Arenavirus on:
[Wikipedia]
[Google]
[Amazon]
An arenavirus is a bisegmented ambisense RNA virus that is a member of the family ''Arenaviridae''. These viruses infect rodents and occasionally humans. A class of novel, highly divergent arenaviruses, properly known as reptarenaviruses, have also been discovered which infect snakes to produce inclusion body disease. At least eight arenaviruses are known to cause human disease. The diseases derived from arenaviruses range in severity. Aseptic meningitis, a severe human disease that causes inflammation covering the brain and spinal cord, can arise from the
 Viewed in cross-section, arenaviruses contain grainy particles that are
Viewed in cross-section, arenaviruses contain grainy particles that are
 Arenaviruses have a segmented
Arenaviruses have a segmented
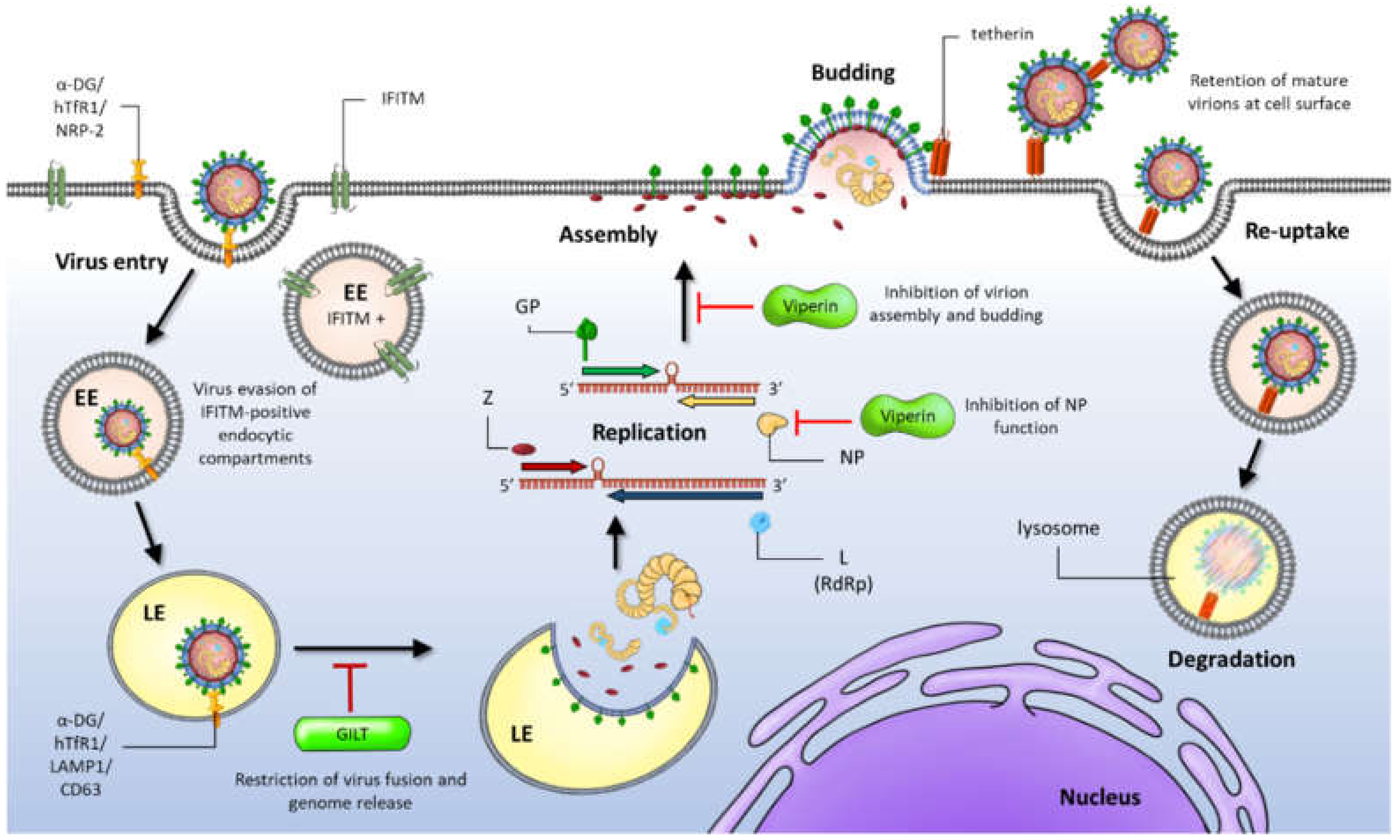 The glycoprotein (GP) is synthesised as a precursor molecule. It is cleaved into three parts - GP1, GP2 and a stable signal peptide (SSP). These reactions are catalysed by cellular signal peptidases and the cellular enzyme Subtilisin Kexin Isozyme-1 (SKI-1)/Site-1 Protease (S1P). These processes are essential for fusion competence and incorporation of mature GP into nascent budding virion particles.
The glycoprotein (GP) is synthesised as a precursor molecule. It is cleaved into three parts - GP1, GP2 and a stable signal peptide (SSP). These reactions are catalysed by cellular signal peptidases and the cellular enzyme Subtilisin Kexin Isozyme-1 (SKI-1)/Site-1 Protease (S1P). These processes are essential for fusion competence and incorporation of mature GP into nascent budding virion particles.
 * Alxa virus (ALXV)
* Dandenong virus (DANV)
* Gairo virus (GAIV)
* Gbagroube virus
* Ippy virus (IPPYV)
* Kodoko virus (KODV)
*
* Alxa virus (ALXV)
* Dandenong virus (DANV)
* Gairo virus (GAIV)
* Gbagroube virus
* Ippy virus (IPPYV)
* Kodoko virus (KODV)
*
 # Lymphocytic choriomeningitis (LCM) viruses cause ''influenza-like febrile illness'', but occasionally they may cause ''meningitis,'' characteristically accompanied by large numbers of lymphocytes in the cerebrospinal fluid (as the name LCM suggests).
# Lassa virus causes
# Lymphocytic choriomeningitis (LCM) viruses cause ''influenza-like febrile illness'', but occasionally they may cause ''meningitis,'' characteristically accompanied by large numbers of lymphocytes in the cerebrospinal fluid (as the name LCM suggests).
# Lassa virus causes
ICTV Report: ''Arenaviridae''
Virus Pathogen Database and Analysis Resource (ViPR): Arenaviridae
Detailed genomic and bioinformatic information about Arenaviridae at NIH-funded database
Arenaviridae Genomes
database search results from th
Viral Bioinformatics Resource Center
info on recent outbreak.
* {{Taxonbar, from=Q2368907 Arthropod-borne viral fevers and viral haemorrhagic fevers de:Arenaviridae
lymphocytic choriomeningitis virus
Lymphocytic choriomeningitis (LCM) is a rodent-borne viral infectious disease that presents as aseptic meningitis, encephalitis or meningoencephalitis. Its causative agent is ''lymphocytic choriomeningitis mammarenavirus'' (LCMV), a member of the ...
. Hemorrhagic fever
Viral hemorrhagic fevers (VHFs) are a diverse group of animal and human illnesses in which fever and hemorrhage are caused by a viral infection. VHFs may be caused by five distinct families of RNA viruses: the families ''Filoviridae'', ''Flav ...
syndromes, including Lassa fever
Lassa fever, also known as Lassa hemorrhagic fever (LHF), is a type of viral hemorrhagic fever caused by the Lassa virus. Many of those infected by the virus do not develop symptoms. When symptoms occur they typically include fever, weakness, ...
, are derived from infections such as Guanarito virus
Venezuelan hemorrhagic fever (VHF) is a zoonotic human illness first identified in 1989. The disease is most prevalent in several rural areas of central Venezuela and is caused by ''Guanarito mammarenavirus'' (GTOV) which belongs to the Arenaviri ...
, Junin virus
''Argentinian mammarenavirus'', better known as the ''Junin virus'' or ''Junín virus'' (JUNV), is an arenavirus in the '' Mammarenavirus'' genus that causes Argentine hemorrhagic fever (AHF). The virus took its original name from the city of Ju ...
, Lassa virus
''Lassa mammarenavirus'' (LASV) is an arenavirus that causes Lassa hemorrhagic fever,
a type of viral hemorrhagic fever (VHF), in humans and other primates. ''Lassa mammarenavirus'' is an emerging virus and a select agent, requiring Biosafet ...
, Lujo virus
Lujo is a bisegmented RNA virus—a member of the family '' Arenaviridae''—and a known cause of viral hemorrhagic fever (VHF) in humans. Its name was suggested by the Special Pathogens Unit of the National Institute for Communicable Diseases of ...
, Machupo virus
Bolivian hemorrhagic fever (BHF), also known as black typhus or Ordog Fever, is a hemorrhagic fever and zoonotic infectious disease originating in Bolivia after infection by ''Machupo mammarenavirus''.Public Health Agency of Canada: ''Machupo V ...
, Sabia virus
Brazilian hemorrhagic fever (BzHF) is an infectious disease caused by ''Brazilian mammarenavirus'', an arenavirus. ''Brazilian mammarenavirus'' is one of the arenaviruses from South America to cause hemorrhagic fever. It shares a common progenit ...
, or Whitewater Arroyo virus
''Whitewater Arroyo mammarenavirus'' (WWAV) is a zoonotic Arenavirus associated with hemorrhagic fever with liver failure.
Discovery
WWAV is an emerging virus; previously, it was widely distributed among woodrats (''Neotoma'' spp.), its rese ...
. Because of the epidemiological association with rodents, some arenaviruses and bunyavirus
''Bunyavirales'' is an order of segmented negative-strand RNA viruses with mainly tripartite genomes. Member viruses infect arthropods, plants, protozoans, and vertebrates. It is the only order in the class ''Ellioviricetes''. The name ''Bunyavi ...
es are designated as roboviruses.
Structure
 Viewed in cross-section, arenaviruses contain grainy particles that are
Viewed in cross-section, arenaviruses contain grainy particles that are ribosome
Ribosomes ( ) are macromolecular machines, found within all cells, that perform biological protein synthesis (mRNA translation). Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to ...
s acquired from their host cells. It is from this characteristic that they acquired the name ''arena
An arena is a large enclosed platform, often circular or oval-shaped, designed to showcase theatre, musical performances, or sporting events. It is composed of a large open space surrounded on most or all sides by tiered seating for spectators ...
'', from the Latin root meaning ''sand
Sand is a granular material composed of finely divided mineral particles. Sand has various compositions but is defined by its grain size. Sand grains are smaller than gravel and coarser than silt. Sand can also refer to a textural class of s ...
''. The ribosomal structures are not believed to be essential for virus replication. Virus particles, or virions, are pleomorphic (variable in shape) but are often spherical, with a diameter of 60–300 nm, and are covered with surface glycoprotein spikes.
The virus contains a beaded nucleocapsid
A capsid is the protein shell of a virus, enclosing its genetic material. It consists of several oligomeric (repeating) structural subunits made of protein called protomers. The observable 3-dimensional morphological subunits, which may or may ...
with two single-stranded RNA
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. RNA and deoxyribonucleic acid ( DNA) are nucleic acids. Along with lipids, proteins, and carbohydra ...
segments. The nucleocapsid consists of a core of nucleic acid enclosed in a protein coat. Although they are categorized as negative-sense
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complementarity (molecular biology), complement in specifying a sequence of am ...
viruses, arenaviruses are ambisense
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context ...
. While sections of their genome encode genes in the negative sense (reverse polarity), other sections encode genes in the opposite (forward/positive sense) direction. This complex gene expression structure is theorized to be a primitive regulatory system, allowing the virus to control what proteins are synthesized at what point in the life cycle. The life cycle of the arenavirus is restricted to the cell cytoplasm.
Genome
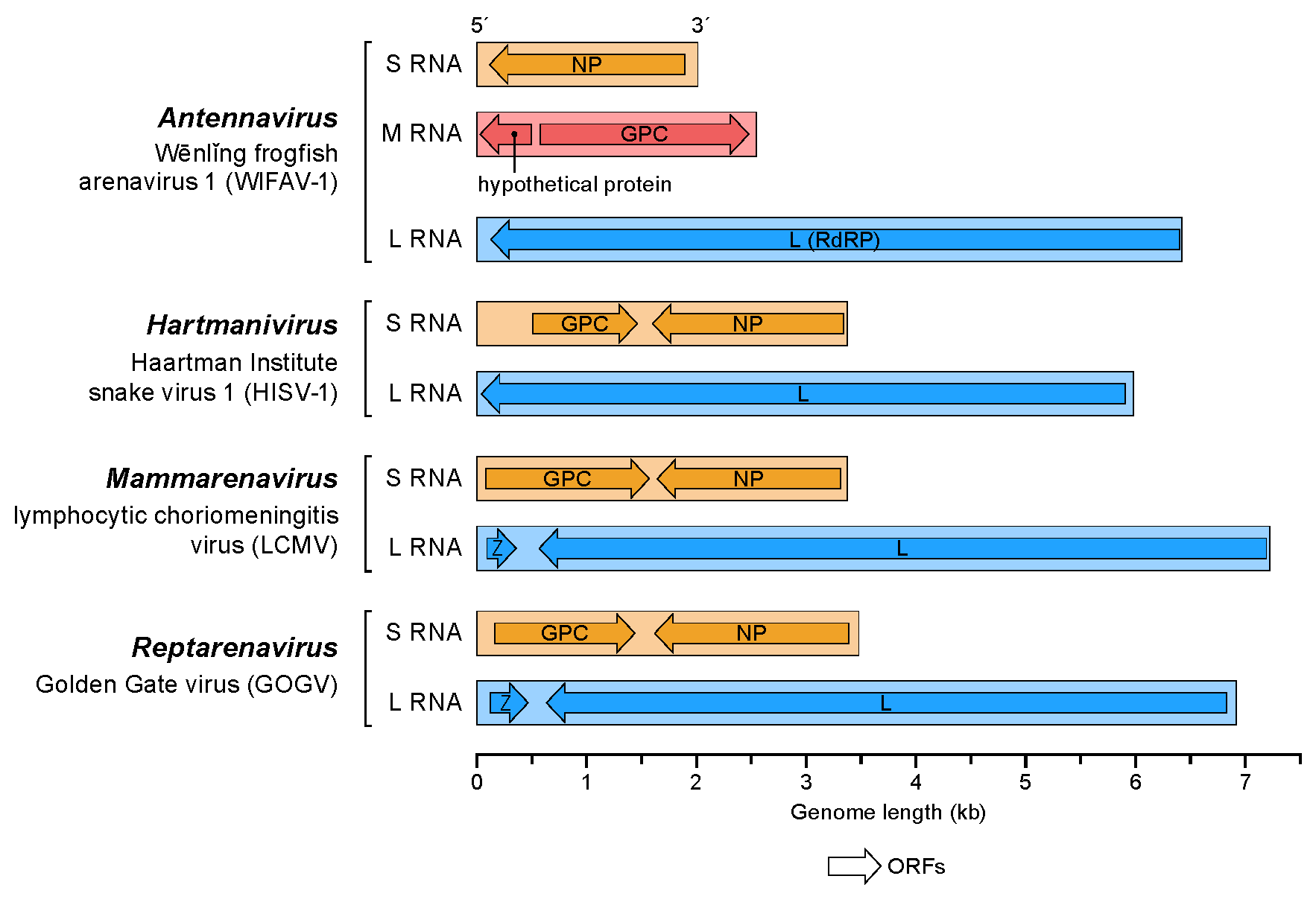
 Arenaviruses have a segmented
Arenaviruses have a segmented RNA
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. RNA and deoxyribonucleic acid ( DNA) are nucleic acids. Along with lipids, proteins, and carbohydra ...
genome that consists of two single-stranded ambisense RNAs. As with all negative-sense RNA viruses, the genomic RNA alone is not infectious and the viral replication machinery is required to initiate infection within a host cell. Genomic sense RNA packaged into the arenavirus virion is designated negative-sense RNA, and must first be copied into a positive-sense mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
in order to produce viral protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respo ...
. The two RNA segments are denoted Small (S) and Large (L), and code for four viral proteins in a unique ambisense coding strategy. Each RNA segment codes for two viral proteins in opposite orientation such that the negative-sense RNA genome serves as the template for transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of a single mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
and the positive-sense copy of the RNA genome templates a second mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
. The separate coding sequences of the two viral proteins are divided by an intergenic region RNA sequence that is predicted to fold into a stable hairpin structure.
The extreme termini of each RNA segment contains a 19 nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules wi ...
highly conserved sequence that is critical for recruitment of the viral replication machinery and initiation of viral mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
and genomic replication. The conserved 5' and 3' RNA termini sequences are complementary and allows each RNA segment to adopt a double-stranded RNA panhandle structure that maintains the termini in close proximity and results in a circular appearance to purified arenavirus genomic templates visualized by electron microscopy
An electron microscope is a microscope that uses a beam of accelerated electrons as a source of illumination. As the wavelength of an electron can be up to 100,000 times shorter than that of visible light photons, electron microscopes have a hi ...
. The double-stranded RNA panhandle structure is critical for efficient viral RNA synthesis, but potential interterminal double-stranded RNA interactions must be transiently relieved in order to recruit the viral polymerase
A polymerase is an enzyme ( EC 2.7.7.6/7/19/48/49) that synthesizes long chains of polymers or nucleic acids. DNA polymerase and RNA polymerase are used to assemble DNA and RNA molecules, respectively, by copying a DNA template strand using base- ...
.
The S-segment RNA is approximately 3.5 kb, and encodes the viral nucleocapsid protein (NP) and glycoprotein (GPC). The L-segment RNA is approximately 7.2 kb, and encodes the viral RNA-dependent RNA-polymerase (L) and a small RING-domain containing protein (Z).
The Z protein forms homo oligomers and a structural component of the virions. The formation of these oligomers is an essential step for particle assembly and budding. Binding between Z and the viral envelope glycoprotein complex is required for virion infectivity. Z also interacts with the L and NP proteins. Polymerase activity appears to be modulated by the association between the L and Z proteins. Interaction between the Z and NP proteins is critical for genome packaging.
Microbiology
 The glycoprotein (GP) is synthesised as a precursor molecule. It is cleaved into three parts - GP1, GP2 and a stable signal peptide (SSP). These reactions are catalysed by cellular signal peptidases and the cellular enzyme Subtilisin Kexin Isozyme-1 (SKI-1)/Site-1 Protease (S1P). These processes are essential for fusion competence and incorporation of mature GP into nascent budding virion particles.
The glycoprotein (GP) is synthesised as a precursor molecule. It is cleaved into three parts - GP1, GP2 and a stable signal peptide (SSP). These reactions are catalysed by cellular signal peptidases and the cellular enzyme Subtilisin Kexin Isozyme-1 (SKI-1)/Site-1 Protease (S1P). These processes are essential for fusion competence and incorporation of mature GP into nascent budding virion particles.
Taxonomy
Within the family ''Arenaviridae'', arenaviruses were formerly all placed in the genus ''Arenavirus'', but in 2014 were divided into the genera ''Mammarenavirus
''Mammarenavirus'' is a genus of viruses in the family ''Arenaviridae''. The name is a portmanteau of mammal and the former name ''Arenavirus'', and differentiates it from the reptile-associated ''Reptarenavirus''. ''Arenavirus'' comes from the ...
'' for those with mammalian hosts and '' Reptarenavirus'' for those infecting snakes. Reptarenaviruses and mammarenavirus are separated by an impenetrable species barrier. Infected rodents cannot pass disease onto snakes, and IBD in captive snakes is not transmissible to humans.
A third genus, '' Hartmanivirus'' (not to be confused with genus ''Haartmanvirus'' of vibrio
''Vibrio'' is a genus of Gram-negative bacteria, possessing a curved-rod (comma) shape, several species of which can cause foodborne infection, usually associated with eating undercooked seafood. Being highly salt tolerant and unable to survive ...
phages
A bacteriophage (), also known informally as a ''phage'' (), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν ('), meaning "to devour". Bacterio ...
in family '' Demerecviridae'', order ''Caudovirales
''Caudovirales'' is an order of viruses known as the tailed bacteriophages (''cauda'' is Latin for "tail"). Under the Baltimore classification scheme, the ''Caudovirales'' are group I viruses as they have double stranded DNA (dsDNA) genomes ...
''), has also been established, including other species that infect snakes. The organisation of the genome of this genus is typical of arenaviruses but their glycoproteins resemble those of filovirus
''Filoviridae'' () is a family of single-stranded negative-sense RNA viruses in the order ''Mononegavirales''. Two members of the family that are commonly known are Ebola virus and Marburg virus. Both viruses, and some of their lesser known re ...
es. Species in this genus lack the matrix protein.
A fourth genus, '' Antennavirus'' has also been established to accommodate two arenaviruses found in striated frogfish ('' Antennarius striatus'').Zhang YZ, Wu WC, Shi M, Holmes EC (2018) The diversity, evolution and origins of vertebrate RNA viruses. Curr Opin Virol 31:9-16
Mammarenaviruses can be divided into two serogroups, which differ genetically and by geographical distribution:
When the virus is classified "Old World" this means it was found in the Eastern Hemisphere in places such as Europe, Asia, and Africa. When it is found in the Western Hemisphere, in places such as Argentina, Bolivia, Venezuela, Brazil, and the United States, it is classified "New World". Lymphocytic choriomeningitis (LCM) virus is the only arenavirus to exist in both areas but is classified as an Old World virus. Old and New World area viruses appear to have diverged ~45,000 years ago.
The Old World Mammarenaviruses originated ~23.1-1.88 thousand years ago, most likely in Southern Africa while the New World Mammarenaviruses evolved in the Latin America-Caribbean region ~41.4-3.3 thousand years ago.
''Mammarenavirus''
Old World complex
 * Alxa virus (ALXV)
* Dandenong virus (DANV)
* Gairo virus (GAIV)
* Gbagroube virus
* Ippy virus (IPPYV)
* Kodoko virus (KODV)
*
* Alxa virus (ALXV)
* Dandenong virus (DANV)
* Gairo virus (GAIV)
* Gbagroube virus
* Ippy virus (IPPYV)
* Kodoko virus (KODV)
* Lassa virus
''Lassa mammarenavirus'' (LASV) is an arenavirus that causes Lassa hemorrhagic fever,
a type of viral hemorrhagic fever (VHF), in humans and other primates. ''Lassa mammarenavirus'' is an emerging virus and a select agent, requiring Biosafet ...
(LASV)
* Lìjiāng virus (LIJV)
* Loei River virus (LORV)
* Lujo virus
Lujo is a bisegmented RNA virus—a member of the family '' Arenaviridae''—and a known cause of viral hemorrhagic fever (VHF) in humans. Its name was suggested by the Special Pathogens Unit of the National Institute for Communicable Diseases of ...
(LUJV)
* Luna virus (LUAV)
* Lunk virus (LNKV)
* Lymphocytic choriomeningitis virus
Lymphocytic choriomeningitis (LCM) is a rodent-borne viral infectious disease that presents as aseptic meningitis, encephalitis or meningoencephalitis. Its causative agent is ''lymphocytic choriomeningitis mammarenavirus'' (LCMV), a member of the ...
(LCMV)
* Mariental virus (MRTV)
* Merino Walk virus (MRWV)
* Menekre virus
* Minu virus
* Mobala virus (MOBV)
* Morogoro virus (MORV)
* Mopeia virus (MOPV)
* Ryukyu virus (RYKV)
* Solwezi virus (SOLV)
* Souris virus (SOUV)
* Okahandja virus (OKAV)
* Wenzhou virus (WENV)
New World complex
* Clade A ** Allpahuayo virus (ALLV) ** Flexal virus (FLEV) ** Paraná virus (PRAV) ** Pichindé virus (PICHV) ** Pirital virus (PIRV) * Clade B ** Amaparí virus (AMAV) ** Aporé virus (APOV) **Chapare virus
''Chapare mammarenavirus'' or Chapare virus is a virus from the family '' Arenaviridae'' which causes a hemorrhagic fever in humans known as Chapare hemorrhagic fever. It was first described after an outbreak of a novel zoonotic mammarenavirus i ...
(CHAPV)
** Cupixi virus (CUPXV)
** Guanarito virus
Venezuelan hemorrhagic fever (VHF) is a zoonotic human illness first identified in 1989. The disease is most prevalent in several rural areas of central Venezuela and is caused by ''Guanarito mammarenavirus'' (GTOV) which belongs to the Arenaviri ...
(GTOV)
** Junín virus (JUNV)
** Machupo virus
Bolivian hemorrhagic fever (BHF), also known as black typhus or Ordog Fever, is a hemorrhagic fever and zoonotic infectious disease originating in Bolivia after infection by ''Machupo mammarenavirus''.Public Health Agency of Canada: ''Machupo V ...
(MACV)
** Ocozocoautla de Espinosa virus
** Real de Catorce virus (RCTV)
** Tacaribe virus (TCRV)
** Xapuri virus (XAPV)
** Sabiá virus (SBAV)
* Clade C
** Latino virus (LATV)
** Oliveros virus (OLVV)
* Clade D
** Bear Canyon virus (BCNV)
** Catarina virus (CTNV)
** Skinner Tank virus (SKTV)
** Tamiami virus (TMMV)
** Whitewater Arroyo virus
''Whitewater Arroyo mammarenavirus'' (WWAV) is a zoonotic Arenavirus associated with hemorrhagic fever with liver failure.
Discovery
WWAV is an emerging virus; previously, it was widely distributed among woodrats (''Neotoma'' spp.), its rese ...
(WWAV)
** Catarina virus (CTNV)
** Big Brushy Tank virus (BBTV)
** Skinner Tank virus
Skinner may refer to:
People and fictional characters
*Skinner (surname), a list of people and fictional characters with that surname
*Skinner (profession), a person who makes a living by working with animal skins or driving mules
*Skinner, a ring ...
(SKTV)
** Tonto Creek virus (TTCV)
* Others
** Patawa virus
''Reptarenavirus''
* California reptarenavirus * Golden reptarenavirus * Rotterdam reptarenavirus * Ordinary reptarenavirus * Giessen reptarenavirus''Hartmanivirus''
* Muikkunen hartmanivirus * Haartman hartmanivirus * Schoolhouse hartmanivirus * Zurich hartmanivirus * Heimat hartmanivirus * Setpatvet hartmanivirusAntennavirus
* Hairy antennavirus * Striated antennavirus * Salmon antennavirusEvolution
The evolution of the Mammarenavirus genus has been studied. The New World and Old World species diverged less than 45,000 years ago. The New World species evolved between 41,400 and 3,300 years ago in the Latin America-Caribbean region. The Old World species evolved between 23,100 and 1,880 years ago, most likely in southern Africa.Reservoirs
Some arenaviruses arezoonotic
A zoonosis (; plural zoonoses) or zoonotic disease is an infectious disease of humans caused by a pathogen (an infectious agent, such as a bacterium, virus, parasite or prion) that has jumped from a non-human (usually a vertebrate) to a human. ...
pathogens and are generally associated with rodent
Rodents (from Latin , 'to gnaw') are mammals of the order Rodentia (), which are characterized by a single pair of continuously growing incisors in each of the upper and lower jaws. About 40% of all mammal species are rodents. They are na ...
—transmitted disease in humans. Each virus usually is associated with a particular rodent host species in which it is maintained. Arenaviruses persist in nature by infecting rodents first and then transmitted into humans. Humans can be infected through mucosal exposure to aerosols, or by direct contact of abraded skin with the infectious material, derived from infected rodents. Aerosols are fine mists or sprays of rodent dried excreta, especially urine that is dropped in the environment. Most of the Arenaviruses caught by humans are within their own homes when these rodents seek shelter. The virus can be caught in factories, from food that has been contaminated, or within agricultural work areas. Humans' risk of contracting the Arenavirus infection is related to age, race, or sex within the degree of contact with the dried rodent excreta.
Epidemiology
Hosts
Clinical diseases
 # Lymphocytic choriomeningitis (LCM) viruses cause ''influenza-like febrile illness'', but occasionally they may cause ''meningitis,'' characteristically accompanied by large numbers of lymphocytes in the cerebrospinal fluid (as the name LCM suggests).
# Lassa virus causes
# Lymphocytic choriomeningitis (LCM) viruses cause ''influenza-like febrile illness'', but occasionally they may cause ''meningitis,'' characteristically accompanied by large numbers of lymphocytes in the cerebrospinal fluid (as the name LCM suggests).
# Lassa virus causes Lassa fever
Lassa fever, also known as Lassa hemorrhagic fever (LHF), is a type of viral hemorrhagic fever caused by the Lassa virus. Many of those infected by the virus do not develop symptoms. When symptoms occur they typically include fever, weakness, h ...
. Lassa fever is endemic in west Africa. The virus was first isolated from Americans stationed in the village of Lassa, Nigeria. The virus can be transmitted person-to-person.
#* Subclinical diseases: Serological studies suggest that inapparent infections particularly among members of hunting tribes are common.
#* Clinical infections: Lassa fever is characterised by high fever, severe myalgia, coagulopathy, haemorrhagic skin rash, and occasional visceral haemorrhage as well as necrosis of liver and spleen.
# Other Arenaviruses like Junin virus, Machupo virus cause haemorrhagic fevers.
All of these diseases pose a great threat to public health in the regions where it is taking place. For example, when the Old World Lassa virus turns into Lassa fever, this usually results in a significant amount of mortality. Similarly the New World Junin virus causes Argentine hemorrhagic fever. This fever is a severe illness with hemorrhagic and neurological manifestations and a case fatality of fifteen to thirty percent. The way this virus spreads is through increased traveling to and from endemic regions. This traveling has led to the importation of Lassa fever into non-endemic metropolitan areas all over the world.
Recent outbreaks
A new species of arenavirus named theLujo virus
Lujo is a bisegmented RNA virus—a member of the family '' Arenaviridae''—and a known cause of viral hemorrhagic fever (VHF) in humans. Its name was suggested by the Special Pathogens Unit of the National Institute for Communicable Diseases of ...
has been linked to five patients who exhibited symptoms of viral hemorrhagic fever in South Africa. The disease originated near Lusaka, Zambia
Zambia (), officially the Republic of Zambia, is a landlocked country at the crossroads of Central Africa, Central, Southern Africa, Southern and East Africa, although it is typically referred to as being in Southern Africa at its most cent ...
and spread to Johannesburg
Johannesburg ( , , ; Zulu and xh, eGoli ), colloquially known as Jozi, Joburg, or "The City of Gold", is the largest city in South Africa, classified as a megacity, and is one of the 100 largest urban areas in the world. According to Demo ...
, South Africa
South Africa, officially the Republic of South Africa (RSA), is the southernmost country in Africa. It is bounded to the south by of coastline that stretch along the South Atlantic and Indian Oceans; to the north by the neighbouring countri ...
, after the first patient was transported to a hospital there. The results of genetic sequencing tests conducted by epidemiologists at Columbia University
Columbia University (also known as Columbia, and officially as Columbia University in the City of New York) is a private research university in New York City. Established in 1754 as King's College on the grounds of Trinity Church in Manhatt ...
in New York City
New York, often called New York City or NYC, is the List of United States cities by population, most populous city in the United States. With a 2020 population of 8,804,190 distributed over , New York City is also the L ...
, USA, and at the Special Pathogens Branch of the Centers for Disease Control
The Centers for Disease Control and Prevention (CDC) is the national public health agency of the United States. It is a United States federal agency, under the Department of Health and Human Services, and is headquartered in Atlanta, Georgi ...
in Atlanta
Atlanta ( ) is the capital and most populous city of the U.S. state of Georgia. It is the seat of Fulton County, the most populous county in Georgia, but its territory falls in both Fulton and DeKalb counties. With a population of 498,715 ...
, USA, provided evidence that the causative agent of the disease is a virus from the family Arenaviridae, which ultimately resulted in the deaths of four out of the five infected in Zambia and South Africa during the outbreak which began in September 2008.
Arenavirus has also pinpointed as the cause of death of three donor organ recipients in Australia who contracted the virus after receiving kidney and a liver donations from a single infected organ donor in late 2006. All three died in the first week of 2007.
WHO
Who or WHO may refer to:
* Who (pronoun), an interrogative or relative pronoun
* Who?, one of the Five Ws in journalism
* World Health Organization
Arts and entertainment Fictional characters
* Who, a creature in the Dr. Seuss book '' Horton He ...
and its Global Outbreak Alert and Response Network (GOARN) partners continue to support the Ministries of Health of the two countries in various facets of the outbreak investigation, including laboratory diagnosis, investigations, active case finding and follow-up of contacts.
Treatments
Very few treatment methods are available. The current lack of a licensed vaccine and limited therapeutic options for the arenavirus make it arguably among the most neglected virus groups. The only licensed drug for the treatment of human arenavirus infection is the nucleoside analogueribavirin
Ribavirin, also known as tribavirin, is an antiviral medication used to treat RSV infection, hepatitis C and some viral hemorrhagic fevers. For hepatitis C, it is used in combination with other medications such as simeprevir, sofosbuvir, pegin ...
. Ribavirin reduces morbidity and mortality in humans infected with certain arenaviruses, such as LASV and JUNV infections, if it is taken in the early stages of the disease. Ribavirin displays mixed success in treating severe arenaviral disease and is associated with significant toxicities.
Experimental approaches
Effective antiviral drugs need to be produced at a low cost, taken orally, and able to withstand tropical climates due to the regions where these infections are occurring. For this reason high throughput screening (HTS) of small molecular libraries could be the answer to finding a better remedy. HTS collects libraries of small synthetic molecules that can be used to identify protein promoting "agonist" molecules or protein inhibiting "antagonist" interactions. With HTS sustainable antiviral drugs can be discovered against possible new human pathogenic viruses.Immunotherapy
Immunotherapy or biological therapy is the treatment of disease by activating or suppressing the immune system. Immunotherapies designed to elicit or amplify an immune response are classified as ''activation immunotherapies,'' while immunotherap ...
is another potential approach. Monoclonal antibodies
A monoclonal antibody (mAb, more rarely called moAb) is an antibody produced from a cell Lineage made by cloning a unique white blood cell. All subsequent antibodies derived this way trace back to a unique parent cell.
Monoclonal antibodies ca ...
against Junin virus
''Argentinian mammarenavirus'', better known as the ''Junin virus'' or ''Junín virus'' (JUNV), is an arenavirus in the '' Mammarenavirus'' genus that causes Argentine hemorrhagic fever (AHF). The virus took its original name from the city of Ju ...
have been tested in animal models. An immunotherapeutic agent active against all tested mammarenaviruses that use the transferrin receptor 1
Transferrin receptor protein 1 (TfR1), also known as Cluster of Differentiation 71 (CD71), is a protein that in humans is encoded by the ''TFRC'' gene. TfR1 is required for iron import from transferrin into cells by endocytosis.
Structure and func ...
as their receptor
Receptor may refer to:
* Sensory receptor, in physiology, any structure which, on receiving environmental stimuli, produces an informative nerve impulse
*Receptor (biochemistry), in biochemistry, a protein molecule that receives and responds to a ...
was under investigation in 2020.
References
External links
ICTV Report: ''Arenaviridae''
Virus Pathogen Database and Analysis Resource (ViPR): Arenaviridae
Detailed genomic and bioinformatic information about Arenaviridae at NIH-funded database
Arenaviridae Genomes
database search results from th
Viral Bioinformatics Resource Center
info on recent outbreak.
* {{Taxonbar, from=Q2368907 Arthropod-borne viral fevers and viral haemorrhagic fevers de:Arenaviridae