
Protein folding is the
physical process by which a
protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, respon ...
chain is
translated to its native
three-dimensional structure, typically a "folded"
conformation by which the protein becomes biologically functional. Via an expeditious and reproducible process, a
polypeptide folds into its characteristic three-dimensional structure from a
random coil.
Each protein exists first as an unfolded polypeptide or random coil after being translated from a sequence of
mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
to a linear chain of
amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha ...
s. At this stage the polypeptide lacks any stable (long-lasting) three-dimensional structure (the left hand side of the first figure). As the polypeptide chain is being synthesized by a
ribosome, the linear chain begins to fold into its three-dimensional structure.
Folding of many proteins begins even during translation of the polypeptide chain. Amino acids interact with each other to produce a well-defined three-dimensional structure, the folded protein (the right hand side of the figure), known as the
native state. The resulting three-dimensional structure is determined by the amino acid sequence or primary structure (
Anfinsen's dogma).
The correct three-dimensional structure is essential to function, although some parts of functional proteins
may remain unfolded, so that
protein dynamics is important. Failure to fold into native structure generally produces inactive proteins, but in some instances misfolded proteins have modified or toxic functionality. Several
neurodegenerative and other
disease
A disease is a particular abnormal condition that negatively affects the structure or function of all or part of an organism, and that is not immediately due to any external injury. Diseases are often known to be medical conditions that a ...
s are believed to result from the accumulation of
amyloid fibrils formed by misfolded proteins, infectious varieties of which are known as
prions.
Many
allergies are caused by incorrect folding of some proteins, because the
immune system
The immune system is a network of biological processes that protects an organism from diseases. It detects and responds to a wide variety of pathogens, from viruses to parasitic worms, as well as Tumor immunology, cancer cells and objects such ...
does not produce
antibodies for certain protein structures.
Denaturation of proteins is a process of transition from the folded to the
unfolded state. It happens in
cooking, in
burn
A burn is an injury to skin, or other tissues, caused by heat, cold, electricity, chemicals, friction, or ultraviolet radiation (like sunburn). Most burns are due to heat from hot liquids (called scalding), solids, or fire. Burns occur mainl ...
s, in
proteinopathies, and in other contexts.
The duration of the folding process varies dramatically depending on the protein of interest. When studied
outside the cell, the slowest folding proteins require many minutes or hours to fold primarily due to
proline isomerization In epigenetics, proline isomerization is the effect that ''cis-trans'' isomerization of the amino acid proline has on the regulation of gene expression. Similar to aspartic acid, the amino acid proline has the rare property of being able to occupy ...
, and must pass through a number of intermediate states, like checkpoints, before the process is complete. On the other hand, very small single-
domain proteins with lengths of up to a hundred amino acids typically fold in a single step. Time scales of milliseconds are the norm and the very fastest known protein folding reactions are complete within a few microseconds. The folding time scale of a protein depends on its size,
contact order and
circuit topology.
Understanding and simulating the protein folding process has been an important challenge for
computational biology since the late 1960s.
Process of protein folding
Primary structure
The
primary structure of a protein, its linear amino-acid sequence, determines its native conformation.
The specific amino acid residues and their position in the polypeptide chain are the determining factors for which portions of the protein fold closely together and form its three-dimensional conformation. The amino acid composition is not as important as the sequence.
The essential fact of folding, however, remains that the amino acid sequence of each protein contains the information that specifies both the native structure and the pathway to attain that state. This is not to say that nearly identical amino acid sequences always fold similarly. Conformations differ based on environmental factors as well; similar proteins fold differently based on where they are found.
Secondary structure


Formation of a
secondary structure is the first step in the folding process that a protein takes to assume its native structure. Characteristic of secondary structure are the structures known as
alpha helices and
beta sheets that fold rapidly because they are stabilized by
intramolecular hydrogen bonds, as was first characterized by
Linus Pauling. Formation of intramolecular hydrogen bonds provides another important contribution to protein stability.
α-helices are formed by hydrogen bonding of the
backbone to form a spiral shape (refer to figure on the right).
The β pleated sheet is a structure that forms with the backbone bending over itself to form the hydrogen bonds (as displayed in the figure to the left). The hydrogen bonds are between the amide hydrogen and carbonyl oxygen of the
peptide bond
In organic chemistry, a peptide bond is an amide type of covalent chemical bond linking two consecutive alpha-amino acids from C1 (carbon number one) of one alpha-amino acid and N2 ( nitrogen number two) of another, along a peptide or protein c ...
. There exists anti-parallel β pleated sheets and parallel β pleated sheets where the stability of the hydrogen bonds is stronger in the anti-parallel β sheet as it hydrogen bonds with the ideal 180 degree angle compared to the slanted hydrogen bonds formed by parallel sheets.
Tertiary structure
The α-Helices and β-Sheets are commonly amphipathic, meaning they have a hydrophilic and a hydrophobic portion. This ability helps in forming tertiary structure of a protein in which folding occurs so that the hydrophilic sides are facing the aqueous environment surrounding the protein and the hydrophobic sides are facing the hydrophobic core of the protein.
Secondary structure hierarchically gives way to tertiary structure formation. Once the protein's tertiary structure is formed and stabilized by the hydrophobic interactions, there may also be
covalent bonding in the form of
disulfide bridges formed between two
cysteine residues. These non-covalent and covalent contacts take a specific
topological arrangement in a native structure of a protein. Tertiary structure of a protein involves a single polypeptide chain; however, additional interactions of folded polypeptide chains give rise to quaternary structure formation.
Quaternary structure
Tertiary structure may give way to the formation of
quaternary structure in some proteins, which usually involves the "assembly" or "coassembly" of subunits that have already folded; in other words, multiple polypeptide chains could interact to form a fully functional quaternary protein.
Driving forces of protein folding

Folding is a
spontaneous process that is mainly guided by hydrophobic interactions, formation of intramolecular
hydrogen bonds,
van der Waals forces, and it is opposed by
conformational entropy
In chemical thermodynamics, conformational entropy is the entropy associated with the number of conformations of a molecule. The concept is most commonly applied to biological macromolecules such as proteins and RNA, but also be used for pol ...
. The process of folding often begins
co-translationally, so that the
N-terminus of the protein begins to fold while the
C-terminal
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is ...
portion of the protein is still being
synthesized by the
ribosome; however, a protein molecule may fold spontaneously during or after
biosynthesis. While these
macromolecules may be regarded as "
folding themselves", the process also depends on the
solvent
A solvent (s) (from the Latin '' solvō'', "loosen, untie, solve") is a substance that dissolves a solute, resulting in a solution. A solvent is usually a liquid but can also be a solid, a gas, or a supercritical fluid. Water is a solvent for ...
(
water
Water (chemical formula ) is an inorganic, transparent, tasteless, odorless, and nearly colorless chemical substance, which is the main constituent of Earth's hydrosphere and the fluids of all known living organisms (in which it acts as ...
or
lipid bilayer), the concentration of
salts, the
pH, the
temperature
Temperature is a physical quantity that expresses quantitatively the perceptions of hotness and coldness. Temperature is measured with a thermometer.
Thermometers are calibrated in various temperature scales that historically have relied on ...
, the possible presence of cofactors and of molecular
chaperones.
Proteins will have limitations on their folding abilities by the restricted bending angles or conformations that are possible. These allowable angles of protein folding are described with a two-dimensional plot known as the
Ramachandran plot, depicted with psi and phi angles of allowable rotation.
Hydrophobic effect
Protein folding must be thermodynamically favorable within a cell in order for it to be a spontaneous reaction. Since it is known that protein folding is a spontaneous reaction, then it must assume a negative
Gibbs free energy value. Gibbs free energy in protein folding is directly related to
enthalpy and
entropy
Entropy is a scientific concept, as well as a measurable physical property, that is most commonly associated with a state of disorder, randomness, or uncertainty. The term and the concept are used in diverse fields, from classical thermodyna ...
.
For a negative delta G to arise and for protein folding to become thermodynamically favorable, then either enthalpy, entropy, or both terms must be favorable.
Minimizing the number of hydrophobic side-chains exposed to water is an important driving force behind the folding process.
The hydrophobic effect is the phenomenon in which the hydrophobic chains of a protein collapse into the core of the protein (away from the hydrophilic environment).
In an aqueous environment, the water molecules tend to aggregate around the hydrophobic regions or side chains of the protein, creating water shells of ordered water molecules. An ordering of water molecules around a hydrophobic region increases order in a system and therefore contributes a negative change in entropy (less entropy in the system). The water molecules are fixed in these water cages which drives the
hydrophobic collapse, or the inward folding of the hydrophobic groups. The hydrophobic collapse introduces entropy back to the system via the breaking of the water cages which frees the ordered water molecules.
The multitude of hydrophobic groups interacting within the core of the globular folded protein contributes a significant amount to protein stability after folding, because of the vastly accumulated van der Waals forces (specifically
London Dispersion forces
London dispersion forces (LDF, also known as dispersion forces, London forces, instantaneous dipole–induced dipole forces, fluctuating induced dipole bonds or loosely as van der Waals forces) are a type of intermolecular force acting between at ...
).
The
hydrophobic effect exists as a driving force in thermodynamics only if there is the presence of an aqueous medium with an
amphiphilic molecule containing a large hydrophobic region. The strength of hydrogen bonds depends on their environment; thus, H-bonds enveloped in a hydrophobic core contribute more than H-bonds exposed to the aqueous environment to the stability of the native state.
In proteins with globular folds, hydrophobic amino acids tend to be interspersed along the primary sequence, rather than randomly distributed or clustered together. However, proteins that have recently been born
de novo, which tend to be
intrinsically disordered, show the opposite pattern of hydrophobic amino acid clustering along the primary sequence.
Chaperones
 Molecular chaperones
Molecular chaperones are a class of proteins that aid in the correct folding of other proteins ''
in vivo''. Chaperones exist in all cellular compartments and interact with the polypeptide chain in order to allow the native three-dimensional conformation of the protein to form; however, chaperones themselves are not included in the final structure of the protein they are assisting in.
Chaperones may assist in folding even when the nascent polypeptide is being synthesized by the ribosome.
Molecular chaperones operate by binding to stabilize an otherwise unstable structure of a protein in its folding pathway, but chaperones do not contain the necessary information to know the correct native structure of the protein they are aiding; rather, chaperones work by preventing incorrect folding conformations.
In this way, chaperones do not actually increase the rate of individual steps involved in the folding pathway toward the native structure; instead, they work by reducing possible unwanted aggregations of the polypeptide chain that might otherwise slow down the search for the proper intermediate and they provide a more efficient pathway for the polypeptide chain to assume the correct conformations.
Chaperones are not to be confused with folding
catalyst proteins, which catalyze chemical reactions responsible for slow steps in folding pathways. Examples of folding catalysts are protein
disulfide isomerase
Protein disulfide isomerase (), or PDI, is an enzyme in the endoplasmic reticulum (ER) in eukaryotes and the periplasm of bacteria that catalyzes the formation and breakage of disulfide bonds between cysteine residues within proteins as t ...
s and
peptidyl-prolyl isomerase
Prolyl isomerase (also known as peptidylprolyl isomerase or PPIase) is an enzyme () found in both prokaryotes and eukaryotes that interconverts the ''cis'' and ''trans'' isomers of peptide bonds with the amino acid proline. Proline has an unusua ...
s that may be involved in formation of
disulfide bonds or interconversion between cis and trans stereoisomers of peptide group.
Chaperones are shown to be critical in the process of protein folding ''in vivo'' because they provide the protein with the aid needed to assume its proper alignments and conformations efficiently enough to become "biologically relevant".
This means that the polypeptide chain could theoretically fold into its native structure without the aid of chaperones, as demonstrated by protein folding experiments conducted ''
in vitro
''In vitro'' (meaning in glass, or ''in the glass'') studies are performed with microorganisms, cells, or biological molecules outside their normal biological context. Colloquially called "test-tube experiments", these studies in biology and ...
'';
however, this process proves to be too inefficient or too slow to exist in biological systems; therefore, chaperones are necessary for protein folding ''in vivo.'' Along with its role in aiding native structure formation, chaperones are shown to be involved in various roles such as protein transport, degradation, and even allow
denatured proteins exposed to certain external denaturant factors an opportunity to refold into their correct native structures.
A fully denatured protein lacks both tertiary and secondary structure, and exists as a so-called
random coil. Under certain conditions some proteins can refold; however, in many cases, denaturation is irreversible.
Cells sometimes protect their proteins against the denaturing influence of heat with
enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different molecule ...
s known as
heat shock proteins (a type of chaperone), which assist other proteins both in folding and in remaining folded.
Heat shock proteins have been found in all species examined, from
bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were am ...
to humans, suggesting that they evolved very early and have an important function. Some proteins never fold in cells at all except with the assistance of chaperones which either isolate individual proteins so that their folding is not interrupted by interactions with other proteins or help to unfold misfolded proteins, allowing them to refold into the correct native structure.
This function is crucial to prevent the risk of
precipitation into
insoluble amorphous aggregates. The external factors involved in protein denaturation or disruption of the native state include temperature, external fields (electric, magnetic),
molecular crowding,
and even the limitation of space (i.e. confinement), which can have a big influence on the folding of proteins. High concentrations of
solutes, extremes of
pH, mechanical forces, and the presence of chemical denaturants can contribute to protein denaturation, as well. These individual factors are categorized together as stresses. Chaperones are shown to exist in increasing concentrations during times of cellular stress and help the proper folding of emerging proteins as well as denatured or misfolded ones.
Under some conditions proteins will not fold into their biochemically functional forms. Temperatures above or below the range that cells tend to live in will cause
thermally unstable proteins to unfold or denature (this is why boiling makes an
egg white turn opaque). Protein thermal stability is far from constant, however; for example,
hyperthermophilic bacteria have been found that grow at temperatures as high as 122 °C, which of course requires that their full complement of vital proteins and protein assemblies be stable at that temperature or above.
The bacterium ''
E. coli'' is the host for
bacteriophage T4, and the phage encoded gp31 protein () appears to be structurally and functionally homologous to ''E. coli''
chaperone protein GroES and able to substitute for it in the assembly of bacteriophage T4
virus
A virus is a wikt:submicroscopic, submicroscopic infectious agent that replicates only inside the living Cell (biology), cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and ...
particles during infection.
[Marusich EI, Kurochkina LP, Mesyanzhinov VV. Chaperones in bacteriophage T4 assembly. Biochemistry (Mosc). 1998;63(4):399-406] Like GroES, gp31 forms a stable complex with
GroEL chaperonin that is absolutely necessary for the folding and assembly in vivo of the bacteriophage T4 major capsid protein gp23.
[
]
Fold switching
Some proteins have multiple native structures, and change their fold based on some external factors. For example, the KaiB protein switches fold throughout the day, acting as a clock for cyanobacteria. It has been estimated that around 0.5–4% of PDB (Protein Data Bank
The Protein Data Bank (PDB) is a database for the three-dimensional structural data of large biological molecules, such as proteins and nucleic acids. The data, typically obtained by X-ray crystallography, NMR spectroscopy, or, increasingly, c ...
) proteins switch folds.
Protein misfolding and neurodegenerative disease
A protein is considered to be misfolded
Protein folding is the physical process by which a protein chain is Translation (biology), translated to its native protein tertiary structure, three-dimensional structure, typically a "folded" Protein structure, conformation by which the prote ...
if it cannot achieve its normal native state. This can be due to mutations in the amino acid sequence or a disruption of the normal folding process by external factors.prion
Prions are misfolded proteins that have the ability to transmit their misfolded shape onto normal variants of the same protein. They characterize several fatal and transmissible neurodegenerative diseases in humans and many other animals. It ...
-related illnesses such as Creutzfeldt–Jakob disease, bovine spongiform encephalopathy (mad cow disease), amyloid-related illnesses such as Alzheimer's disease and familial amyloid cardiomyopathy or polyneuropathy,Huntington's
Huntington's disease (HD), also known as Huntington's chorea, is a neurodegenerative disease that is mostly inherited. The earliest symptoms are often subtle problems with mood or mental abilities. A general lack of coordination and an unst ...
and Parkinson's disease
Parkinson's disease (PD), or simply Parkinson's, is a long-term degenerative disorder of the central nervous system that mainly affects the motor system. The symptoms usually emerge slowly, and as the disease worsens, non-motor symptoms becom ...
.
Experimental techniques for studying protein folding
While inferences about protein folding can be made through mutation studies, typically, experimental techniques for studying protein folding rely on the gradual unfolding or folding of proteins and observing conformational changes using standard non-crystallographic techniques.
X-ray crystallography
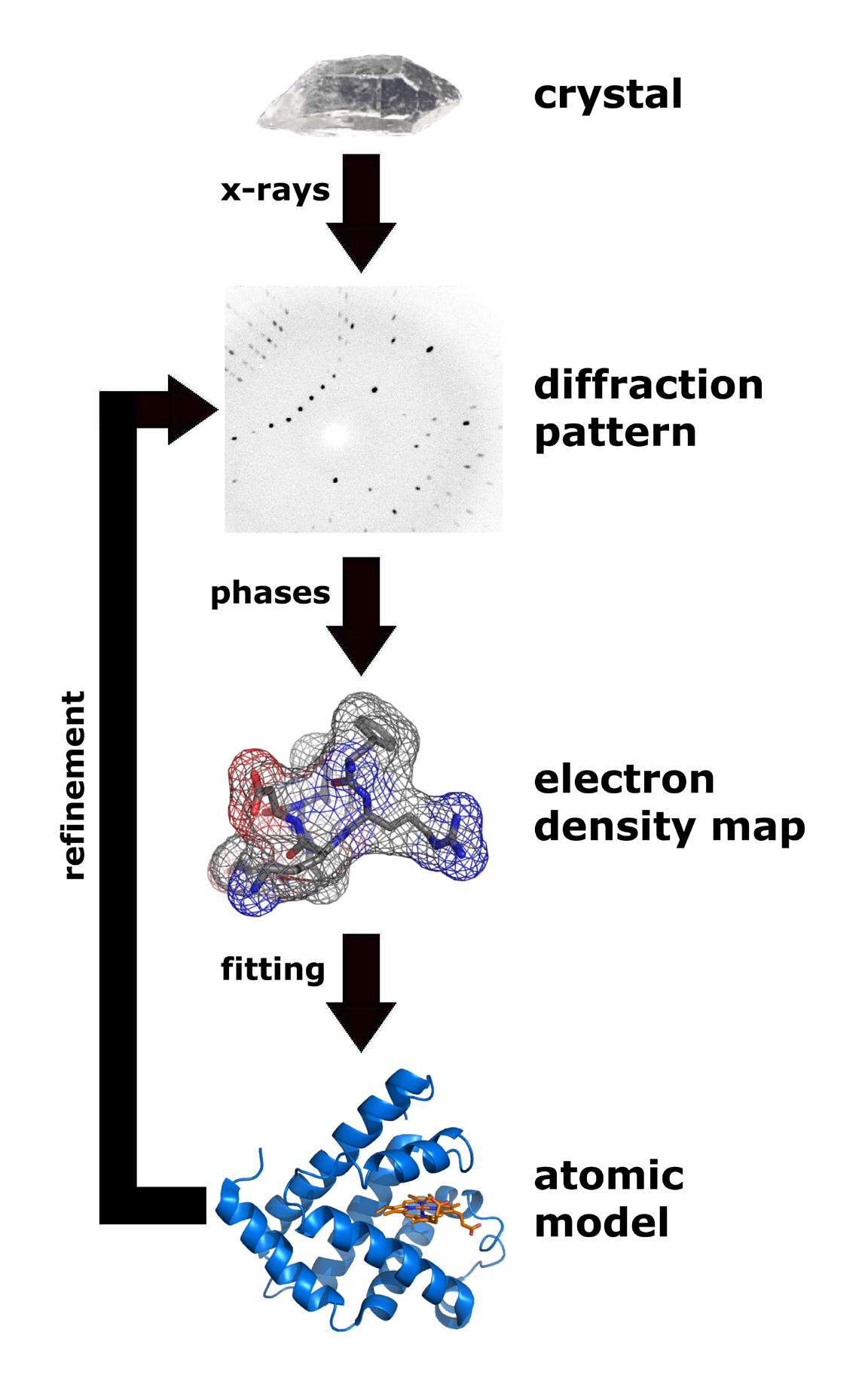
X-ray crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angle ...
is one of the more efficient and important methods for attempting to decipher the three dimensional configuration of a folded protein.
Fluorescence spectroscopy
Fluorescence spectroscopy is a highly sensitive method for studying the folding state of proteins. Three amino acids, phenylalanine (Phe), tyrosine (Tyr) and tryptophan (Trp), have intrinsic fluorescence properties, but only Tyr and Trp are used experimentally because their quantum yields are high enough to give good fluorescence signals. Both Trp and Tyr are excited by a wavelength of 280 nm, whereas only Trp is excited by a wavelength of 295 nm. Because of their aromatic character, Trp and Tyr residues are often found fully or partially buried in the hydrophobic core of proteins, at the interface between two protein domains, or at the interface between subunits of oligomeric proteins. In this apolar environment, they have high quantum yields and therefore high fluorescence intensities. Upon disruption of the protein's tertiary or quaternary structure, these side chains become more exposed to the hydrophilic environment of the solvent, and their quantum yields decrease, leading to low fluorescence intensities. For Trp residues, the wavelength of their maximal fluorescence emission also depend on their environment.
Fluorescence spectroscopy can be used to characterize the equilibrium unfolding of proteins by measuring the variation in the intensity of fluorescence emission or in the wavelength of maximal emission as functions of a denaturant value.
Circular dichroism
Circular dichroism is one of the most general and basic tools to study protein folding. Circular dichroism spectroscopy measures the absorption of circularly polarized light
In electrodynamics, circular polarization of an electromagnetic wave is a polarization state in which, at each point, the electromagnetic field of the wave has a constant magnitude and is rotating at a constant rate in a plane perpendicular to t ...
. In proteins, structures such as alpha helices and beta sheets are chiral, and thus absorb such light. The absorption of this light acts as a marker of the degree of foldedness of the protein ensemble. This technique has been used to measure equilibrium unfolding of the protein by measuring the change in this absorption as a function of denaturant concentration or temperature
Temperature is a physical quantity that expresses quantitatively the perceptions of hotness and coldness. Temperature is measured with a thermometer.
Thermometers are calibrated in various temperature scales that historically have relied on ...
. A denaturant melt measures the free energy of unfolding as well as the protein's m value, or denaturant dependence. A temperature melt measures the denaturation temperature (Tm) of the protein.kinetics
Kinetics ( grc, κίνησις, , kinesis, ''movement'' or ''to move'') may refer to:
Science and medicine
* Kinetics (physics), the study of motion and its causes
** Rigid body kinetics, the study of the motion of rigid bodies
* Chemical k ...
and to generate chevron plots.
Vibrational circular dichroism of proteins
The more recent developments of vibrational circular dichroism (VCD) techniques for proteins, currently involving Fourier transform (FT) instruments, provide powerful means for determining protein conformations in solution even for very large protein molecules. Such VCD studies of proteins can be combined with X-ray diffraction data for protein crystals, FT-IR data for protein solutions in heavy water (D2O), or quantum computations.
Protein nuclear magnetic resonance spectroscopy
Protein nuclear magnetic resonance (NMR) is able to collect protein structural data by inducing a magnet field through samples of concentrated protein. In NMR, depending on the chemical environment, certain nuclei will absorb specific radio-frequencies.COSY
Cosy may refer to
* Tea cosy, a cover for a teapot
* Cozy mystery, a subgenre of crime fiction
* Cosy catastrophe, post-apocalyptic science fiction style
* Correlation spectroscopy (COSY)
* CoSy (Conferencing System), an early computer conferenc ...
, TOCSY, HSQC The heteronuclear single quantum coherence or heteronuclear single quantum correlation experiment, normally abbreviated as HSQC, is used frequently in NMR spectroscopy of organic molecules and is of particular significance in the field of protein ...
, time relaxation (T1 & T2), and NOE. Because protein folding takes place in about 50 to 3000 s−1 CPMG Relaxation dispersion and chemical exchange saturation transfer have become some of the primary techniques for NMR analysis of folding.
Because protein folding takes place in about 50 to 3000 s−1 CPMG Relaxation dispersion and chemical exchange saturation transfer have become some of the primary techniques for NMR analysis of folding.X-ray crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angle ...
.amyotrophic lateral sclerosis
Amyotrophic lateral sclerosis (ALS), also known as motor neuron disease (MND) or Lou Gehrig's disease, is a neurodegenerative disease that results in the progressive loss of motor neurons that control voluntary muscles. ALS is the most comm ...
involved protein SOD1, excited intermediates were studied with relaxation dispersion and Saturation transfer.
Dual-polarization interferometry
Dual polarisation interferometry is a surface-based technique for measuring the optical properties of molecular layers. When used to characterize protein folding, it measures the conformation by determining the overall size of a monolayer of the protein and its density in real time at sub-Angstrom resolution,temperature
Temperature is a physical quantity that expresses quantitatively the perceptions of hotness and coldness. Temperature is measured with a thermometer.
Thermometers are calibrated in various temperature scales that historically have relied on ...
.
Studies of folding with high time resolution
The study of protein folding has been greatly advanced in recent years by the development of fast, time-resolved techniques. Experimenters rapidly trigger the folding of a sample of unfolded protein and observe the resulting dynamics. Fast techniques in use include neutron scattering,Martin Gruebele
Martin Gruebele (born January 10, 1964 in Stuttgart, Germany) is a German-born American physical chemist and biophysicist who is currently James R. Eiszner Professor of Chemistry, Professor of Physics, Professor of Biophysics and Computational Biol ...
, Brian Dyer, William Eaton, Sheena Radford
Sheena Elizabeth Radford FRS FMedSci is a British biophysicist, and Astbury Professor of Biophysics in the Astbury Centre for Structural Molecular Biology, School of Molecular and Cellular Biology at the University of Leeds. Radford is the A ...
, Chris Dobson, Alan Fersht, Bengt Nölting and Lars Konermann.
Proteolysis
Proteolysis is routinely used to probe the fraction unfolded under a wide range of solution conditions (e.g. fast parallel proteolysis (FASTpp)
Fast parallel proteolysis (FASTpp) is a method to determine the thermostability of proteins by measuring which fraction of protein resists rapid proteolytic digestion.
History and background
Proteolysis is widely used in biochemistry and cell b ...
.
Single-molecule force spectroscopy
Single molecule techniques such as optical tweezers and AFM have been used to understand protein folding mechanisms of isolated proteins as well as proteins with chaperones.
Biotin painting
Biotin painting enables condition-specific cellular snapshots of (un)folded proteins. Biotin 'painting' shows a bias towards predicted Intrinsically disordered proteins.
Computational studies of protein folding
Computational studies of protein folding includes three main aspects related to the prediction of protein stability, kinetics, and structure. A 2013 review summarizes the available computational methods for protein folding.
Levinthal's paradox
In 1969, Cyrus Levinthal noted that, because of the very large number of degrees of freedom in an unfolded polypeptide chain, the molecule has an astronomical number of possible conformations. An estimate of 3300 or 10143 was made in one of his papers. Levinthal's paradox is a thought experiment based on the observation that if a protein were folded by sequential sampling of all possible conformations, it would take an astronomical amount of time to do so, even if the conformations were sampled at a rapid rate (on the nanosecond or picosecond scale). Based upon the observation that proteins fold much faster than this, Levinthal then proposed that a random conformational search does not occur, and the protein must, therefore, fold through a series of meta-stable intermediate states.
Energy landscape of protein folding
 The configuration space of a protein during folding can be visualized as an energy landscape. According to Joseph Bryngelson and Peter Wolynes, proteins follow the ''principle of minimal frustration'', meaning that naturally evolved proteins have optimized their folding energy landscapes,
The configuration space of a protein during folding can be visualized as an energy landscape. According to Joseph Bryngelson and Peter Wolynes, proteins follow the ''principle of minimal frustration'', meaning that naturally evolved proteins have optimized their folding energy landscapes,design
A design is a plan or specification for the construction of an object or system or for the implementation of an activity or process or the result of that plan or specification in the form of a prototype, product, or process. The verb ''to design'' ...
.
Modeling of protein folding
'' De novo'' or '' ab initio'' techniques for computational protein structure prediction can be used for simulating various aspects of protein folding. Molecular Dynamics
Molecular dynamics (MD) is a computer simulation method for analyzing the physical movements of atoms and molecules. The atoms and molecules are allowed to interact for a fixed period of time, giving a view of the dynamic "evolution" of th ...
(MD) was used in simulations of protein folding and dynamics in silico.
/ref> target protein folding.
Long continuous-trajectory simulations have been performed on Anton, a massively parallel supercomputer designed and built around custom ASICs and interconnects by D. E. Shaw Research
D. E. Shaw Research (DESRES) is a privately held biochemistry research company based in New York City. Under the scientific direction of David E. Shaw, the group's chief scientist, D. E. Shaw Research develops technologies for molecular dynami ...
. The longest published result of a simulation performed using Anton is a 2.936 millisecond simulation of NTL9 at 355 K.
See also
* Chevron plot
* Denaturation midpoint
* Downhill folding
Downhill folding is a process in which a protein folds without encountering any significant macroscopic free energy barrier. It is a key prediction of the folding funnel hypothesis of the energy landscape theory of proteins.
Overview
Downhil ...
* Folding (chemistry)
* Phi value analysis
* Potential energy of protein
* Protein dynamics
* Protein misfolding cyclic amplification
* Protein structure prediction software
This list of protein structure prediction software summarizes notable used software tools in protein structure prediction, including homology modeling, protein threading, ''ab initio'' methods, secondary structure prediction, and transmembrane h ...
* Proteopathy
* Time-resolved mass spectrometry
Time-resolved mass spectrometry (TRMS) is a strategy in analytical chemistry that uses mass spectrometry platform to collect data with temporal resolution. Implementation of TRMS builds on the ability of mass spectrometers to process ions within s ...
References
External links
Human Proteome Folding Project
{{Authority control
Protein structure
 Protein folding is the physical process by which a
Protein folding is the physical process by which a 
 Formation of a secondary structure is the first step in the folding process that a protein takes to assume its native structure. Characteristic of secondary structure are the structures known as alpha helices and beta sheets that fold rapidly because they are stabilized by intramolecular hydrogen bonds, as was first characterized by Linus Pauling. Formation of intramolecular hydrogen bonds provides another important contribution to protein stability. α-helices are formed by hydrogen bonding of the backbone to form a spiral shape (refer to figure on the right). The β pleated sheet is a structure that forms with the backbone bending over itself to form the hydrogen bonds (as displayed in the figure to the left). The hydrogen bonds are between the amide hydrogen and carbonyl oxygen of the
Formation of a secondary structure is the first step in the folding process that a protein takes to assume its native structure. Characteristic of secondary structure are the structures known as alpha helices and beta sheets that fold rapidly because they are stabilized by intramolecular hydrogen bonds, as was first characterized by Linus Pauling. Formation of intramolecular hydrogen bonds provides another important contribution to protein stability. α-helices are formed by hydrogen bonding of the backbone to form a spiral shape (refer to figure on the right). The β pleated sheet is a structure that forms with the backbone bending over itself to form the hydrogen bonds (as displayed in the figure to the left). The hydrogen bonds are between the amide hydrogen and carbonyl oxygen of the  Folding is a spontaneous process that is mainly guided by hydrophobic interactions, formation of intramolecular hydrogen bonds, van der Waals forces, and it is opposed by
Folding is a spontaneous process that is mainly guided by hydrophobic interactions, formation of intramolecular hydrogen bonds, van der Waals forces, and it is opposed by  Molecular chaperones are a class of proteins that aid in the correct folding of other proteins '' in vivo''. Chaperones exist in all cellular compartments and interact with the polypeptide chain in order to allow the native three-dimensional conformation of the protein to form; however, chaperones themselves are not included in the final structure of the protein they are assisting in. Chaperones may assist in folding even when the nascent polypeptide is being synthesized by the ribosome. Molecular chaperones operate by binding to stabilize an otherwise unstable structure of a protein in its folding pathway, but chaperones do not contain the necessary information to know the correct native structure of the protein they are aiding; rather, chaperones work by preventing incorrect folding conformations. In this way, chaperones do not actually increase the rate of individual steps involved in the folding pathway toward the native structure; instead, they work by reducing possible unwanted aggregations of the polypeptide chain that might otherwise slow down the search for the proper intermediate and they provide a more efficient pathway for the polypeptide chain to assume the correct conformations. Chaperones are not to be confused with folding catalyst proteins, which catalyze chemical reactions responsible for slow steps in folding pathways. Examples of folding catalysts are protein
Molecular chaperones are a class of proteins that aid in the correct folding of other proteins '' in vivo''. Chaperones exist in all cellular compartments and interact with the polypeptide chain in order to allow the native three-dimensional conformation of the protein to form; however, chaperones themselves are not included in the final structure of the protein they are assisting in. Chaperones may assist in folding even when the nascent polypeptide is being synthesized by the ribosome. Molecular chaperones operate by binding to stabilize an otherwise unstable structure of a protein in its folding pathway, but chaperones do not contain the necessary information to know the correct native structure of the protein they are aiding; rather, chaperones work by preventing incorrect folding conformations. In this way, chaperones do not actually increase the rate of individual steps involved in the folding pathway toward the native structure; instead, they work by reducing possible unwanted aggregations of the polypeptide chain that might otherwise slow down the search for the proper intermediate and they provide a more efficient pathway for the polypeptide chain to assume the correct conformations. Chaperones are not to be confused with folding catalyst proteins, which catalyze chemical reactions responsible for slow steps in folding pathways. Examples of folding catalysts are protein 
 Because protein folding takes place in about 50 to 3000 s−1 CPMG Relaxation dispersion and chemical exchange saturation transfer have become some of the primary techniques for NMR analysis of folding. In addition, both techniques are used to uncover excited intermediate states in the protein folding landscape. To do this, CPMG Relaxation dispersion takes advantage of the spin echo phenomenon. This technique exposes the target nuclei to a 90 pulse followed by one or more 180 pulses. As the nuclei refocus, a broad distribution indicates the target nuclei is involved in an intermediate excited state. By looking at Relaxation dispersion plots the data collect information on the thermodynamics and kinetics between the excited and ground. Saturation Transfer measures changes in signal from the ground state as excited states become perturbed. It uses weak radio frequency irradiation to saturate the excited state of a particular nuclei which transfers its saturation to the ground state. This signal is amplified by decreasing the magnetization (and the signal) of the ground state.
The main limitations in NMR is that its resolution decreases with proteins that are larger than 25 kDa and is not as detailed as
Because protein folding takes place in about 50 to 3000 s−1 CPMG Relaxation dispersion and chemical exchange saturation transfer have become some of the primary techniques for NMR analysis of folding. In addition, both techniques are used to uncover excited intermediate states in the protein folding landscape. To do this, CPMG Relaxation dispersion takes advantage of the spin echo phenomenon. This technique exposes the target nuclei to a 90 pulse followed by one or more 180 pulses. As the nuclei refocus, a broad distribution indicates the target nuclei is involved in an intermediate excited state. By looking at Relaxation dispersion plots the data collect information on the thermodynamics and kinetics between the excited and ground. Saturation Transfer measures changes in signal from the ground state as excited states become perturbed. It uses weak radio frequency irradiation to saturate the excited state of a particular nuclei which transfers its saturation to the ground state. This signal is amplified by decreasing the magnetization (and the signal) of the ground state.
The main limitations in NMR is that its resolution decreases with proteins that are larger than 25 kDa and is not as detailed as