proteogenomics on:
[Wikipedia]
[Google]
[Amazon]
 Proteogenomics is a field of biological research that utilizes a combination of proteomics, genomics, and transcriptomics to aid in the discovery and identification of peptides. Proteogenomics is used to identify new peptides by comparing
Proteogenomics is a field of biological research that utilizes a combination of proteomics, genomics, and transcriptomics to aid in the discovery and identification of peptides. Proteogenomics is used to identify new peptides by comparing
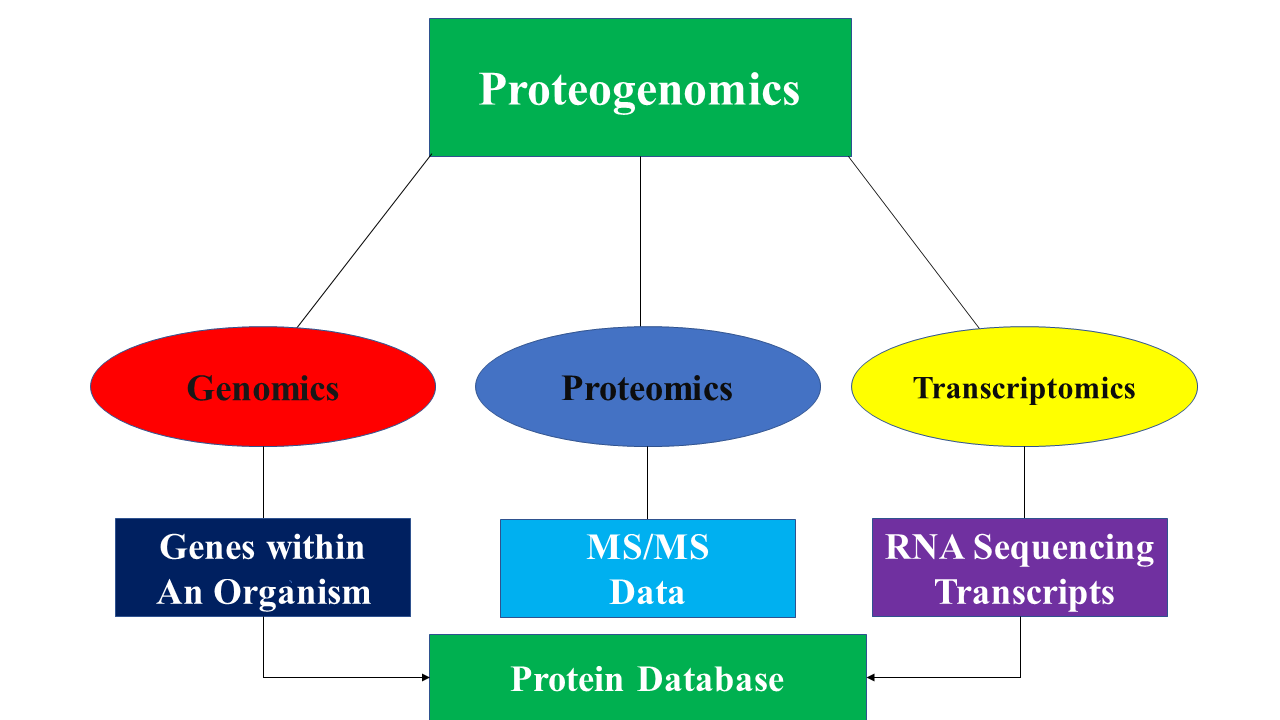
 The main idea behind the proteogenomic approach is to identify peptides by comparing MS/MS data to protein databases that contain predicted protein sequences. The protein database is generated in a variety of ways through the utilization of genomic and transcriptomic data. Below are some of the ways in which protein databases are generated:
The main idea behind the proteogenomic approach is to identify peptides by comparing MS/MS data to protein databases that contain predicted protein sequences. The protein database is generated in a variety of ways through the utilization of genomic and transcriptomic data. Below are some of the ways in which protein databases are generated:
 Proteogenomics is a field of biological research that utilizes a combination of proteomics, genomics, and transcriptomics to aid in the discovery and identification of peptides. Proteogenomics is used to identify new peptides by comparing
Proteogenomics is a field of biological research that utilizes a combination of proteomics, genomics, and transcriptomics to aid in the discovery and identification of peptides. Proteogenomics is used to identify new peptides by comparing MS/MS
Tandem mass spectrometry, also known as MS/MS or MS2, is a technique in instrumental analysis where two or more mass analyzers are coupled together using an additional reaction step to increase their abilities to analyse chemical samples. A comm ...
spectra against a protein database that has been derived from genomic and transcriptomic information. Proteogenomics often refers to studies that use proteomic information, often derived from mass spectrometry, to improve gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
annotations. The utilization of both proteomics and genomics data alongside advances in the availability and power of spectrographic and chromatographic technology led to the emergence of proteogenomics as its own field in 2004.
Proteomics deals with proteins in the same way that Genomics studies the genetic code of entire organisms, while Transcriptomics
Transcriptomics technologies are the techniques used to study an organism's transcriptome, the sum of all of its RNA transcripts. The information content of an organism is recorded in the DNA of its genome and expressed through transcription. H ...
deals with the study of RNA sequencing and transcripts. While all three fields might use forms of mass spectrometry and chromatography
In chemical analysis, chromatography is a laboratory technique for the separation of a mixture into its components. The mixture is dissolved in a fluid solvent (gas or liquid) called the ''mobile phase'', which carries it through a system ( ...
to identify and study the functions of DNA, RNA, and proteins, proteomics relies on the assumption that current gene models are correct and that all relevant protein sequences can be found in a reference database such as the Proteomics Identifications Database. Proteogenomics helps eliminate this reliance on existing, limited genetic models by combining datasets from multiple fields in order to produce a database of proteins or genetic markers. In addition, the emergence of novel protein sequences due to mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA replication, DNA or viral repl ...
s often cannot be accounted for in traditional proteomic databases, but can be predicted and studied using a synthesis of genomic and transcriptomic data.
The resulting research has applications in improving gene annotations, studying mutations, and understanding the effects of genetic manipulation
Genetic engineering, also called genetic modification or genetic manipulation, is the modification and manipulation of an organism's genes using technology. It is a set of technologies used to change the genetic makeup of cells, including ...
.
More recently, the joint profiling of surface proteins and mRNA transcripts from single cells by methods such as CITE-Seq and ESCAPE has been referred to as single-cell proteogenomics, although the goals of these studies are not related to peptide identification. Since 2019 these methods are more commonly referred to as multimodal omics or multi-omics.
History
Proteogenomics emerged as an independent field in 2004, based on the integration of technological advancements in next-generation sequencing genomics, and mass spectrometry proteomics. The term itself came into use that year, with the publication of a paper by George Church’s research group describing their discovery of a proteogenomic mapping technique that utilized proteomics data to better annotate the genome of the bacteria '' M. pneumoniae.'' By using a modern protein database, the lab mapped peptides detected in a whole cell onto a genetic scaffold using tandem mass spectrometry, then used the generated "hits" in order to create a "proteogenomic map" based on traditional genetic signals. The resulting map proved extremely accurate, with over 81% of predicted genomic reading frames being detected in the bacterial cells studied. In addition, the lab discovered several new frames not predicted via purely genetic methods, as well as some evidence supporting the idea that several predictions based genetic models could be false, proving the accuracy and cost-effectiveness of the hybrid technique. ''      '' The field expanded over the next two decades, initially using proteomics data to aid in refining genetic models via protein databases. In 2020s, one of the most common technique for identifying peptides involves usingtandem mass spectrometry
Tandem mass spectrometry, also known as MS/MS or MS2, is a technique in instrumental analysis where two or more mass analyzers are coupled together using an additional reaction step to increase their abilities to analyse chemical samples. A com ...
. This technique originated with Eng and Yates in 1994 which involves comparing a theoretical peptide fragment spectrum to compare an experimentally derived peptide spectrum to and outputting the most likely matches found. However, in the absence of an established peptide database, Proteogenomics instead compares the experimental spectrum to a genomic database instead which can then be used for genome annotation - as described in George Church's work. /sup> The latter technique has become more widely used over the last decade in large part due to the increasing affordability and speed of genomic sequencing techniques coupled with the increasing sensitivity of mass spectrometry-based proteomics.
Methodology
 The main idea behind the proteogenomic approach is to identify peptides by comparing MS/MS data to protein databases that contain predicted protein sequences. The protein database is generated in a variety of ways through the utilization of genomic and transcriptomic data. Below are some of the ways in which protein databases are generated:
The main idea behind the proteogenomic approach is to identify peptides by comparing MS/MS data to protein databases that contain predicted protein sequences. The protein database is generated in a variety of ways through the utilization of genomic and transcriptomic data. Below are some of the ways in which protein databases are generated:
Six-frame translation
Six-frame translations can be utilized to generate a database that predicts protein sequences. The limitation of this method is that databases will be very large due to the number of sequences that are generated, some of which do not exist in nature.Ab initio gene prediction
In this method, a protein base is generated by gene predicting algorithms that enable the identification of protein coding regions. The database is similar to one generated through six-frame translation in regards to the fact that the databases can be very large.Expressed sequence tag data
Six-frame translations can utilize anexpressed sequence tag In genetics, an expressed sequence tag (EST) is a short sub-sequence of a cDNA sequence. ESTs may be used to identify gene transcripts, and were instrumental in gene discovery and in gene-sequence determination. The identification of ESTs has proc ...
(EST) to generate protein databases. EST data provide transcription information that can aid in the creation of the database. The database can be very large and has the disadvantage of having multiple copies of a given sequence present; however, this problem can be circumvented by compressing the protein sequence generated through computational strategies.
Other methods
Protein databases can also be created by using RNA sequencing data, annotated RNA transcripts, and variant protein sequences. Also, there are other more specialized protein databases that can be made to appropriately identify the peptide of interest. Another method in the identification of proteins through proteogenomics is comparative proteogenomics. Comparative proteogenomics compares proteomic data from multiple related species concurrently and exploits the homology between their proteins to improve annotations with higher statistical confidence.Gupta N., Benhamida J., Bhargava V., Goodman D., Kain E., Kerman I., Nguyen N., Ollikainen N., Rodriguez J., Wang J., et al. Comparative proteogenomics: Combining mass spectrometry and comparative genomics to analyze multiple genomes. Genome Res. 2008;18:1133–1142.Gallien S., Perrodou E., Carapito C., Deshayes C., Reyrat J. M., Van Dorsselaer A., Poch O., Schaeffer C., Lecompte O. ( 2009) Ortho-proteogenomics: multiple proteomes investigation through orthology and a new MS-based protocol. Genome Res 19, 128– 135.Applications
Proteogenomics can be applied in different ways. One application is the improvement of gene annotations in various organisms. Gene annotation involves discovering genes and their functions. Proteogenomics has become especially useful in the discovery and improvement of gene annotations in prokaryotic organisms. For example, various microorganisms have had their genomic annotation studied through the proteogenomic approach including, ''Escherichia coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus '' Esc ...
'', '' Mycobacterium'', and multiple species of ''Shewanella
''Shewanella'' is the sole genus included in the marine bacteria family Shewanellaceae. Some species within it were formerly classed as '' Alteromonas''. ''Shewanella'' consists of facultatively anaerobic Gram-negative rods, most of which are f ...
'' bacteria.
Besides improving gene annotations, proteogenomic studies can also provide valuable information about the presence of programmed frameshift
Ribosomal frameshifting, also known as translational frameshifting or translational recoding, is a biological phenomenon that occurs during translation that results in the production of multiple, unique proteins from a single mRNA. The process can ...
s, N-terminal methionine excision, signal peptides, proteolysis and other post-translational modification
Post-translational modification (PTM) is the covalent and generally enzymatic modification of proteins following protein biosynthesis. This process occurs in the endoplasmic reticulum and the golgi apparatus. Proteins are synthesized by ribos ...
s.Gupta N., Tanner S., Jaitly N., Adkins J.N., Lipton M., Edwards R., Romine M., Osterman A., Bafna V., Smith R.D., et al. Whole proteome analysis of post-translational modifications: Applications of mass-spectrometry for proteogenomic annotation. Genome Res. 2007;17:1362–1377. Proteogenomics has potential applications in medicine, especially to oncology
Oncology is a branch of medicine that deals with the study, treatment, diagnosis and prevention of cancer. A medical professional who practices oncology is an ''oncologist''. The name's etymological origin is the Greek word ὄγκος (''à ...
research. Cancer occurs through genetic mutations such as methylation, translocation, and somatic mutations. Research has shown that both genomic and proteomic information are needed to understand the molecular variations that lead to cancer. Proteogenomics has aided in this through the identification of protein sequences that may have functional roles in cancer. A specific example of this occurred in a study involving colon cancer that resulted in the discovery of potential targets for cancer treatment. Proteogenomics has also led to personalized cancer targeting immunotherapies, where antibody epitopes for cancer antigens are predicted using proteogenomics to create medicines that act on the patient's specific tumor. In addition to treatment, proteogenonomics may provide insight into cancer diagnosis. In studies involving colon and rectal cancer, proteogenomics was utilized to identify somatic mutations. The identification of somatic mutations in patients could be used to diagnose cancer in patients. In addition to direct applications in cancer treatment and diagnosis, a proteogenomic approach can be used to study proteins that result in resistance to chemotherapy
Chemotherapy (often abbreviated to chemo and sometimes CTX or CTx) is a type of cancer treatment that uses one or more anti-cancer drugs ( chemotherapeutic agents or alkylating agents) as part of a standardized chemotherapy regimen. Chemothe ...
.
Challenges
Proteogenomics may offer methods of peptide identification without having the disadvantage of incomplete or inaccurate protein databases faced by proteomics; however, there are incurring challenges with the proteogenomic approach. One of the biggest challenges of proteogenomics is the sheer size of protein databases generated. statistically, a large protein database is more likely to result in the incorrect matching of the data from the protein database to the MS/MS data, this issue can hinder the identification of new peptides. False positives are also an issue through proteogenomic approaches. false positives can occur as a result of extremely large protein data bases where miss-matched data leads to incorrect identification. Another issue is the incorrect matching of MS/MS spectra to protein sequence data that corresponds to a similar peptide instead of the actual peptide. There are cases of receiving data of a peptide located at multiple gene sites, this can lead to data that can be interpreted in different ways. Despite these challenges, there are ways to reduce many of the errors that occur. For example, when dealing with a very large protein database, one could compare the identified novel peptide sequences to all of the sequences within the database and then compare the post translational modifications. Next it can be determined if the two sequences represent the same peptide or if they are two different peptides.References
{{Genomics Proteomics Genomics Mass spectrometry