Transcription Translation Feedback Loop on:
[Wikipedia]
[Google]
[Amazon]
Transcription-translation feedback loop (TTFL) is a cellular model for explaining
 Secondary feedback loops interact with this primary feedback loop. CLOCKWORK ORANGE (CWO) binds the E-boxes to act as a direct competitor of CYC-CLK, therefore inhibiting transcription. PAR-DOMAIN PROTEIN 1 ε (PDP1ε) is a feedback activator and VRILLE (VRI) is a feedback inhibitor of the Clk promoter, and their expression is activated by dCLK-dCYC. Ecdysone-induced protein 75 (E75) inhibits ''clk'' expression and ''tim''-dependently activates ''pe''r transcription. All of these secondary loops act to reinforce the primary TTFL.
Cryptochrome in ''Drosophila'' is a blue-light photoreceptor that triggers degradation of TIM, indirectly leading to the clock phase being reset and the renewed promotion of ''per'' expression.
Secondary feedback loops interact with this primary feedback loop. CLOCKWORK ORANGE (CWO) binds the E-boxes to act as a direct competitor of CYC-CLK, therefore inhibiting transcription. PAR-DOMAIN PROTEIN 1 ε (PDP1ε) is a feedback activator and VRILLE (VRI) is a feedback inhibitor of the Clk promoter, and their expression is activated by dCLK-dCYC. Ecdysone-induced protein 75 (E75) inhibits ''clk'' expression and ''tim''-dependently activates ''pe''r transcription. All of these secondary loops act to reinforce the primary TTFL.
Cryptochrome in ''Drosophila'' is a blue-light photoreceptor that triggers degradation of TIM, indirectly leading to the clock phase being reset and the renewed promotion of ''per'' expression.
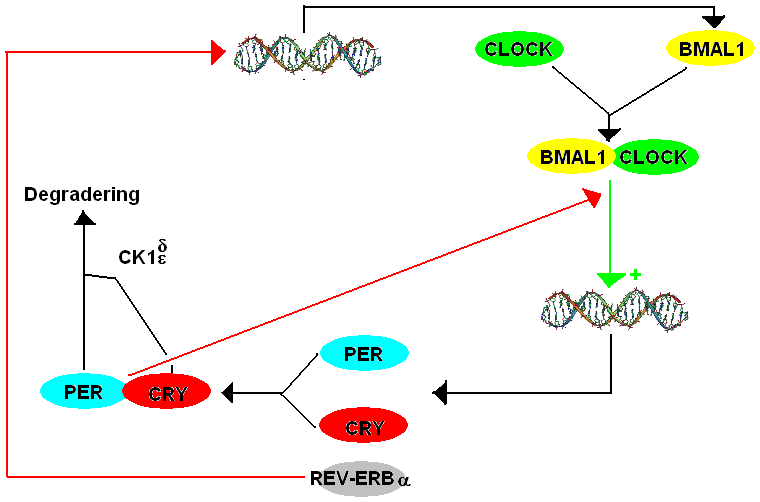 The mammalian TTFL model contains many components that are homologs of the ones found in ''Drosophila''. The way the mammalian system works is that
The mammalian TTFL model contains many components that are homologs of the ones found in ''Drosophila''. The way the mammalian system works is that
 The gene ''frequency'' (''frq'') in ''
The gene ''frequency'' (''frq'') in ''
 A second loop exists involving PRR9, PRR7, and PRR5, which are all homologs of TOC1 and repress CCA1 and LHY expression. These PRR genes are directly repressed by LHY and TOC1. These genes are also regulated by the “evening complex” (EC), which is formed by LUX ARRHYTHMO (LUX), EARLY FLOWERING 3 (ELF3) and EARLY FLOWERING 4 (ELF4). LUX is a transcription factor with a similar function to MYB, while ELF3 and ELF4 are nuclear proteins whose functions are unknown. The "evening complex" indirectly promotes the expression of LHY and CCA1, which repress transcription of its own components. Since this model consists of two inhibitions leading to an activation, it is also referred to as a repressilator.
A recently discovered loop includes the ''reveille'' (''reveille'') family of genes, which are expressed in the morning and induce transcription of evening genes such as PRR5, TOC1, LUX, and ELF4. Once the resulting proteins are translated, PRR9, PRR7, and PRR5 repress RVE8. RVE8 also interacts with the NIGHT LIGHT-INDUCIBLE AND CLOCK-REGULATED (LNK1, 2, 3, and 4) morning components, with LNKs either antagonizing or co-activating RVE8.
Although GIGANTEA (GI) is not known as a core part of the ''Arabdopsis'' TTFL model, it is repressed by CCA1, LHY and TOC1. Additionally, GI activates CCA1 and LHY expression.
A second loop exists involving PRR9, PRR7, and PRR5, which are all homologs of TOC1 and repress CCA1 and LHY expression. These PRR genes are directly repressed by LHY and TOC1. These genes are also regulated by the “evening complex” (EC), which is formed by LUX ARRHYTHMO (LUX), EARLY FLOWERING 3 (ELF3) and EARLY FLOWERING 4 (ELF4). LUX is a transcription factor with a similar function to MYB, while ELF3 and ELF4 are nuclear proteins whose functions are unknown. The "evening complex" indirectly promotes the expression of LHY and CCA1, which repress transcription of its own components. Since this model consists of two inhibitions leading to an activation, it is also referred to as a repressilator.
A recently discovered loop includes the ''reveille'' (''reveille'') family of genes, which are expressed in the morning and induce transcription of evening genes such as PRR5, TOC1, LUX, and ELF4. Once the resulting proteins are translated, PRR9, PRR7, and PRR5 repress RVE8. RVE8 also interacts with the NIGHT LIGHT-INDUCIBLE AND CLOCK-REGULATED (LNK1, 2, 3, and 4) morning components, with LNKs either antagonizing or co-activating RVE8.
Although GIGANTEA (GI) is not known as a core part of the ''Arabdopsis'' TTFL model, it is repressed by CCA1, LHY and TOC1. Additionally, GI activates CCA1 and LHY expression.
circadian rhythm
A circadian rhythm (), or circadian cycle, is a natural oscillation that repeats roughly every 24 hours. Circadian rhythms can refer to any process that originates within an organism (i.e., Endogeny (biology), endogenous) and responds to the env ...
s in behavior and physiology
Physiology (; ) is the science, scientific study of function (biology), functions and mechanism (biology), mechanisms in a life, living system. As a branches of science, subdiscipline of biology, physiology focuses on how organisms, organ syst ...
. Widely conserved across species, the TTFL is auto-regulatory, in which transcription of clock genes is regulated by their own protein products.
Discovery
Circadian rhythm
A circadian rhythm (), or circadian cycle, is a natural oscillation that repeats roughly every 24 hours. Circadian rhythms can refer to any process that originates within an organism (i.e., Endogeny (biology), endogenous) and responds to the env ...
s have been documented for centuries. For example, French astronomer Jean-Jacques d’Ortous de Mairan noted the periodic 24-hour movement of ''Mimosa'' plant leaves as early as 1729. However, science has only recently begun to uncover the cellular mechanisms responsible for driving observed circadian rhythms. The cellular basis of circadian rhythms is supported by the fact that rhythms have been observed in single-celled organisms
A unicellular organism, also known as a single-celled organism, is an organism that consists of a single cell, unlike a multicellular organism that consists of multiple cells. Organisms fall into two general categories: prokaryotic organisms and ...
Beginning in the 1970s, experiments conducted by Ron Konopka and colleagues, in which forward genetic methods were used to induce mutation, revealed that ''Drosophila melanogaster
''Drosophila melanogaster'' is a species of fly (an insect of the Order (biology), order Diptera) in the family Drosophilidae. The species is often referred to as the fruit fly or lesser fruit fly, or less commonly the "vinegar fly", "pomace fly" ...
'' specimens with altered ''period'' (''Per'') genes also demonstrated altered periodicity. As genetic and molecular biology experimental tools improved, researchers further identified genes involved in sustaining normal rhythmic behavior, giving rise to the concept that internal rhythms are modified by a small subset of core clock genes. Hardin and colleagues (1990) were the first to propose that the mechanism driving these rhythms was a negative feedback loop. Subsequent major discoveries confirmed this model; notably experiments led by Thomas K. Darlington and Nicholas Gekakis in the late 1990s that identified clock proteins and characterized their methods in ''Drosophila'' and mice, respectively. These experiments gave rise to the transcription-translation feedback loop (TTFL) model that has now become the dominant paradigm for explaining circadian behavior in a wide array of species.
General mechanisms of TTFL
The TTFL is anegative feedback loop
Negative feedback (or balancing feedback) occurs when some function of the output of a system, process, or mechanism is fed back in a manner that tends to reduce the fluctuations in the output, whether caused by changes in the input or by o ...
, in which clock genes are regulated by their protein products. Generally, the TTFL involves two main arms: positive regulatory elements that promote transcription and protein products that suppress transcription. When a positive regulatory element binds to a clock gene promoter
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription (genetics), transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can hav ...
, transcription proceeds, resulting in the creation of an mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
transcript, and then translation
Translation is the communication of the semantics, meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The English la ...
proceeds, resulting in a protein product. There are characteristic delays between mRNA transcript accumulation, protein accumulation, and gene suppression due to translation dynamics, post-translational protein modification, protein dimerization, and intracellular travel to the nucleus
Nucleus (: nuclei) is a Latin word for the seed inside a fruit. It most often refers to:
*Atomic nucleus, the very dense central region of an atom
*Cell nucleus, a central organelle of a eukaryotic cell, containing most of the cell's DNA
Nucleu ...
. Across species, proteins involved in the TTFL contain common structural motifs such PAS domain
A Per-Arnt-Sim (PAS) domain is a protein domain found in all kingdoms of life. Generally, the PAS domain acts as a molecular sensor, whereby small molecules and other proteins associate via binding of the PAS domain. Due to this sensing capabilit ...
s, involved in protein-protein interactions, and bHLH domains, involved in DNA binding.
Once enough modified protein products accumulate in the cytoplasm
The cytoplasm describes all the material within a eukaryotic or prokaryotic cell, enclosed by the cell membrane, including the organelles and excluding the nucleus in eukaryotic cells. The material inside the nucleus of a eukaryotic cell a ...
, they are transported into the nucleus where they inhibit the positive element from the promoter to stop transcription of clock genes. The clock gene is thus transcribed at low levels until its protein products are degraded, allowing for positive regulatory elements to bind to the promoter and restart transcription. The negative feedback loop of the TTFL has multiple properties important for the cellular circadian clock. First, it results in daily rhythms in both gene transcription and protein abundance and size, caused by the delay between translation and negative regulation of the gene. The cycle's period, or time required to complete one cycle, remains consistent in each individual and, barring mutation, is typically near 24 hours. This enables stable entrainment
Entrainment may refer to:
* Air entrainment, the intentional creation of tiny air bubbles in concrete
* Brainwave entrainment, the practice of entraining one's brainwaves to a desired frequency
* Entrainment (biomusicology), the synchronization o ...
to the 24 hour light-dark cycle that Earth experiences. Additionally, the protein products of clock genes control downstream genes that are not part of the feedback loop, allowing clock genes to create daily rhythms in other processes, such as metabolism, within the organism. Lastly, the TTFL is a limit cycle, meaning that it is a closed loop that will return to its fixed trajectory even if it is disturbed, maintaining the oscillatory path on its fixed 24-hour period.
Prominent models
The presence of the TTFL is highly conserved across animal species; however, many of the players involved in the process have changed across evolutionary time within different species. There are differences in the genes and proteins involved in the TTFL when comparing plants, animals, fungi and other eukaryotes. This suggests that a clock that follows the TTFL model has evolved multiple times during the existence of life.''Drosophila melanogaster''
The TTFL was first discovered in ''Drosophila'', and the system shares several components with the mammalian TTFL. Transcription of the clock genes, ''Period'' ''(per)'' and ''Timeless (tim)'', is initiated when positive elements ''Cycle'' (dCYC) and ''Clock'' (dCLK) form a heterodimer and bind E-box promoters, initiating transcription. During the day TIM is degraded; light exposure facilitates CRY binging to TIM, which leads to TIM's ubiquitination and eventual degradation. During the night, TIM and PER are able to form heterodimers and accumulate slowly in the cytoplasm, where PER is phosphorylated by the kinaseDOUBLETIME
''Doubletime'' is a documentary film about the sport of modern-day jump roping and Double Dutch. The film follows two disparate teams—one suburban white and one inner-city black—as they train to compete against each other for the very first ...
(DBT). The post-transcriptional modification of multiple phosphate groups both targets the complex for degradation and facilitates nuclear localization. In the nucleus, the PER-TIM dimer binds to the CYC-CLK dimer, which makes the CYC-CLK dimer release from the E-boxes and inhibits transcription. Once PER and TIM degrade, CYC-CLK dimers are able to bind the E-boxes again to initiate transcription, closing the negative feedback loop.
Mammals
BMAL1
Basic helix-loop-helix ARNT-like protein 1 or aryl hydrocarbon receptor nuclear translocator-like protein 1 (ARNTL), or brain and muscle ARNT-like 1 is a protein that in humans is encoded by the ''BMAL1'' gene on chromosome 11, region p15.3. It's ...
forms a heterodimer with CLOCK
A clock or chronometer is a device that measures and displays time. The clock is one of the oldest Invention, human inventions, meeting the need to measure intervals of time shorter than the natural units such as the day, the lunar month, a ...
to initiate transcription of ''mPer'' and ''cryptochrome
Cryptochromes (from the Greek κρυπτός χρώμα, "hidden colour") are a class of flavoproteins found in plants and animals that are sensitive to blue light. They are involved in the circadian rhythms and the sensing of magnetic fiel ...
'' (''cry''). There are three paralogs, or historically similar genes that now appear as a duplication, of the ''period'' gene in mammals listed as ''mPer1'', ''mPer2'', and ''mPer3''. There are also two paralogs of cryptochrome in mammals. PER and CRY proteins form a heterodimer, and PER's phosphorylation by CK1δ and CK1ε regulates the localization of the dimer to the nucleus. In the nucleus, PER-CRY negatively regulates the transcription of their cognate genes by binding BMAL1-CLOCK and causing their release from the E-box promoter.
Although the ''mPer'' paralogs work together as a functional ortholog of ''dPer'', they each have a distinguished function. ''mPer1'' and ''mPer2'' are necessary for clock function in the brain, while ''mPer3'' only plays a discernible role in the circadian rhythms of peripheral tissues. Knocking out
A knockout (abbreviated to KO or K.O.) is a fight-ending, winning criterion in several full-contact combat sports, such as boxing, kickboxing, Muay Thai, mixed martial arts, karate, some forms of taekwondo and other sports involving striking, a ...
either ''mPer1'' or ''mPer2'' causes a change in period, with ''mPer1'' knockouts free-running with a shorter period and ''mPer2'' knockouts free running with a longer period compared to the original tau before eventually becoming arrhythmic. Similarly, ''mCry1'' knockouts result in a shortened period and ''mCry2'' knockouts result in a lengthened period, with a double ''mCry1''/m''Cry2'' knockouts result in arrhythmicity.
There are also secondary loops in mammals, although they are more complex than those seen in ''Drosophila''. Like CWO in ''Drosophila'', Deleted in esophageal cancer1,2 (Dec1 Dec2) repress ''mPer'' expression by binding E-boxes which prevents CLOCK-BMAL1 from binding their targets. The receptors REV-ERB and retinoic acid-related orphan receptor (ROR) play a similar role to PDP1ε and VRI in ''Drosophila'', except they regulate CLOCK's binding partner BMAL1 instead of directly regulating CLOCK. D site-binding protein (DBP) and E4-binding protein (E4BP4) bind to the D-Box promoter sequence to regulate ''mPer'' expression.
The way these genes relate to ''Drosophila melanogaster'' is seen in the function of each of the genes and how they have evolutionarily changed. BMAL1
Basic helix-loop-helix ARNT-like protein 1 or aryl hydrocarbon receptor nuclear translocator-like protein 1 (ARNTL), or brain and muscle ARNT-like 1 is a protein that in humans is encoded by the ''BMAL1'' gene on chromosome 11, region p15.3. It's ...
is an ortholog
Sequence homology is the biological homology between DNA, RNA, or protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speci ...
of CYCLE. This means that BMAL1 and CYCLE appear to have a common history, but are found in different species. Another example of the parallels between ''Drosophila melanogaster'' and mammals is also seen in ''cry'' and ''mPer'' since they are functional orthologs of ''per'' and ''tim''.
Fungi: ''Neurospora''
Neurospora
''Neurospora'' is a genus of Ascomycete fungi. The genus name, meaning "nerve spore" refers to the characteristic striations on the spores that resemble axons.
The best known species in this genus is '' Neurospora crassa'', a common model organ ...
'' was identified as the second known clock gene in 1979 by JF Feldman and his colleagues. ''Frq'' was first cloned in 1989 by CR McClung and his colleagues. This gene was of particular interest because its expression is very complex compared to other known microbial genes. Two positive regulator proteins, White Collar-1 (WC-1) and White Collar-2 (WC-2) bind the frq promoter, which is called the Clock Box, during late subjective night to activate transcription. Light is also important for inducing FRQ expression; WC-1 is a photopigment, and light allows WC-1 and WC-2 to bind another promoter called the proximal light-response element (PLRE). FRQ protein negatively regulates the activity of WC-1 and WC-2. Several kinases (CK1, CK2, and PRD-4/checkpoint kinase 2) and phosphatases (PP1 and PP2A) regulate the ability of FRQ to translocate to the nucleus and FRQ, WC-1 and WC-2 stability.
Plants: ''Arabidopsis thaliana''
The first TTFL model was proposed for ''Arabidopsis thaliana
''Arabidopsis thaliana'', the thale cress, mouse-ear cress or arabidopsis, is a small plant from the mustard family (Brassicaceae), native to Eurasia and Africa. Commonly found along the shoulders of roads and in disturbed land, it is generally ...
'' in 2001 and included two MYB transcription factors, LATE ELONGATED HYPOCOTYL (LHY), CIRCADIAN CLOCK ASSOCIATED 1 (CCA1), and TIMING OF CAB EXPRESSION 1 (TOC1). CCA1 and LHY are expressed in the morning, and interact together to repress the expression of TOC1. CCA1 and LHY expression decreases in the darkness, allowing for TOC1 to express and negatively regulate CCA1 and LHY expression. CCA1 and LHY can also bind to their own promoter to repress their own transcription.
Cyanobacteria
Studies of the cyanobacteria clock led to the discovery of three essential clock genes: KaiA,KaiB Kaib, KaiB, or KAIB may refer to:
* KAIB (FM) Kaib, KaiB, or KAIB may refer to:
* KAIB (FM), one of the radio stations of Air 1
* KaiB, a gene
* KAI Bandara, an Indonesian railway operator
* Korea Aviation Accident Investigation Board
* Rami K ...
, and KaiC. Initially, these proteins were thought to follow the TTFL model similar to that proposed for eukarya
The eukaryotes ( ) constitute the domain of Eukaryota or Eukarya, organisms whose cells have a membrane-bound nucleus. All animals, plants, fungi, seaweeds, and many unicellular organisms are eukaryotes. They constitute a major group of l ...
, as there was a daily pattern in mRNA and protein abundance and level of phosphorylation, negative feedback of proteins on their cognate genes, resetting of clock phase in response to KaiC over-expression, and modified Kai activity through interactions with one another. Each of these results was consistent with understandings of the TTFL at the time. However, later studies have since concluded that post translational modifications such as phosphorylation are more important for clock control. When promoters for the Kai proteins were replaced with non-specific promoters, there was no interruption of the central feedback loop, as would be expected if inhibition occurred through the proteins’ feedback onto their specific promoters. As a consequence, the TTFL model has largely been determined to be inaccurate for cyanobacteria; transcriptional regulation is not the central process driving cyanobacteria rhythms. Though transcriptional and translational regulation are present, they were deemed to be effects of the clock rather than necessary for clock function.
Alternatives to the TTFL model
Post-translational feedback loops (PTFLs) involved in clock gene regulation have also been uncovered, often working in tandem with the TTFL model. In both mammals and plants, post-translational modifications such as phosphorylation and acetylation regulate the abundance and/or activity of clock genes and proteins. For example, levels of phosphorylation of TTFL components have been shown to vary rhythmically. These post-translational modifications can serve as degradation signals, binding regulators, and signals for the recruitment of additional factors. Notably, cyanobacteria demonstrate rhythmic 24-hour changes in phosphorylation in a feedback loop that is independent of transcription and translation: circadian rhythms in phosphorylation are observed when the feedback loop Kai proteins are placed in a test tube with ATP, independent of any other cellular machinery. This three-protein post-translational system is widely accepted to be the core oscillator, both necessary and sufficient to drive daily rhythms. In addition to the Kai system in cyanobacteria, oxidation of peroxiredoxin proteins has been shown to occur independently of transcription and translation in both mammalian red blood cells and algae ''Ostreococcus tauri'' cells; this system has been seen to be conserved in many organisms. It is not clear whether the peroxiredoxin system interacts with TTFL-based clocks or is itself a part of a new PTFL-based clock. However, both of these findings imply that in some organisms or cell types, PTFLs are sufficient to drive circadian rhythms.See also
*Chronobiology
Chronobiology is a field of biology that examines timing processes, including periodic (cyclic) phenomena in living organisms, such as their adaptation to solar- and lunar-related rhythms. These cycles are known as biological rhythms. Chron ...
* Circadian Rhythm
A circadian rhythm (), or circadian cycle, is a natural oscillation that repeats roughly every 24 hours. Circadian rhythms can refer to any process that originates within an organism (i.e., Endogeny (biology), endogenous) and responds to the env ...
* Familial sleep traits
Familial sleep traits are heritable variations in sleep patterns, resulting in abnormal sleep-wake times and/or abnormal sleep length.
Circadian rhythms are coordinated physiological and biological changes that oscillate on an approximately 24-h ...
* Period (Per) gene
References
{{Reflist Cell biology Circadian rhythm