be assessed on the basis of several criteria:
* efficiency (how long does it take to compute the answer, how much memory does it need?)
* power (does it make good use of the data, or is information being wasted?)
* consistency (will it converge on the same answer repeatedly, if each time given different data for the same model problem?)
* robustness (does it cope well with violations of the assumptions of the underlying model?)
* falsifiability (does it alert us when it is not good to use, i.e. when assumptions are violated?)
Tree-building techniques have also gained the attention of mathematicians. Trees can also be built using T-theory.
File formats
Trees can be encoded in a number of different formats, all of which must represent the nested structure of a tree. They may or may not encode branch lengths and other features. Standardized formats are critical for distributing and sharing trees without relying on graphics output that is hard to import into existing software. Commonly used formats are * Nexus file format * Newick formatLimitations of phylogenetic analysis
Although phylogenetic trees produced on the basis of sequencedgene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protei ...
s or genomic data in different species can provide evolutionary insight, these analyses have important limitations. Most importantly, the trees that they generate are not necessarily correct – they do not necessarily accurately represent the evolutionary history of the included taxa. As with any scientific result, they are subject to falsification by further study (e.g., gathering of additional data, analyzing the existing data with improved methods). The data on which they are based may be noisy; the analysis can be confounded by genetic recombination
Genetic recombination (also known as genetic reshuffling) is the exchange of genetic material between different organisms which leads to production of offspring with combinations of traits that differ from those found in either parent. In eukaryot ...
, horizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). HGT is an important factor in the e ...
, hybridisation between species that were not nearest neighbors on the tree before hybridisation takes place, and conserved sequences.
Also, there are problems in basing an analysis on a single type of character, such as a single gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protei ...
or protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metab ...
or only on morphological analysis, because such trees constructed from another unrelated data source often differ from the first, and therefore great care is needed in inferring phylogenetic relationships among species. This is most true of genetic material that is subject to lateral gene transfer and recombination, where different haplotype blocks can have different histories. In these types of analysis, the output tree of a phylogenetic analysis of a single gene is an estimate of the gene's phylogeny (i.e. a gene tree) and not the phylogeny of the taxa (i.e. species tree) from which these characters were sampled, though ideally, both should be very close. For this reason, serious phylogenetic studies generally use a combination of genes that come from different genomic sources (e.g., from mitochondrial or plastid vs. nuclear genomes), or genes that would be expected to evolve under different selective regimes, so that homoplasy (false homology) would be unlikely to result from natural selection.
When extinct species are included as terminal nodes in an analysis (rather than, for example, to constrain internal nodes), they are considered not to represent direct ancestors of any extant species. Extinct species do not typically contain high-quality DNA.
The range of useful DNA materials has expanded with advances in extraction and sequencing technologies. Development of technologies able to infer sequences from smaller fragments, or from spatial patterns of DNA degradation products, would further expand the range of DNA considered useful.
Phylogenetic trees can also be inferred from a range of other data types, including morphology, the presence or absence of particular types of genes, insertion and deletion events – and any other observation thought to contain an evolutionary signal.
Phylogenetic networks are used when bifurcating trees are not suitable, due to these complications which suggest a more reticulate evolutionary history of the organisms sampled.
See also
*Clade
In biology, a clade (), also known as a Monophyly, monophyletic group or natural group, is a group of organisms that is composed of a common ancestor and all of its descendants. Clades are the fundamental unit of cladistics, a modern approach t ...
* Cladistics
Cladistics ( ; from Ancient Greek 'branch') is an approach to Taxonomy (biology), biological classification in which organisms are categorized in groups ("clades") based on hypotheses of most recent common ancestry. The evidence for hypothesiz ...
* Computational phylogenetics
Computational phylogenetics, phylogeny inference, or phylogenetic inference focuses on computational and optimization algorithms, Heuristic (computer science), heuristics, and approaches involved in Phylogenetics, phylogenetic analyses. The goal i ...
* Evolutionary biology
Evolutionary biology is the subfield of biology that studies the evolutionary processes such as natural selection, common descent, and speciation that produced the diversity of life on Earth. In the 1930s, the discipline of evolutionary biolo ...
* Evolutionary taxonomy
* Generalized tree alignment
* List of phylogenetics software
* List of phylogenetic tree visualization software
* PANDIT, a biological database covering protein domains
* Phylogenetic comparative methods
* Phylogenetic reconciliation
* Taxonomic rank
In biology, taxonomic rank (which some authors prefer to call nomenclatural rank because ranking is part of nomenclature rather than taxonomy proper, according to some definitions of these terms) is the relative or absolute level of a group of or ...
* Tokogeny
References
Further reading
* Schuh, R. T. and A. V. Z. Brower. 2009. ''Biological Systematics: principles and applications (2nd edn.)'' * Manuel Lima, ''The Book of Trees: Visualizing Branches of Knowledge'', 2014, Princeton Architectural Press, New York. * MEGA, a free software to draw phylogenetic trees. * Gontier, N. 2011. "Depicting the Tree of Life: the Philosophical and Historical Roots of Evolutionary Tree Diagrams." Evolution, Education, Outreach 4: 515–538. * Jan Sapp, ''The New Foundations of Evolution: On the Tree of Life'', 2009, Oxford University Press, New York.External links
Images
Human Y-Chromosome 2002 Phylogenetic Tree
iTOL: Interactive Tree Of Life
Phylogenetic Tree of Artificial Organisms Evolved on Computers
Miyamoto and Goodman's Phylogram of Eutherian Mammals
General
* An overview of different methods of tree visualization is available atOneZoom: Tree of Life – all living species as intuitive and zoomable fractal explorer (responsive design)
Discover Life
An interactive tree based on the U.S. National Science Foundation's Assembling the Tree of Life Project
* ttp://tolweb.org/tree Tree of Life Web Project
Phylogenetic inferring on the T-REX server
NCBI's Taxonomy Database
ETE: A Python Environment for Tree Exploration
This is a programming library to analyze, manipulate and visualize phylogenetic trees
Ref.
A daily-updated tree of (sequenced) life
{{DEFAULTSORT:Phylogenetic Tree Phylogenetics Trees (data structures)