|
HMGA
HMGA is a family of high mobility group proteins characterized by an AT-hook. They code for a "small, nonhistone, chromatin-associated protein that has no intrinsic transcriptional activity but can modulate transcription by altering the chromatin architecture". Mammals have two orthologs: HMGA1 and HMGA2. Genomic distribution In mouse embryonic stem cells it has been demonstrated that both HMGA proteins binds uniformly to the DNA due to their AT-hook domains, with a slight preference for AT-rich regions/ Such regions tend to lack coding genes, an observation that argues against a direct role in transcriptional control and in agreement with previous studies, suggest that these proteins have a structural role in the chromatin, similar to histone. Association with human traits Variations in HMGA2 to have a moderate association with adult height. See also * HMGA1 High-mobility group protein HMG-I/HMG-Y is a protein that in humans is encoded by the ''HMGA1'' gene. Function ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
HMGA2
High-mobility group AT-hook 2, also known as HMGA2, is a protein that, in humans, is encoded by the ''HMGA2'' gene. Function This gene encodes a protein that belongs to the non-histone chromosomal high-mobility group (HMG) protein family. HMG proteins function as architectural factors and are essential components of the enhanceosome. This protein contains structural DNA-binding domains and may act as a transcriptional regulating factor. Identification of the deletion, amplification, and rearrangement of this gene that are associated with lipomas suggests a role in adipogenesis and mesenchymal differentiation. A gene knock-out study of the mouse counterpart demonstrated that this gene is involved in diet-induced obesity. Alternate transcriptional splice variants, encoding different isoforms, have been characterized. The expression of HMGA2 in adult tissues is commonly associated with both malignant and benign tumor formation, as well as certain characteristic cancer-promoting mu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
HMGA1
High-mobility group protein HMG-I/HMG-Y is a protein that in humans is encoded by the ''HMGA1'' gene. Function This gene encodes a non-histone chromatin protein involved in many cellular processes, including regulation of inducible gene transcription, DNA replication, heterochromatin organization, integration of retroviruses into chromosomes, and the metastatic progression of cancer cells. HMGA1 proteins are quite small (~10-12 kDa) and basic molecules, and consist of three AT-hooks with the RGRP (Arg-Gly-Arg-Pro) core motif, a novel cross-linking domain located between the second and third AT-hook, and a C-terminal acidic tail characteristic for the HMG family comprising HMGA, HMGB and HMGN proteins. HMGA1-GFP fusion proteins are highly dynamic in vivo (determined using FRAP analysis), but in contrast also show nanomolar affinity to AT-rich DNA in vitro (determined biochemically), which might be explained due to the extensive post-transcriptional modifications in vivo. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
High Mobility Group
High-Mobility Group or HMG is a group of chromosomal proteins that are involved in the regulation of DNA-dependent processes such as transcription, replication, recombination, and DNA repair. Families The HMG proteins are subdivided into 3 superfamilies each containing a characteristic functional domain: * HMGA – contains an AT-hook domain ** HMGA1 ** HMGA2 * HMGB – contains a HMG-box domain ** HMGB1 ** HMGB2 ** HMGB3 ** HMGB4 * HMGN – contains a nucleosomal binding domain ** HMGN1 ** HMGN2 ** HMGN3 ** HMGN4 ** HMGN5 Proteins containing any of these embedded in their sequence are known as HMG motif proteins. HMG-box proteins are found in a variety of eukaryotic organisms. They were originally isolated from mammalian cells, and named according to their electrophoretic mobility in polyacrylamide gels. Other families with HMG-box domain * SOX gene family ** Sex-Determining Region Y Protein ** SOX1, SOX2, etc. * TCF/LEF family (T cell factor/lymphoid enhancer f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
AT-hook
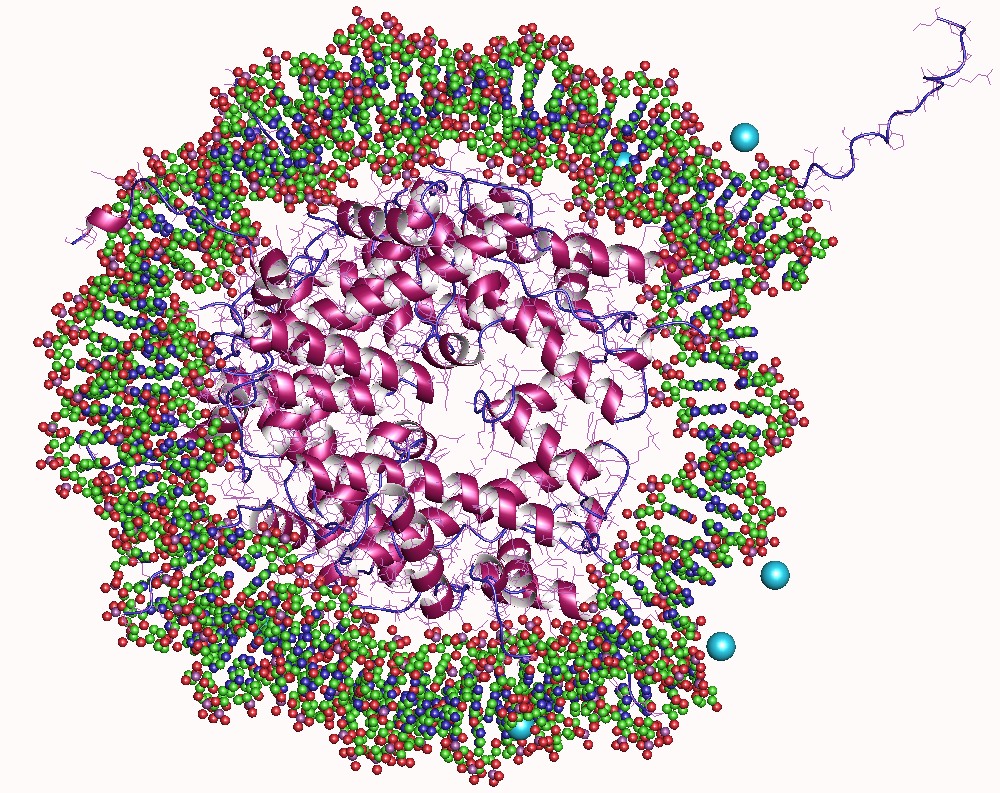
The second AT-hook of HMGA1 (black ribbon) bound to the minor-groove of AT-rich DNA. The amino-acid side chains and nucleotides have been hidden. The AT-hook is a DNA-binding motif present in many proteins, including the high mobility group (HMG) proteins, DNA-binding proteins from plants and hBRG1 protein, a central ATPase of the human switching/sucrose non-fermenting (SWI/SNF) remodeling complex. This motif consists of a conserved, palindromic, core sequence of proline-arginine-glycine-arginine-proline, although some AT-hooks contain only a single proline in the core sequence. AT-hooks also include a variable number of positively charged lysine and arginine residues on either side of the core sequence. The AT-hook binds to the minor groove of adenine-thymine (AT) rich DNA, hence the AT in the name. The rest of the name derives from a predicted asparagine/aspartate "hook" in the earliest AT-hooks reported in 1990. In 1997 structural studies using NMR Nuclear magnetic r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wrapped into 30- nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 90 micrometers (0.09 mm) of 30 nm diameter chromatin fibers. There are five families of histones which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B dimers and a H3-H4 tetramer. The tight wrapping of DNA around h ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |