Mycobacterium tuberculosis on:
[Wikipedia]
[Google]
[Amazon]
 ''Mycobacterium tuberculosis'' (M. tb) is a species of
''Mycobacterium tuberculosis'' (M. tb) is a species of

TB database: an integrated platform for Tuberculosis research
*
Database on Mycobacterium tuberculosis genetics
{{Authority control Acid-fast bacilli
 ''Mycobacterium tuberculosis'' (M. tb) is a species of
''Mycobacterium tuberculosis'' (M. tb) is a species of pathogenic bacteria
Pathogenic bacteria are bacteria that can cause disease. This article focuses on the bacteria that are pathogenic to humans. Most species of bacteria are harmless and are often Probiotic, beneficial but others can cause infectious diseases. The n ...
in the family Mycobacteriaceae
''Mycobacteriaceae'' is a family of bacteria in the phylum Actinomycetota
The ''Actinomycetota'' (or ''Actinobacteria'') are a phylum of all gram-positive bacteria. They can be terrestrial or aquatic. They are of great economic importance ...
and the causative agent
In biology, a pathogen ( el, πάθος, "suffering", "passion" and , "producer of") in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ ...
of tuberculosis
Tuberculosis (TB) is an infectious disease usually caused by '' Mycobacterium tuberculosis'' (MTB) bacteria. Tuberculosis generally affects the lungs, but it can also affect other parts of the body. Most infections show no symptoms, in ...
. First discovered in 1882 by Robert Koch
Heinrich Hermann Robert Koch ( , ; 11 December 1843 – 27 May 1910) was a German physician and microbiologist. As the discoverer of the specific causative agents of deadly infectious diseases including tuberculosis, cholera (though the Vibrio ...
, ''M. tuberculosis'' has an unusual, waxy coating on its cell surface primarily due to the presence of mycolic acid
Mycolic acids are long fatty acids found in the cell walls of the Mycolata taxon, a group of bacteria that includes ''Mycobacterium tuberculosis'', the causative agent of the disease tuberculosis. They form the major component of the cell wall of ...
. This coating makes the cells impervious to Gram staining
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish bac ...
, and as a result, ''M. tuberculosis'' can appear weakly Gram-positive. Acid-fast
Acid-fastness is a physical property of certain bacterial and eukaryotic cells, as well as some sub-cellular structures, specifically their resistance to decolorization by acids during laboratory staining procedures. Once stained as part of a sam ...
stains such as Ziehl–Neelsen, or fluorescent
Fluorescence is the emission of light by a substance that has absorbed light or other electromagnetic radiation. It is a form of luminescence. In most cases, the emitted light has a longer wavelength, and therefore a lower photon energy, tha ...
stains such as auramine are used instead to identify ''M. tuberculosis'' with a microscope. The physiology of ''M. tuberculosis'' is highly aerobic
Aerobic means "requiring air," in which "air" usually means oxygen.
Aerobic may also refer to
* Aerobic exercise, prolonged exercise of moderate intensity
* Aerobics, a form of aerobic exercise
* Aerobic respiration, the aerobic process of cel ...
and requires high levels of oxygen. Primarily a pathogen of the mammalian respiratory system
The respiratory system (also respiratory apparatus, ventilatory system) is a biological system consisting of specific organs and structures used for gas exchange in animals and plants. The anatomy and physiology that make this happen varies grea ...
, it infects the lungs. The most frequently used diagnostic methods for tuberculosis are the tuberculin skin test
The Mantoux test or Mendel–Mantoux test (also known as the Mantoux screening test, tuberculin sensitivity test, Pirquet test, or PPD test for purified protein derivative) is a tool for screening for tuberculosis (TB) and for tuberculosis diagn ...
, acid-fast stain
Acid-fastness is a physical property of certain bacterial and Eukaryote, eukaryotic cell (biology), cells, as well as some Sub-cellular, sub-cellular structures, specifically their resistance to decolorization by acids during laboratory staining p ...
, culture
Culture () is an umbrella term which encompasses the social behavior, institutions, and norms found in human societies, as well as the knowledge, beliefs, arts, laws, customs, capabilities, and habits of the individuals in these groups.Tyl ...
, and polymerase chain reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) t ...
.
The ''M. tuberculosis'' genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding ge ...
was sequenced
In genetics and biochemistry, sequencing means to determine the primary structure (sometimes incorrectly called the primary sequence) of an unbranched biopolymer. Sequencing results in a symbolic linear depiction known as a sequence which suc ...
in 1998.
Microbiology
In 2019, M. tuberculosis was found in a genetically related complex group of Mycobacterium species calledMycobacterium tuberculosis complex
The ''Mycobacterium tuberculosis'' complex (MTC or MTBC) is a genetically related group of ''Mycobacterium'' species that can cause tuberculosis in humans or other animals.
It includes:
* ''Mycobacterium tuberculosis''
* '' Mycobacterium african ...
that has at least 9 members:
*''M. tuberculosis'' ''sensu stricto''
*'' M. africanum''
*'' M. canetti''
*'' M. bovis''
*'' M. caprae''
*'' M. microti''
*'' M. pinnipedii''
*'' M. mungi''
*'' M. orygis''
It requires oxygen to grow, and is nonmotile. ''M. tuberculosis'' divides every 18–24 hours. This is extremely slow compared with other bacteria, which tend to have division times measured in minutes (''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escher ...
'' can divide roughly every 20 minutes). It is a small bacillus
''Bacillus'' (Latin "stick") is a genus of Gram-positive, rod-shaped bacteria, a member of the phylum ''Bacillota'', with 266 named species. The term is also used to describe the shape (rod) of other so-shaped bacteria; and the plural ''Bacilli ...
that can withstand weak disinfectant
A disinfectant is a chemical substance or compound used to inactivate or destroy microorganisms on inert surfaces. Disinfection does not necessarily kill all microorganisms, especially resistant bacterial spores; it is less effective than st ...
s and can survive in a dry state for weeks. Its unusual cell wall is rich in lipids
Lipids are a broad group of naturally-occurring molecules which includes fats, waxes, sterols, fat-soluble vitamins (such as vitamins A, D, E and K), monoglycerides, diglycerides, phospholipids, and others. The functions of lipids include ...
such as mycolic acid
Mycolic acids are long fatty acids found in the cell walls of the Mycolata taxon, a group of bacteria that includes ''Mycobacterium tuberculosis'', the causative agent of the disease tuberculosis. They form the major component of the cell wall of ...
and cord factor
Cord factor, or trehalose dimycolate, is a glycolipid molecule found in the cell wall of '' Mycobacterium tuberculosis'' and similar species. It is the primary lipid found on the exterior of ''M. tuberculosis'' cells. Cord factor influences the ...
glycolipid
Glycolipids are lipids with a carbohydrate attached by a glycosidic (covalent) bond. Their role is to maintain the stability of the cell membrane and to facilitate cellular recognition, which is crucial to the immune response and in the connec ...
, is likely responsible for its resistance to desiccation
Desiccation () is the state of extreme dryness, or the process of extreme drying. A desiccant is a hygroscopic (attracts and holds water) substance that induces or sustains such a state in its local vicinity in a moderately sealed container.
...
and is a key virulence factor
Virulence factors (preferably known as pathogenicity factors or effectors in plant science) are cellular structures, molecules and regulatory systems that enable microbial pathogens (bacteria, viruses, fungi, and protozoa) to achieve the following ...
.
Microscopy
Other bacteria are commonly identified with a microscope by staining them withGram stain
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish ...
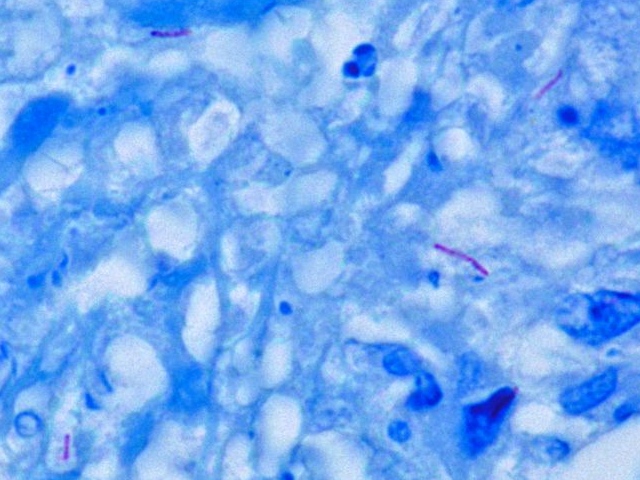
. However, the mycolic acid in the cell wall of ''M. tuberculosis'' does not absorb the stain. Instead, acid-fast stains such as Ziehl–Neelsen stain
Ziehl–Neelsen staining is a type of acid-fast stain, first introduced by Paul Ehrlich. Ziehl–Neelsen staining is a bacteriological stain used to identify acid-fast organisms, mainly Mycobacteria. It is named for two German doctors who modif ...
, or fluorescent stains such as auramine are used. Cells are curved rod-shaped and are often seen wrapped together, due to the presence of fatty acids in the cell wall that stick together. This appearance is referred to as cording, like strands of cord that make up a rope. ''M. tuberculosis'' is characterized in tissue by caseating granulomas
A granuloma is an aggregation of macrophages that forms in response to chronic inflammation. This occurs when the immune system attempts to isolate foreign substances that it is otherwise unable to eliminate. Such substances include infectious o ...
containing Langhans giant cell
Langhans giant cells are large cells found in granulomatous conditions.
They are formed by the fusion of epithelioid cells ( macrophages), and contain nuclei arranged in a horseshoe-shaped pattern in the cell periphery.
Although traditionally ...
s, which have a "horseshoe" pattern of nuclei.
Culture
''M. tuberculosis'' can be grown in the laboratory. Compared to other commonly studied bacteria, ''M. tuberculosis'' has a remarkably slow growth rate, doubling roughly once per day. Commonly usedmedia
Media may refer to:
Communication
* Media (communication), tools used to deliver information or data
** Advertising media, various media, content, buying and placement for advertising
** Broadcast media, communications delivered over mass el ...
include liquids such as Middlebrook 7H9 or 7H12, egg-based solid media such as Lowenstein-Jensen, and solid agar-based such as Middlebrook 7H11 or 7H10. Visible colonies require several weeks to grow on agar plates. It is distinguished from other mycobacteria by its production of catalase
Catalase is a common enzyme found in nearly all living organisms exposed to oxygen (such as bacteria, plants, and animals) which catalyzes the decomposition of hydrogen peroxide to water and oxygen. It is a very important enzyme in protecting t ...
and niacin
Niacin, also known as nicotinic acid, is an organic compound and a form of vitamin B3, an essential human nutrient. It can be manufactured by plants and animals from the amino acid tryptophan. Niacin is obtained in the diet from a variet ...
. Other tests to confirm its identity include gene probe
In molecular biology, a hybridization probe (HP) is a fragment of DNA or RNA of usually 15–10000 nucleotide long which can be radioactively or fluorescently labeled. HP can be used to detect the presence of nucleotide sequences in analyzed RN ...
s and MALDI-TOF
In mass spectrometry, matrix-assisted laser desorption/ionization (MALDI) is an ionization technique that uses a laser energy absorbing matrix to create ions from large molecules with minimal fragmentation. It has been applied to the analysis of ...
.
Morphology
The slices of the ''Mycobacterium tuberculosis'' analyzed under a scanning electron microscope by a Japan-based research group has revealed that bacteria is about 2.71 ± 1.05μm in length with an average diameter of the cell approximately 0.345 ± 0.029 μm. The outer membrane and plasma membrane surface areas were measured to be 3.04 ± 1.33 μm2 and 2.67 ± 1.19 μm2, respectively. The cell, outer membrane, periplasm, plasma membrane, and cytoplasm volumes were 0.293 ± 0.113 fl (= μm3), 0.006 ± 0.003 fl, 0.060 ± 0.021 fl, 0.019 ± 0.008 fl, and 0.210 ± 0.091 fl, respectively. The average totalribosome
Ribosomes ( ) are macromolecular machines, found within all cells, that perform biological protein synthesis (mRNA translation). Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to ...
number was 1,672 ± 568 with ribosome density about 716.5 ± 171.4/0.1 fl.
Pathophysiology
Humans are the only known reservoirs of ''M. tuberculosis''. A misconception is that ''M. tuberculosis'' can be spread by shaking hands, making contact with toilet seats, sharing food or drink, or sharing toothbrushes. However, major spread is throughair droplets
An aerosol is a suspension (chemistry), suspension of fine solid particles or liquid Drop (liquid), droplets in air or another gas. Aerosols can be natural or Human impact on the environment, anthropogenic. Examples of natural aerosols are fog o ...
originating from a person who has the disease either coughing, sneezing, speaking, or singing.
When in the lungs, ''M. tuberculosis'' is phagocytosed
Phagocytosis () is the process by which a cell uses its plasma membrane to engulf a large particle (≥ 0.5 μm), giving rise to an internal compartment called the phagosome. It is one type of endocytosis. A cell that performs phagocytosis is c ...
by alveolar macrophage
An alveolar macrophage, pulmonary macrophage, (or dust cell) is a type of macrophage, a professional phagocyte, found in the airways and at the level of the alveoli in the lungs, but separated from their walls.
Activity of the alveolar macropha ...
s, but they are unable to kill and digest the bacterium. Its cell wall that is made of a cord factor
Cord factor, or trehalose dimycolate, is a glycolipid molecule found in the cell wall of '' Mycobacterium tuberculosis'' and similar species. It is the primary lipid found on the exterior of ''M. tuberculosis'' cells. Cord factor influences the ...
glycoplipds that inhibit the fusion of the phagosome
In cell biology, a phagosome is a vesicle formed around a particle engulfed by a phagocyte via phagocytosis. Professional phagocytes include macrophages, neutrophils, and dendritic cells (DCs).
A phagosome is formed by the fusion of the cell mem ...
with the lysosome
A lysosome () is a membrane-bound organelle found in many animal cells. They are spherical vesicles that contain hydrolytic enzymes that can break down many kinds of biomolecules. A lysosome has a specific composition, of both its membrane prot ...
, which contains a host of antibacterial factors.
Specifically, ''M. tuberculosis'' blocks the bridging molecule, early endosomal autoantigen 1 ( EEA1); however, this blockade does not prevent fusion of vesicles filled with nutrients. In addition, production of the diterpene isotuberculosinol
Isotuberculosinol, also called nosyberkol or edaxadiene is a diterpene molecule produced by the bacterium '' Mycobacterium tuberculosis'', the causative agent of TB, which aids in its pathogenesis. Isotuberculosinol functions by preventing mat ...
prevents maturation of the phagosome. The bacteria also evades macrophage-killing by neutralizing reactive nitrogen intermediates. More recently, ''M. tuberculosis'' has been shown to secrete and cover itself in 1-tuberculosinyladenosine (1-TbAd), a special nucleoside
Nucleosides are glycosylamines that can be thought of as nucleotides without a phosphate group. A nucleoside consists simply of a nucleobase (also termed a nitrogenous base) and a five-carbon sugar (ribose or 2'-deoxyribose) whereas a nucleotide ...
that acts as an antacid
An antacid is a substance which neutralizes stomach acidity and is used to relieve heartburn, indigestion or an upset stomach. Some antacids have been used in the treatment of constipation and diarrhea. Marketed antacids contain salts of alumi ...
, allowing it to neutralize pH and induce swelling in lysosomes.
In ''M. tuberculosis'' infections, PPM1A levels were found to be upregulated, and this, in turn, would impact the normal apoptotic response of macrophages to clear pathogens, as PPM1A is involved in the intrinsic and extrinsic apoptotic pathways. Hence, when PPM1A levels were increased, the expression of it inhibits the two apoptotic pathways. With kinome analysis, the JNK/AP-1 signalling pathway was found to be a downstream effector that PPM1A has a part to play in, and the apoptotic pathway in macrophages are controlled in this manner. As a result of having apoptosis being suppressed, it provides ''M. tuberculosis'' with a safe replicative niche, and so the bacteria are able to maintain a latent state for a prolonged time.
Granuloma
A granuloma is an aggregation of macrophages that forms in response to chronic inflammation. This occurs when the immune system attempts to isolate foreign substances that it is otherwise unable to eliminate. Such substances include infectious ...
s, organized aggregates of immune cells, are a hallmark feature of tuberculosis infection. Granulomas play dual roles during infection: they regulate the immune response and minimize tissue damage, but also can aid in the expansion of infection.
The ability to construct ''M. tuberculosis'' mutants and test individual gene products for specific functions has significantly advanced the understanding of its pathogenesis
Pathogenesis is the process by which a disease or disorder develops. It can include factors which contribute not only to the onset of the disease or disorder, but also to its progression and maintenance. The word comes from Greek πάθος ''pat ...
and virulence factors
Virulence factors (preferably known as pathogenicity factors or effectors in plant science) are cellular structures, molecules and regulatory systems that enable microbial pathogens (bacteria, viruses, fungi, and protozoa) to achieve the followin ...
. Many secreted and exported proteins are known to be important in pathogenesis. For example, one such virulence factor is cord factor
Cord factor, or trehalose dimycolate, is a glycolipid molecule found in the cell wall of '' Mycobacterium tuberculosis'' and similar species. It is the primary lipid found on the exterior of ''M. tuberculosis'' cells. Cord factor influences the ...
(trehalose dimycolate), which serves to increase survival within its host. Resistant strains of ''M. tuberculosis'' have developed resistance to more than one TB drug, due to mutations in their genes. In addition, pre-existing first-line TB drugs such as rifampicin and streptomycin have decreased efficiency in clearing intracellular
This glossary of biology terms is a list of definitions of fundamental terms and concepts used in biology, the study of life and of living organisms. It is intended as introductory material for novices; for more specific and technical definitions ...
''M. tuberculosis'' due not being able to effectively penetrate the macrophage niche.
JNK plays a key role in the control of apoptotic pathways—intrinsic and extrinsic. In addition, it is also found to be a substrate of PPM1A activity, hence the phosphorylation of JNK would cause apoptosis to occur. Since PPM1A levels are elevated during ''M. tuberculosis'' infections, by inhibiting the PPM1A signalling pathways, it could potentially be a therapeutic method to kill ''M. tuberculosis''-infected macrophages by restoring its normal apoptotic function in defence of pathogens. By targeting the PPM1A-JNK signalling axis pathway, then, it could eliminate ''M. tuberculosis''-infected macrophages.
The ability to restore macrophage apoptosis to ''M. tuberculosis''-infected ones could improve the current tuberculosis chemotherapy treatment, as TB drugs can gain better access to the bacteria in the niche. thus decreasing the treatment times for ''M. tuberculosis'' infections.
Symptoms of ''M. tuberculosis'' include coughing that lasts for more than three weeks, hemoptysis
Hemoptysis is the coughing up of blood or blood-stained mucus from the bronchi, larynx, trachea, or lungs. In other words, it is the airway bleeding. This can occur with lung cancer, infections such as tuberculosis, bronchitis, or pneumonia, and ...
, chest pain when breathing or coughing, weight loss, fatigue, fever, night sweats, chills, and loss of appetite. ''M. tuberculosis'' also has the potential of spreading to other parts of the body. This can cause blood in urine if the kidneys are affected, and back pain if the spine is affected.
Strain variation
Typing of strains is useful in the investigation of tuberculosis outbreaks, because it gives the investigator evidence for or against transmission from person to person. Consider the situation where person A has tuberculosis and believes he acquired it from person B. If the bacteria isolated from each person belong to different types, then transmission from B to A is definitively disproven; however, if the bacteria are the same strain, then this supports (but does not definitively prove) the hypothesis that B infected A. Until the early 2000s, ''M. tuberculosis'' strains were typed bypulsed field gel electrophoresis
Pulsed field gel electrophoresis is a technique used for the separation of large DNA molecules by applying to a gel matrix an electric field that periodically changes direction.
Historical background
Standard gel electrophoresis techniques for s ...
. This has now been superseded by variable numbers of tandem repeats (VNTR), which is technically easier to perform and allows better discrimination between strains. This method makes use of the presence of repeated DNA sequences within the ''M. tuberculosis'' genome.
Three generations of VNTR typing for ''M. tuberculosis'' are noted. The first scheme, called exact tandem repeat, used only five loci, but the resolution afforded by these five loci was not as good as PFGE. The second scheme, called mycobacterial interspersed repetitive unit, had discrimination as good as PFGE. The third generation (mycobacterial interspersed repetitive unit – 2) added a further nine loci to bring the total to 24. This provides a degree of resolution greater than PFGE and is currently the standard for typing ''M. tuberculosis''. However, with regard to archaeological remains, additional evidence may be required because of possible contamination from related soil bacteria.
Antibiotic resistance in ''M. tuberculosis'' typically occurs due to either the accumulation of mutations in the genes targeted by the antibiotic or a change in titration of the drug. ''M. tuberculosis'' is considered to be multidrug-resistant (MDR TB) if it has developed drug resistance to both rifampicin and isoniazid, which are the most important antibiotics used in treatment. Additionally, extensively drug-resistant ''M. tuberculosis'' (XDR TB) is characterized by resistance to both isoniazid and rifampin, plus any fluoroquinolone and at least one of three injectable second-line drugs (i.e., amikacin, kanamycin, or capreomycin).


Genome
The genome of theH37Rv
Mycobacterium tuberculosis strain H37Rv is the most studied strain of tuberculosis in research laboratories. It was first isolated by Dr. Edward R. Baldwin in 1905. The strain came from a 19 year old patient with chronic pulmonary tuberculosis at ...
strain was published in 1998. Its size is 4 million base pairs, with 3,959 genes; 40% of these genes have had their function characterized, with possible function postulated for another 44%. Within the genome are also six pseudogene
Pseudogenes are nonfunctional segments of DNA that resemble functional genes. Most arise as superfluous copies of functional genes, either directly by DNA duplication or indirectly by Reverse transcriptase, reverse transcription of an mRNA trans ...
s.
The genome contains 250 genes involved in fatty acid
In chemistry, particularly in biochemistry, a fatty acid is a carboxylic acid with an aliphatic chain, which is either saturated or unsaturated. Most naturally occurring fatty acids have an unbranched chain of an even number of carbon atoms, fr ...
metabolism, with 39 of these involved in the polyketide
Polyketides are a class of natural products derived from a precursor molecule consisting of a chain of alternating ketone (or reduced forms of a ketone) and methylene groups: (-CO-CH2-). First studied in the early 20th century, discovery, biosynth ...
metabolism generating the waxy coat. Such large numbers of conserved genes show the evolutionary importance of the waxy coat to pathogen survival. Furthermore, experimental studies have since validated the importance of a lipid metabolism for'' M. tuberculosis'', consisting entirely of host-derived lipids such as fats and cholesterol. Bacteria isolated from the lungs of infected mice were shown to preferentially use fatty acids over carbohydrate substrates. ''M. tuberculosis'' can also grow on the lipid cholesterol
Cholesterol is any of a class of certain organic molecules called lipids. It is a sterol (or modified steroid), a type of lipid. Cholesterol is biosynthesized by all animal cells and is an essential structural component of animal cell mem ...
as a sole source of carbon, and genes involved in the cholesterol use pathway(s) have been validated as important during various stages of the infection lifecycle of ''M. tuberculosis'', especially during the chronic phase of infection when other nutrients are likely not available.
About 10% of the coding capacity is taken up by the ''PE''/''PPE'' gene families that encode acidic, glycine-rich proteins. These proteins have a conserved N-terminal motif, deletion of which impairs growth in macrophages and granulomas.
Nine noncoding sRNAs have been characterised in ''M. tuberculosis'', with a further 56 predicted in a bioinformatics
Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combi ...
screen.
In 2013, a study on the genome of several sensitive, ultraresistant, and multiresistant ''M. tuberculosis'' strains was made to study antibiotic resistance mechanisms. Results reveal new relationships and drug resistance genes not previously associated and suggest some genes and intergenic regions associated with drug resistance may be involved in the resistance to more than one drug. Noteworthy is the role of the intergenic regions in the development of this resistance, and most of the genes proposed in this study to be responsible for drug resistance have an essential role in the development of ''M. tuberculosis''.
Evolution
The ''M. tuberculosis'' complex evolved in Africa and most probably in theHorn of Africa
The Horn of Africa (HoA), also known as the Somali Peninsula, is a large peninsula and geopolitical region in East Africa.Robert Stock, ''Africa South of the Sahara, Second Edition: A Geographical Interpretation'', (The Guilford Press; 2004), ...
. In addition to ''M. tuberculosis'', the ''M. tuberculosis'' complex (MTBC) has a number of members infecting various animal species, these include ''M. africanum'', ''M. bovis'' (Dassie's bacillus), ''M. caprae'', ''M. microti'', ''M. mungi, M. orygis'', and ''M. pinnipedii''. This group may also include the ''M. canettii'' clade. These animal strains of MTBC do not strictly deserve species status, as they are all closely related and embedded in the ''M. tuberculosis'' phylogeny, but for historic reasons, they currently hold species status.
The ''M. canettii'' clade – which includes ''M. prototuberculosis'' – is a group of smooth-colony ''Mycobacterium'' species. Unlike the established members of the ''M. tuberculosis'' group, they undergo recombination with other species. The majority of the known strains of this group have been isolated from the Horn of Africa. The ancestor of ''M. tuberculosis'' appears to be ''M. canettii'', first described in 1969.
The established members of the ''M. tuberculosis'' complex are all clonal in their spread. The main human-infecting species have been classified into seven lineages. Translating these lineages into the terminology used for spoligotyping, a very crude genotyping methodology, lineage 1 contains the East Africa
East Africa, Eastern Africa, or East of Africa, is the eastern subregion of the African continent. In the United Nations Statistics Division scheme of geographic regions, 10-11-(16*) territories make up Eastern Africa:
Due to the historical ...
n-India
India, officially the Republic of India (Hindi: ), is a country in South Asia. It is the seventh-largest country by area, the second-most populous country, and the most populous democracy in the world. Bounded by the Indian Ocean on the so ...
n (EAI), the Manila family of strains and some Manu (Indian) strains; lineage 2 is the Beijing
}
Beijing ( ; ; ), alternatively romanized as Peking ( ), is the capital of the People's Republic of China. It is the center of power and development of the country. Beijing is the world's most populous national capital city, with over 21 ...
group; lineage 3 includes the Central Asia
Central Asia, also known as Middle Asia, is a subregion, region of Asia that stretches from the Caspian Sea in the west to western China and Mongolia in the east, and from Afghanistan and Iran in the south to Russia in the north. It includes t ...
n (CAS) strains; lineage 4 includes the Ghana
Ghana (; tw, Gaana, ee, Gana), officially the Republic of Ghana, is a country in West Africa. It abuts the Gulf of Guinea and the Atlantic Ocean to the south, sharing borders with Ivory Coast in the west, Burkina Faso in the north, and To ...
and Haarlem
Haarlem (; predecessor of ''Harlem'' in English) is a city and municipality in the Netherlands. It is the capital of the province of North Holland. Haarlem is situated at the northern edge of the Randstad, one of the most populated metropoli ...
(H/T), Latin America
Latin America or
* french: Amérique Latine, link=no
* ht, Amerik Latin, link=no
* pt, América Latina, link=no, name=a, sometimes referred to as LatAm is a large cultural region in the Americas where Romance languages — languages derived f ...
-Mediterranean
The Mediterranean Sea is a sea connected to the Atlantic Ocean, surrounded by the Mediterranean Basin and almost completely enclosed by land: on the north by Western and Southern Europe and Anatolia, on the south by North Africa, and on the e ...
(LAM) and X strains; types 5 and 6 correspond to ''M. africanum'' and are observed predominantly and at high frequencies in West Africa
West Africa or Western Africa is the westernmost region of Africa. The United Nations defines Western Africa as the 16 countries of Benin, Burkina Faso, Cape Verde, The Gambia, Ghana, Guinea, Guinea-Bissau, Ivory Coast, Liberia, Mali, Maurit ...
. A seventh type has been isolated from the Horn of Africa. The other species of this complex belong to a number of spoligotypes and do not normally infect humans.
Lineages 2, 3 and 4 all share a unique deletion event (tbD1) and thus form a monophyletic group. Types 5 and 6 are closely related to the animal strains of MTBC, which do not normally infect humans. Lineage 3 has been divided into two clades: CAS-Kili (found in Tanzania
Tanzania (; ), officially the United Republic of Tanzania ( sw, Jamhuri ya Muungano wa Tanzania), is a country in East Africa within the African Great Lakes region. It borders Uganda to the north; Kenya to the northeast; Comoro Islands and ...
) and CAS-Delhi (found in India and Saudi Arabia
Saudi Arabia, officially the Kingdom of Saudi Arabia (KSA), is a country in Western Asia. It covers the bulk of the Arabian Peninsula, and has a land area of about , making it the fifth-largest country in Asia, the second-largest in the A ...
).
Lineage 4 is also known as the Euro-American lineage. Subtypes within this type include Latin American Mediterranean, Uganda I, Uganda II, Haarlem, X, and Congo.
A much cited study reported that ''M. tuberculosis'' has co-evolved with human populations, and that the most recent common ancestor
In biology and genetic genealogy, the most recent common ancestor (MRCA), also known as the last common ancestor (LCA) or concestor, of a set of organisms is the most recent individual from which all the organisms of the set are descended. The ...
of the ''M. tuberculosis'' complex evolved between 40,000 and 70,000 years ago. However, a later study that included genome sequences from ''M. tuberculosis'' complex members extracted from three 1,000-year-old Peruvian mummies, came to quite different conclusions. If the most recent common ancestor
In biology and genetic genealogy, the most recent common ancestor (MRCA), also known as the last common ancestor (LCA) or concestor, of a set of organisms is the most recent individual from which all the organisms of the set are descended. The ...
of the ''M. tuberculosis'' complex were 40,000 to 70,000 years old, this would necessitate an evolutionary rate much lower than any estimates produced by genomic analyses of heterochronous samples, suggesting a far more recent common ancestor of the ''M. tuberculosis'' complex as little as 6000 years ago.
An analysis of over 3000 strains of ''M. bovis'' from 35 countries suggested an Africa origin for this species.Loiseau C, Menardo F, Aseffa A, Hailu E, Gumi B, Ameni G, Berg S, Rigouts L, Robbe-Austerman S, Zinsstag J, Gagneux S, Brites D (2020) An African origin for ''Mycobacterium bovis''. Evol Med Public Health. 2020 Jan 31;2020(1):49–59
Co-evolution with modern humans
There are currently two narratives existing in parallel regarding the age of MTBC and how it has spread and co-evolved with humans through time. One study compared the ''M. tuberculosis'' phylogeny to a human mitochondrial genome phylogeny and interpreted these as being highly similar. Based on this, the study suggested that ''M. tuberculosis'', like humans, evolved in Africa and subsequently spread with anatomically modern humans out of Africa across the world. By calibrating the mutation rate of M. tuberculosis to match this narrative, the study suggested that MTBC evolved 40,000–70,000 years ago. Applying this time scale, the study found that the ''M. tuberculosis''effective population size
The effective population size (''N'e'') is a number that, in some simplified scenarios, corresponds to the number of breeding individuals in the population. More generally, ''N'e'' is the number of individuals that an idealised population wo ...
expanded during the Neolithic Demographic Transition
The Neolithic demographic transition was a period of rapid population growth following the adoption of agriculture by prehistoric societies (the Neolithic Revolution). It was a demographic transition caused by an abrupt increase in birth rates du ...
(around 10,000 years ago) and suggested that ''M. tuberculosis'' was able to adapt to changing human populations and that the historical success of this pathogen was driven at least in part by dramatic increases in human host population density. It has also been demonstrated that after emigrating from one continent to another, a human host's region of origin is predictive of which TB lineage they carry, which could reflect either a stable association between host populations and specific ''M. tuberculosis'' lineages and/or social interactions that are shaped by shared cultural and geographic histories.
Regarding the congruence between human and ''M. tuberculosis'' phylogenies, a study relying on ''M. tuberculosis'' and human Y chromosome
The Y chromosome is one of two sex chromosomes (allosomes) in therian mammals, including humans, and many other animals. The other is the X chromosome. Y is normally the sex-determining chromosome in many species, since it is the presence or abse ...
DNA sequences to formally assess the correlation between them, concluded that they are not congruent. Also, a more recent study which included genome sequences from ''M. tuberculosis'' complex members extracted from three 1,000-year-old Peruvian mummies, estimated that the most recent common ancestor
In biology and genetic genealogy, the most recent common ancestor (MRCA), also known as the last common ancestor (LCA) or concestor, of a set of organisms is the most recent individual from which all the organisms of the set are descended. The ...
of the ''M. tuberculosis'' complex lived only 4,000 – 6,000 years ago. The ''M. tuberculosis'' evolutionary rate estimated by the Bos et al. study is also supported by a study on Lineage 4 relying on genomic aDNA sequences from Hungarian mummies more than 200 years old. In total, the evidence thus favors this more recent estimate of the age of the MTBC most recent common ancestor, and thus that the global evolution and dispersal of ''M. tuberculosis'' has occurred over the last 4,000–6,000 years.
Among the seven recognized lineages of ''M. tuberculosis'', only two are truly global in their distribution: Lineages 2 and 4. Among these, Lineage 4 is the most well dispersed, and almost totally dominates in the Americas. Lineage 4 was shown to have evolved in or in the vicinity of Europe, and to have spread globally with Europeans starting around the 13th century. This study also found that Lineage 4 tuberculosis spread to the Americas shortly after the European discovery of the continent in 1492, and suggests that this represented the first introduction of human TB on the continent (although animal strains have been found in human remains predating Columbus. Similarly, Lineage 4 was found to have spread from Europe to Africa during the Age of Discovery
The Age of Discovery (or the Age of Exploration), also known as the early modern period, was a period largely overlapping with the Age of Sail, approximately from the 15th century to the 17th century in European history, during which seafarin ...
, starting in the early 15th century.
It has been suggested that ancestral mycobacteria may have infected early hominids in East Africa as early as three million years ago.
DNA fragments from ''M. tuberculosis'' and tuberculosis disease indications were present in human bodies dating from 9250 to 8160 years ago found at Atlit-Yam
Atlit Yam is an ancient submerged Neolithic village off the coast of Atlit, Israel. It has been carbon-dated as to be between 8,900 and 8,300 years old. Among the features of the 10-acre site is a stone circle.
History
Atlit-Yam provides the ear ...
in the Levant
The Levant () is an approximate historical geographical term referring to a large area in the Eastern Mediterranean region of Western Asia. In its narrowest sense, which is in use today in archaeology and other cultural contexts, it is eq ...
.
Antibiotic resistance (ABR)
''M. tuberculosis'' is a clonal organism and does not exchange DNA viahorizontal gene transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between Unicellular organism, unicellular and/or multicellular organisms other than by the ("vertical") transmission of DNA from parent to offsprin ...
. Despite an additionally slow evolution rate, the emergence and spread of antibiotic resistance in ''M. tuberculosis'' poses an increasing threat to global public health. In 2019, the WHO reported the estimated incidence of antibiotic resistant TB to be 3.4% in new cases, and 18% in previously treated cases. Geographical discrepancies exist in the incidence rates of drug-resistant TB. Countries facing the highest rates of ABR TB China, India, Russia, and South Africa. Recent trends reveal an increase in drug-resistant cases in a number of regions, with Papua New Guinea, Singapore, and Australia undergoing significant increases.
Multidrug-resistant Tuberculosis (MDR-TB) is characterised by resistance to at least the two front-line drugs isoniazid
Isoniazid, also known as isonicotinic acid hydrazide (INH), is an antibiotic used for the treatment of tuberculosis. For active tuberculosis it is often used together with rifampicin, pyrazinamide, and either streptomycin or ethambutol. For l ...
and rifampin
Rifampicin, also known as rifampin, is an ansamycin antibiotic used to treat several types of bacterial infections, including tuberculosis (TB), ''Mycobacterium avium'' complex, leprosy, and Legionnaires’ disease. It is almost always used tog ...
. MDR is associated with a relatively poor treatment success rate of 52%. Isoniazid and rifampin resistance are tightly linked, with 78% of the reported rifampin-resistant TB cases in 2019 being resistant to isoniazid as well. Rifampin-resistance is primarily due to resistance-conferring mutations in the rifampin-resistance determining region (RRDR) within the rpoB gene. The most frequently observed mutations of the codons in RRDR are 531, 526 and 516. However, alternative more elusive resistance-conferring mutations have been detected. Isoniazid function occurs through the inhibition of mycolic acid synthesis through the NADH-dependent enoyl-acyl carrier protein (ACP)-reductase. This is encoded by the ''inhA'' gene. As a result, isoniazid resistance is primarily due to mutations within inhA and the KatG gene or its promoter region - a catalase peroxidase which is required to activate Isoniazid. As MDR in ''M. tuberculosis'' becomes increasingly common, the emergence of pre-extensively drug resistant (pre-XDR) and extensively drug resistant (XDR-) TB threatens to exasperate the public health crisis. XDR-TB is characterised by resistance to both rifampin and Isoniazid, as well second-line fluoroquinolones and at least one additional front-line drug. Thus, the development of alternative therapeutic measures is of utmost priority.
An intrinsic contributor to the antibiotic resistant nature of ''M. tuberculosis'' is its unique cell wall. Saturated with long-chain fatty acids or mycolic acids, the mycobacterial cell presents a robust, relatively insoluble barrier. This has led to its synthesis being the target of many antibiotics - such as Isoniazid. However, resistance has emerged to the majority of them. A novel, promising therapeutic target is mycobacterial membrane protein large 3 (MmpL3). The mycobacterial membrane protein large (MmpL) proteins are transmembrane proteins which play a key role in the synthesis of the cell wall and the transport of the associated lipids. Of these, MmpL3 is essential; knock-out of which has been shown to be bactericidal. Due to its essential nature, MmpL3 inhibitors show promise as alternative therapeutic measures in the age of antibiotic resistance. Inhibition of MmpL3 function showed an inability to transport trehalose monomycolate - an essential cell wall lipid - across the plasma membrane. The recently reported structure of MmpL3 revealed resistance-conferring mutations to associate primarily with the transmembrane domain. Although resistance to pre-clinical MmpL3 inhibitors has been detected, analysis of the widespread mutational landscape revealed a low level of environmental resistance. This suggests that MmpL3 inhibitors currently undergoing clinical trials would face little resistance if made available. Additionally, the ability of many MmpL3 inhibitors to work synergistically with other antitubercular drugs presents a ray of hope in combatting the TB crisis.
Host genetics
The nature of the host-pathogen interaction between humans and ''M. tuberculosis'' is considered to have a genetic component. A group of rare disorders called Mendelian susceptibility to mycobacterial diseases was observed in a subset of individuals with a genetic defect that results in increased susceptibility to mycobacterial infection. Early case and twin studies have indicated that genetic components are important in host susceptibility to ''M. tuberculosis''. Recent genome-wide association studies (GWAS) have identified three genetic risk loci, including at positions 11p13 and 18q11. As is common in GWAS, the variants discovered have moderate effect sizes.DNA repair
As anintracellular pathogen
Intracellular parasites are microparasites that are capable of growing and reproducing inside the cells of a host.
Types of parasites
There are two main types of intracellular parasites: Facultative and Obligate.
Facultative intracellular para ...
, ''M. tuberculosis'' is exposed to a variety of DNA-damaging assaults, primarily from host-generated antimicrobial toxic radicals. Exposure to reactive oxygen species and/or reactive nitrogen species causes different types of DNA damage including oxidation, depurination, methylation, and deamination that can give rise to single- and double-strand breaks (DSBs).
DnaE2 polymerase is upregulated in ''M. tuberculosis'' by several DNA-damaging agents, as well as during infection of mice. Loss of this DNA polymerase reduces the virulence of ''M. tuberculosis'' in mice. DnaE2 is an error-prone DNA repair polymerase that appears to contribute to ''M. tuberculosis'' survival during infection.
The two major pathways employed in repair of DSBs are homologous recombination
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids (usually DNA as in cellular organisms but may ...
al repair (HR) and nonhomologous end joining
Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. NHEJ is referred to as "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology direc ...
(NHEJ). Macrophage-internalized ''M. tuberculosis'' is able to persist if either of these pathways is defective, but is attenuated when both pathways are defective. This indicates that intracellular exposure of ''M. tuberculosis'' to reactive oxygen and/or reactive nitrogen species results in the formation of DSBs that are repaired by HR or NHEJ. However deficiency of DSB repair does not appear to impair ''M. tuberculosis'' virulence in animal models.
History
''M. tuberculosis'', then known as the "tubercle
In anatomy, a tubercle (literally 'small tuber', Latin for 'lump') is any round nodule, small eminence, or warty outgrowth found on external or internal organs of a plant or an animal.
In plants
A tubercle is generally a wart-like projection ...
bacillus
''Bacillus'' (Latin "stick") is a genus of Gram-positive, rod-shaped bacteria, a member of the phylum ''Bacillota'', with 266 named species. The term is also used to describe the shape (rod) of other so-shaped bacteria; and the plural ''Bacilli ...
", was first described on 24 March 1882 by Robert Koch
Heinrich Hermann Robert Koch ( , ; 11 December 1843 – 27 May 1910) was a German physician and microbiologist. As the discoverer of the specific causative agents of deadly infectious diseases including tuberculosis, cholera (though the Vibrio ...
, who subsequently received the Nobel Prize in Physiology or Medicine
The Nobel Prize in Physiology or Medicine is awarded yearly by the Nobel Assembly at the Karolinska Institute for outstanding discoveries in physiology or medicine. The Nobel Prize is not a single prize, but five separate prizes that, accord ...
for this discovery in 1905; the bacterium is also known as "Koch's bacillus".
''M. tuberculosis'' has existed throughout history, but the name has changed frequently over time. In 1720, though, the history of tuberculosis started to take shape into what is known of it today; as the physician Benjamin Marten
Benjamin Marten (c.1690–1752) was an English physician from "Theobald's Row" near Red Lyon Square, Holborn, and one of several sons of a tailor.
In 1720 he conjectured in ''"A New Theory of Consumptions - More Especially a Phthisis or Consump ...
described in his ''A Theory of Consumption'', tuberculosis may be caused by small living creatures transmitted through the air to other patients.
Vaccine
TheBCG vaccine
Bacillus Calmette–Guérin (BCG) vaccine is a vaccine primarily used against tuberculosis (TB). It is named after its inventors Albert Calmette and Camille Guérin. In countries where tuberculosis or leprosy is common, one dose is recommended ...
(bacille Calmette-Guerin), which was derived from ''M. bovis,'' while effective against childhood and severe forms of tuberculosis, has limited success in preventing the most common form of the disease today, adult pulmonary tuberculosis. Because of this, it is primarily used in high tuberculosis incidence regions, and is not a recommended vaccine in the United States due to the low risk of infection. To receive this vaccine in the United States, an individual is required to go through a consultation process with an expert in M. tuberculosis and is only given to those who meet the specific criteria.
The BCG, according to an article of the Kyodo News (April 14, 2020) titled "Tuberculosis vaccine drawing attention in fight against coronavirus" indicates a possible correlation between BCG vaccination, and better immune response to the COVID-19.
See also
*Philip D'Arcy Hart
Philip Montagu D'Arcy Hart, CBE (25 June 1900 – 30 July 2006) was a seminal British medical researcher and pioneer in tuberculosis treatment.
Personal life
Philip D'Arcy Hart was the grandson of Samuel Montagu, 1st Baron Swaythling. He was ...
References
External links
TB database: an integrated platform for Tuberculosis research
*
Database on Mycobacterium tuberculosis genetics
{{Authority control Acid-fast bacilli
tuberculosis
Tuberculosis (TB) is an infectious disease usually caused by '' Mycobacterium tuberculosis'' (MTB) bacteria. Tuberculosis generally affects the lungs, but it can also affect other parts of the body. Most infections show no symptoms, in ...
Tuberculosis
Pathogenic bacteria
Bacteria described in 1882