Haplogroup A-M51 (Y-DNA) on:
[Wikipedia]
[Google]
[Amazon]
A  Each haplogroup originates from, and remains part of, a preceding single haplogroup (or paragroup). As such, any related group of haplogroups may be precisely modelled as a nested hierarchy, in which each set (haplogroup) is also a
Each haplogroup originates from, and remains part of, a preceding single haplogroup (or paragroup). As such, any related group of haplogroups may be precisely modelled as a nested hierarchy, in which each set (haplogroup) is also a
 Human mtDNA haplogroups are lettered:
A,
B,
C,
CZ,
D,
E,
F,
G,
H,
HV,
I,
J,
pre-JT,
JT,
K,
L0,
L1,
L2,
L3,
L4,
L5,
L6,
M,
N,
P,
Q,
R,
R0,
S,
T,
U,
V,
W,
X,
Y, and
Z. The most up-to-date version of the mtDNA tree is maintained by Mannis van Oven on the PhyloTree website.
Mitochondrial Eve is the name given by researchers to the woman who is the most recent common matrilineal (female-lineage) ancestor of all living humans.
Human mtDNA haplogroups are lettered:
A,
B,
C,
CZ,
D,
E,
F,
G,
H,
HV,
I,
J,
pre-JT,
JT,
K,
L0,
L1,
L2,
L3,
L4,
L5,
L6,
M,
N,
P,
Q,
R,
R0,
S,
T,
U,
V,
W,
X,
Y, and
Z. The most up-to-date version of the mtDNA tree is maintained by Mannis van Oven on the PhyloTree website.
Mitochondrial Eve is the name given by researchers to the woman who is the most recent common matrilineal (female-lineage) ancestor of all living humans.
 Haplogroups can be used to define genetic populations and are often geographically oriented. For example, the following are common divisions for mtDNA haplogroups:
* African: L0, L1, L2, L3, L4, L5, L6
* West Eurasian: H, T, U, V, X, K, I, J, W (all listed West Eurasian haplogroups are derived from macro-haplogroup N)
* East Eurasian: A, B, C, D, E, F, G, Y, Z (note: C, D, E, G, and Z belong to macro-haplogroup M)
* Native American: A, B, C, D, X
* Australo-Melanesian: P, Q, S
The mitochondrial haplogroups are divided into three main groups, which are designated by the sequential letters L, M, N.
Humanity first split within the L group between L0 and L1-6. L1-6 gave rise to other L groups, one of which, L3, split into the M and N group.
The M group comprises the first wave of human migration which is thought to have evolved outside of Africa, following an eastward route along southern coastal areas. Descendant lineages of haplogroup M are now found throughout Asia, the Americas, and Melanesia, as well as in parts of the Horn of Africa and North Africa; almost none have been found in Europe. The N haplogroup may represent another macrolineage that evolved outside of Africa, heading northward instead of eastward. Shortly after the migration, the large R group split off from the N.
Haplogroup R consists of two subgroups defined on the basis of their geographical distributions, one found in southeastern Asia and Oceania and the other containing almost all of the modern European populations. Haplogroup N(xR), i.e. mtDNA that belongs to the N group but not to its R subgroup, is typical of Australian aboriginal populations, while also being present at low frequencies among many populations of Eurasia and the Americas.
The L type consists of nearly all Africans.
The M type consists of:
M1 – Ethiopian, Somali and Indian populations. Likely due to much gene flow between the Horn of Africa and the Arabian Peninsula (Saudi Arabia, Yemen, Oman), separated only by a narrow strait between the Red Sea and the Gulf of Aden.
CZ – Many Siberians; branch C – Some Amerindian; branch Z – Many Saami, some Korean, some North Chinese, some Central Asian populations.
D – Some Amerindians, many Siberians and northern East Asians
E – Malay, Borneo, Philippines, Taiwanese aborigines, Papua New Guinea
G – Many Northeast Siberians, northern East Asians, and Central Asians
Q – Melanesian, Polynesian, New Guinean populations
The N type consists of:
A – Found in many Amerindians and some East Asians and Siberians
I – 10% frequency in Northern, Eastern Europe
S – Some Australian aborigines
W – Some Eastern Europeans, South Asians, and southern East Asians
X – Some Amerindians, Southern Siberians, Southwest Asians, and Southern Europeans
Y – Most Nivkhs and people of Nias; many Ainus, Tungusic people, and Austronesians; also found with low frequency in some other populations of Siberia, East Asia, and Central Asia
R – Large group found within the N type. Populations contained therein can be divided geographically into West Eurasia and East Eurasia. Almost all European populations and a large number of Middle-Eastern population today are contained within this branch. A smaller percentage is contained in other N type groups (See above). Below are subclades of R:
B – Some Chinese, Tibetans, Mongolians, Central Asians, Koreans, Amerindians, South Siberians, Japanese, Austronesians
F – Mainly found in southeastern Asia, especially Vietnam; 8.3% in Hvar Island in Croatia.
R0 – Found in Arabia and among Ethiopians and Somalis; branch HV (branch H; branch V) – Europe, Western Asia, North Africa;
Pre-JT – Arose in the Levant (modern Lebanon area), found in 25% frequency in Bedouin populations; branch JT (branch J; branch T) – North, Eastern Europe, Indus, Mediterranean
U – High frequency in West Eurasia, Indian sub-continent, and Algeria, found from India to the Mediterranean and to the rest of Europe; U5 in particular shows high frequency in Scandinavia and Baltic countries with the highest frequency in the
Haplogroups can be used to define genetic populations and are often geographically oriented. For example, the following are common divisions for mtDNA haplogroups:
* African: L0, L1, L2, L3, L4, L5, L6
* West Eurasian: H, T, U, V, X, K, I, J, W (all listed West Eurasian haplogroups are derived from macro-haplogroup N)
* East Eurasian: A, B, C, D, E, F, G, Y, Z (note: C, D, E, G, and Z belong to macro-haplogroup M)
* Native American: A, B, C, D, X
* Australo-Melanesian: P, Q, S
The mitochondrial haplogroups are divided into three main groups, which are designated by the sequential letters L, M, N.
Humanity first split within the L group between L0 and L1-6. L1-6 gave rise to other L groups, one of which, L3, split into the M and N group.
The M group comprises the first wave of human migration which is thought to have evolved outside of Africa, following an eastward route along southern coastal areas. Descendant lineages of haplogroup M are now found throughout Asia, the Americas, and Melanesia, as well as in parts of the Horn of Africa and North Africa; almost none have been found in Europe. The N haplogroup may represent another macrolineage that evolved outside of Africa, heading northward instead of eastward. Shortly after the migration, the large R group split off from the N.
Haplogroup R consists of two subgroups defined on the basis of their geographical distributions, one found in southeastern Asia and Oceania and the other containing almost all of the modern European populations. Haplogroup N(xR), i.e. mtDNA that belongs to the N group but not to its R subgroup, is typical of Australian aboriginal populations, while also being present at low frequencies among many populations of Eurasia and the Americas.
The L type consists of nearly all Africans.
The M type consists of:
M1 – Ethiopian, Somali and Indian populations. Likely due to much gene flow between the Horn of Africa and the Arabian Peninsula (Saudi Arabia, Yemen, Oman), separated only by a narrow strait between the Red Sea and the Gulf of Aden.
CZ – Many Siberians; branch C – Some Amerindian; branch Z – Many Saami, some Korean, some North Chinese, some Central Asian populations.
D – Some Amerindians, many Siberians and northern East Asians
E – Malay, Borneo, Philippines, Taiwanese aborigines, Papua New Guinea
G – Many Northeast Siberians, northern East Asians, and Central Asians
Q – Melanesian, Polynesian, New Guinean populations
The N type consists of:
A – Found in many Amerindians and some East Asians and Siberians
I – 10% frequency in Northern, Eastern Europe
S – Some Australian aborigines
W – Some Eastern Europeans, South Asians, and southern East Asians
X – Some Amerindians, Southern Siberians, Southwest Asians, and Southern Europeans
Y – Most Nivkhs and people of Nias; many Ainus, Tungusic people, and Austronesians; also found with low frequency in some other populations of Siberia, East Asia, and Central Asia
R – Large group found within the N type. Populations contained therein can be divided geographically into West Eurasia and East Eurasia. Almost all European populations and a large number of Middle-Eastern population today are contained within this branch. A smaller percentage is contained in other N type groups (See above). Below are subclades of R:
B – Some Chinese, Tibetans, Mongolians, Central Asians, Koreans, Amerindians, South Siberians, Japanese, Austronesians
F – Mainly found in southeastern Asia, especially Vietnam; 8.3% in Hvar Island in Croatia.
R0 – Found in Arabia and among Ethiopians and Somalis; branch HV (branch H; branch V) – Europe, Western Asia, North Africa;
Pre-JT – Arose in the Levant (modern Lebanon area), found in 25% frequency in Bedouin populations; branch JT (branch J; branch T) – North, Eastern Europe, Indus, Mediterranean
U – High frequency in West Eurasia, Indian sub-continent, and Algeria, found from India to the Mediterranean and to the rest of Europe; U5 in particular shows high frequency in Scandinavia and Baltic countries with the highest frequency in the
The Genographic Project
Y Chromosome Consortium
PhyloTree's Y-tree
A minimal reference phylogeny for the human Y-chromosome
* The Y Chromosome Consortium (2002)
A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups
Genome Research, Vol. 12(2), 339–48, February 2002. (Detailed hierarchica
chart
has conversions from previous naming schemes) * Semino et al. (2000)
The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans
Science, Vol 290 (paper which introduced the "Eu" haplogroups).
Y-DNA Ethnographic and Genographic Atlas and Open-Source Data Compilation
PhyloD3 – D3.js-based phylogenetic tree based on PhyloTree
MitoTool – a web server for the analysis and retrieval of human mitochondrial DNA sequence variations
HaploGrep – automatic classification of mitochondrial DNA haplogroups based on PhyloTree
{{Webarchive, url=https://web.archive.org/web/20160612075145/http://haplogrep.uibk.ac.at/ , date=2016-06-12
HaploFind – fast automatic haplogroup assignment pipeline for human mitochondrial DNA
graphical mtDNA haplogroup skeleton
The Making of the African mtDNA Landscape
Do the Four Clades of the mtDNA Haplogroup L2 Evolve at Different Rates?
Y-DNA Haplogroup Browser
DNA Human evolution Phylogenetics Population genetics Classical genetics
haplotype
A haplotype ( haploid genotype) is a group of alleles in an organism that are inherited together from a single parent.
Many organisms contain genetic material ( DNA) which is inherited from two parents. Normally these organisms have their DNA or ...
is a group of alleles in an organism that are inherited together from a single parent, and a haplogroup (haploid
Ploidy () is the number of complete sets of chromosomes in a cell, and hence the number of possible alleles for autosomal and pseudoautosomal genes. Sets of chromosomes refer to the number of maternal and paternal chromosome copies, respectively ...
from the el, ἁπλοῦς, ''haploûs'', "onefold, simple" and en, group) is a group of similar haplotypes that share a common ancestor with a single-nucleotide polymorphism mutation. More specifically, a haplogroup is a combination of alleles at different chromosomal regions that are closely linked and that tend to be inherited together. As a haplogroup consists of similar haplotypes, it is usually possible to predict a haplogroup from haplotypes. Haplogroups pertain to a single line of descent
In anthropology, kinship is the web of social relationships that form an important part of the lives of all humans in all societies, although its exact meanings even within this discipline are often debated. Anthropologist Robin Fox says that ...
. As such, membership of a haplogroup, by any individual, relies on a relatively small proportion of the genetic material possessed by that individual.
 Each haplogroup originates from, and remains part of, a preceding single haplogroup (or paragroup). As such, any related group of haplogroups may be precisely modelled as a nested hierarchy, in which each set (haplogroup) is also a
Each haplogroup originates from, and remains part of, a preceding single haplogroup (or paragroup). As such, any related group of haplogroups may be precisely modelled as a nested hierarchy, in which each set (haplogroup) is also a subset
In mathematics, Set (mathematics), set ''A'' is a subset of a set ''B'' if all Element (mathematics), elements of ''A'' are also elements of ''B''; ''B'' is then a superset of ''A''. It is possible for ''A'' and ''B'' to be equal; if they are ...
of a single broader set (as opposed, that is, to biparental models, such as human family trees).
Haplogroups are normally identified by an initial letter of the alphabet, and refinements consist of additional number and letter combinations, such as (for example) .
In human genetics, the haplogroups most commonly studied are Y-chromosome (Y-DNA) haplogroups and mitochondrial DNA (mtDNA) haplogroups, each of which can be used to define genetic populations. Y-DNA is passed solely along the patrilineal line, from father to son, while mtDNA is passed down the matrilineal line, from mother to offspring of both sexes. Neither recombines, and thus Y-DNA and mtDNA change only by chance mutation at each generation with no intermixture between parents' genetic material.
Haplogroup formation

Mitochondria
A mitochondrion (; ) is an organelle found in the Cell (biology), cells of most Eukaryotes, such as animals, plants and Fungus, fungi. Mitochondria have a double lipid bilayer, membrane structure and use aerobic respiration to generate adenosi ...
are small organelle
In cell biology, an organelle is a specialized subunit, usually within a cell, that has a specific function. The name ''organelle'' comes from the idea that these structures are parts of cells, as organs are to the body, hence ''organelle,'' the ...
s that lie in the cytoplasm of eukaryotic cells, such as those of humans. Their primary function is to provide energy to the cell. Mitochondria are thought to be reduced descendants of symbiotic
Symbiosis (from Greek , , "living together", from , , "together", and , bíōsis, "living") is any type of a close and long-term biological interaction between two different biological organisms, be it mutualistic, commensalistic, or parasit ...
bacteria that were once free living. One indication that mitochondria were once free living is that each contains a circular DNA, called mitochondrial DNA
Mitochondrial DNA (mtDNA or mDNA) is the DNA located in mitochondria, cellular organelles within eukaryotic cells that convert chemical energy from food into a form that cells can use, such as adenosine triphosphate (ATP). Mitochondrial D ...
(mtDNA), whose structure is more similar to bacteria than eukaryotic organisms (see endosymbiotic theory
Symbiogenesis (endosymbiotic theory, or serial endosymbiotic theory,) is the leading evolutionary theory of the origin of eukaryotic cells from prokaryotic organisms. The theory holds that mitochondria, plastids such as chloroplasts, and possibl ...
). The overwhelming majority of a human's DNA is contained in the chromosomes in the nucleus of the cell, but mtDNA is an exception.
An individual inherits their cytoplasm and the organelles contained by that cytoplasm exclusively from the maternal ovum (egg cell); sperm
Sperm is the male reproductive cell, or gamete, in anisogamous forms of sexual reproduction (forms in which there is a larger, female reproductive cell and a smaller, male one). Animals produce motile sperm with a tail known as a flagellum, whi ...
only pass on the chromosomal DNA, all paternal mitochondria are digested in the oocyte. When a mutation arises in a mtDNA molecule, the mutation is therefore passed in a direct female line of descent. Mutations are copying mistakes in the DNA sequence
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. Th ...
. Single mistakes are called single nucleotide polymorphism
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently larg ...
s (SNPs).
Human Y chromosomes are male-specific sex chromosomes; nearly all humans that possess a Y chromosome will be morphologically male. Although Y chromosomes are situated in the cell nucleus and paired with X chromosomes, they only recombine with the X chromosome at the ends of the Y chromosome; the remaining 95% of the Y chromosome does not recombine. Therefore, the Y chromosome and any mutations that arise in it are passed on from father to son in a direct male line of descent. This means the Y chromosome and mtDNA share specific properties.
Other chromosomes, autosomes
An autosome is any chromosome that is not a sex chromosome. The members of an autosome pair in a diploid cell have the same morphology, unlike those in allosomal (sex chromosome) pairs, which may have different structures. The DNA in autosomes ...
and X chromosomes in women, share their genetic material (called crossing over leading to recombination) during meiosis (a special type of cell division that occurs for the purposes of sexual reproduction). Effectively this means that the genetic material from these chromosomes gets mixed up in every generation, and so any new mutations are passed down randomly from parents to offspring.
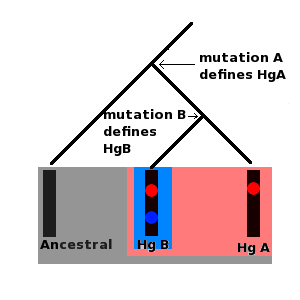
The special feature that both Y chromosomes and mtDNA display is that mutations can accrue along a certain segment of both molecules and these mutations remain fixed in place on the DNA. Furthermore, the historical sequence of these mutations can also be inferred. For example, if a set of ten Y chromosomes (derived from ten different men) contains a mutation, A, but only five of these chromosomes contain a second mutation, B, then it must be the case that mutation B occurred after mutation A.
Furthermore, all ten men who carry the chromosome with mutation A are the direct male line descendants of the same man who was the first person to carry this mutation. The first man to carry mutation B was also a direct male line descendant of this man, but is also the direct male line ancestor of all men carrying mutation B. Series of mutations such as this form molecular lineages. Furthermore, each mutation defines a set of specific Y chromosomes called a haplogroup.
All men carrying mutation A form a single haplogroup, and all men carrying mutation B are part of this haplogroup, but mutation B also defines a more recent haplogroup (which is a subgroup or subclade) of its own to which men carrying only mutation A do not belong. Both mtDNA and Y chromosomes are grouped into lineages and haplogroups; these are often presented as tree like diagrams.
Haplogroup population genetics
It is usually assumed that there is little natural selection for or against a particular haplotype mutation which has survived to the present day, so apart from mutation rates (which may vary from one marker to another) the main driver of population genetics affecting the proportions of haplotypes in a population is genetic drift—random fluctuation caused by the sampling randomness of which members of the population happen to pass their DNA on to members of the next generation of the appropriate sex. This causes the prevalence of a particular marker in a population to continue to fluctuate, until it either hits 100%, or falls out of the population entirely. In a large population with efficient mixing the rate of genetic drift for common alleles is very low; however, in a very small interbreeding population the proportions can change much more quickly. The marked geographical variations and concentrations of particular haplotypes and groups of haplotypes therefore witness the distinctive effects of repeatedpopulation bottleneck
A population bottleneck or genetic bottleneck is a sharp reduction in the size of a population due to environmental events such as famines, earthquakes, floods, fires, disease, and droughts; or human activities such as specicide, widespread violen ...
s or founder events followed by population separations and increases.
The lineages which can be traced back from the present will not reflect the full genetic variation of the older population: genetic drift means that some of the variants will have died out. The cost of full Y-DNA and mtDNA sequence tests has limited the availability of data; however, their cost has dropped dramatically in the last decade. Haplotype coalescence times and current geographical prevalences both carry considerable error uncertainties. This is especially troublesome for coalescence times, because most population geneticists still continue (albeit decreasing a little bit) to use the "Zhivotovski method", which is heavily criticised by DNA-genealogists for its falsehood. The eusocial wasp ''Angiopolybia pallens
''Angiopolybia pallens'' is a species of social wasp predominantly found in South America. The wasp is generally seen in Brazilian rainforests. This species was discovered by Lepeletier in 1836. It typically feeds on nectar and carrion. In fa ...
''presents with 8 haplogroups depending on its location. This displays the idea of genetic drift.
Human Y-chromosome DNA haplogroups
Human Y chromosome DNA (Y-DNA) haplogroups are named from A to T, and are further subdivided using numbers and lower case letters. Y chromosome haplogroup designations are established by the Y Chromosome Consortium. Y-chromosomal Adam is the name given by researchers to the male who is the most recent common patrilineal (male-lineage) ancestor of all living humans. Major Y-chromosome haplogroups, and their geographical regions of occurrence (prior to the recent European colonization), include:Groups without mutation M168
* Haplogroup A (M91) (Africa, especially theKhoisan
Khoisan , or (), according to the contemporary Khoekhoegowab orthography, is a catch-all term for those indigenous peoples of Southern Africa who do not speak one of the Bantu languages, combining the (formerly "Khoikhoi") and the or ( in t ...
and Nilotes)
* Haplogroup B (M60) (Africa, especially the Pygmies and Hadzabe
The Hadza, or Hadzabe (''Wahadzabe'' in Swahili), are a Tanzanian indigenous ethnic group mostly based in southwest Karatu District of Arusha Region. They live around Lake Eyasi in the central Rift Valley and in the neighboring Serengeti Plateau ...
)
Groups with mutation M168
(mutation M168 occurred ~50,000 bp) * Haplogroup C (M130) (Oceania, North/Central/East Asia, North America and a minor presence in South America, Southeast Asia, South Asia, West Asia, and Europe) * YAP+ haplogroups ** Haplogroup DE (M1, M145, M203) ***Haplogroup D Haplogroup D may refer to:
* Haplogroup D (mtDNA), a human mitochondrial DNA (mtDNA) haplogroup
* Haplogroup D (Y-DNA), a human Y-chromosome (Y-DNA) haplogroup
{{Disambiguation ...
(CTS3946) (Tibet, Japan, the Andaman Islands, Central Asia, and a sporadic presence in Nigeria, Syria, and Saudi Arabia)
*** Haplogroup E (M96)
**** Haplogroup E1b1a (V38) West Africa and surrounding regions; formerly known as E3a
**** Haplogroup E1b1b
E-M215, also known as E1b1b and formerly E3b, is a major human Y-chromosome DNA haplogroup. It is a division of the macro-haplogroup E-M96, which is defined by the single-nucleotide polymorphism (SNP) mutation M215. In other words, it is one of ...
(M215) Associated with the spread of Afroasiatic languages; now concentrated in North Africa and the Horn of Africa, as well as parts of the Middle East, the Mediterranean, and the Balkans; formerly known as E3b
Groups with mutation M89
(mutation M89 occurred ~45,000 bp) * Haplogroup F (M89) Oceania, Europe, Asia, North and South America ** Haplogroup FT (P14, M213) (China, Vietnam, Singapore) *Haplogroup G Haplogroup G may refer to:
* Haplogroup G (mtDNA)
In human mitochondrial genetics, Haplogroup G is a human mitochondrial DNA (mtDNA) haplogroup.
Origin
Haplogroup G is a descendant of haplogroup M. Haplogroup G is divided into subclades G1, G2, ...
(M201) (present among many ethnic groups in Eurasia, usually at low frequency; most common in the Caucasus, the Iranian plateau, and Anatolia; in Europe mainly in Greece, Italy, Iberia, the Tyrol, Bohemia; rare in Northern Europe)
* Haplogroup H (L901/M2939)
**H1'3 (Z4221/M2826, Z13960)
***H1 (L902/M3061)
****H1a (M69/Page45) India, Sri Lanka, Nepal, Pakistan, Iran, Central Asia
****H1b (B108) Found in a Burmese
Burmese may refer to:
* Something of, from, or related to Myanmar, a country in Southeast Asia
* Burmese people
* Burmese language
* Burmese alphabet
* Burmese cuisine
* Burmese culture
Animals
* Burmese cat
* Burmese chicken
* Burmese (hor ...
individual in Myanmar
Myanmar, ; UK pronunciations: US pronunciations incl. . Note: Wikipedia's IPA conventions require indicating /r/ even in British English although only some British English speakers pronounce r at the end of syllables. As John C. Wells, Joh ...
.
***H3 (Z5857) India, Sri Lanka, Pakistan, Bahrain, Qatar
**H2 (P96) Formerly known as haplogroup F3. Found with low frequency in Europe and western Asia.
* Haplogroup IJK (L15, L16)
Groups with mutations L15 & L16
* Haplogroup IJK (L15, L16) ** Haplogroup IJ (S2, S22) *** Haplogroup I (M170, P19, M258) (widespread in Europe, found infrequently in parts of the Middle East, and virtually absent elsewhere) **** Haplogroup I1 (M253, M307, P30, P40) (Northern Europe, dominant in Scandinavia) **** Haplogroup I2 (S31) (Central and Southeast Europe, Sardinia, Balkans) *** Haplogroup J (M304) (the Middle East, Turkey, Caucasus, Italy, Greece, the Balkans, North Africa) **** Haplogroup J* (Mainly found in Socotra, with a few observations in Pakistan, Oman, Greece, the Czech Republic, and among Turkic peoples) **** Haplogroup J1 (M267) (Mostly associated with Semitic peoples in the Middle East but also found in; Mediterranean Europe, Ethiopia, North Africa, Iran, Pakistan, India and with Northeast Caucasian peoples inDagestan
Dagestan ( ; rus, Дагеста́н, , dəɡʲɪˈstan, links=yes), officially the Republic of Dagestan (russian: Респу́блика Дагеста́н, Respúblika Dagestán, links=no), is a republic of Russia situated in the North C ...
; J1 with DYS388=13 is associated with eastern Anatolia)
**** Haplogroup J2 (M172) (Mainly found in West Asia, Central Asia, Southern Europe, and North Africa)
** Haplogroup K (M9, P128, P131, P132)
Groups with mutation M9
(mutation M9 occurred ~40,000 bp) * Haplogroup K ** Haplogroup LT (L298/P326) *** Haplogroup L (M11, M20, M22, M61, M185, M295) (South Asia, Central Asia, Southwestern Asia, the Mediterranean) *** Haplogroup T (M70, M184/USP9Y+3178, M193, M272) (North Africa, Horn of Africa, Southwest Asia, the Mediterranean, South Asia); formerly known as Haplogroup K2 ** Haplogroup K(xLT) (rs2033003/M526)= Groups with mutation M526
= ** Haplogroup M (P256) (New Guinea, Melanesia, eastern Indonesia) ** Haplogroup NO (M214) *** Haplogroup N (M231) (northernmost Eurasia) *** Haplogroup O (M175) (East Asia, Southeast Asia, the South Pacific, South Asia, Central Asia) **** Haplogroup O1 (F265) ***** Haplogroup O1a (M119) ***** Haplogroup O1b (P31, M268) **** Haplogroup O2 (M122) ** Haplogroup P-M45 (M45) (M45 occurred ~35,000 bp) *** Haplogroup Q-M242 (M242) (Occurred ~15,000–20,000 bp. Found in Asia and the Americas) **** Haplogroup Q-M3 (M3) (North America, Central America, and South America) *** Haplogroup R (M207) **** Haplogroup R1 (M173) ***** Haplogroup R1a (M17) (Central Asia, South Asia, and Central, Northern, and Eastern Europe) *****Haplogroup R1b
Haplogroup R1b (R-M343), previously known as Hg1 and Eu18, is a human Y-chromosome haplogroup.
It is the most frequently occurring paternal lineage in Western Europe, as well as some parts of Russia (e.g. the Bashkirs) and pockets of Central A ...
(M343) (Europe, Caucasus, Central Asia, South Asia, North Africa, Central Africa)
**** Haplogroup R2 (M124) (South Asia, Caucasus, Central Asia)
** Haplogroup S (M230, P202, P204) (New Guinea, Melanesia, eastern Indonesia)
Human mitochondrial DNA haplogroups
Defining populations
 Haplogroups can be used to define genetic populations and are often geographically oriented. For example, the following are common divisions for mtDNA haplogroups:
* African: L0, L1, L2, L3, L4, L5, L6
* West Eurasian: H, T, U, V, X, K, I, J, W (all listed West Eurasian haplogroups are derived from macro-haplogroup N)
* East Eurasian: A, B, C, D, E, F, G, Y, Z (note: C, D, E, G, and Z belong to macro-haplogroup M)
* Native American: A, B, C, D, X
* Australo-Melanesian: P, Q, S
The mitochondrial haplogroups are divided into three main groups, which are designated by the sequential letters L, M, N.
Humanity first split within the L group between L0 and L1-6. L1-6 gave rise to other L groups, one of which, L3, split into the M and N group.
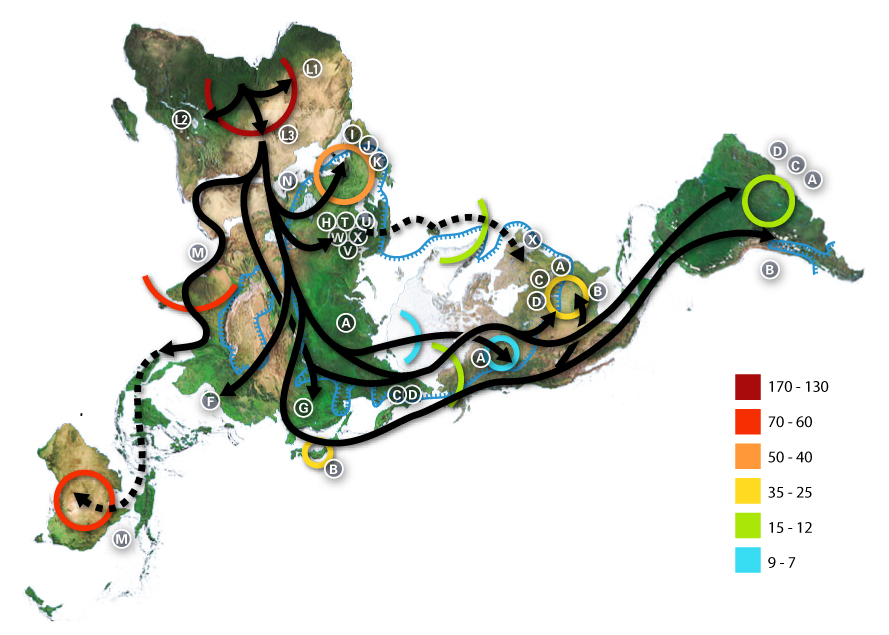
The M group comprises the first wave of human migration which is thought to have evolved outside of Africa, following an eastward route along southern coastal areas. Descendant lineages of haplogroup M are now found throughout Asia, the Americas, and Melanesia, as well as in parts of the Horn of Africa and North Africa; almost none have been found in Europe. The N haplogroup may represent another macrolineage that evolved outside of Africa, heading northward instead of eastward. Shortly after the migration, the large R group split off from the N.
Haplogroup R consists of two subgroups defined on the basis of their geographical distributions, one found in southeastern Asia and Oceania and the other containing almost all of the modern European populations. Haplogroup N(xR), i.e. mtDNA that belongs to the N group but not to its R subgroup, is typical of Australian aboriginal populations, while also being present at low frequencies among many populations of Eurasia and the Americas.
The L type consists of nearly all Africans.
The M type consists of:
M1 – Ethiopian, Somali and Indian populations. Likely due to much gene flow between the Horn of Africa and the Arabian Peninsula (Saudi Arabia, Yemen, Oman), separated only by a narrow strait between the Red Sea and the Gulf of Aden.
CZ – Many Siberians; branch C – Some Amerindian; branch Z – Many Saami, some Korean, some North Chinese, some Central Asian populations.
D – Some Amerindians, many Siberians and northern East Asians
E – Malay, Borneo, Philippines, Taiwanese aborigines, Papua New Guinea
G – Many Northeast Siberians, northern East Asians, and Central Asians
Q – Melanesian, Polynesian, New Guinean populations
The N type consists of:
A – Found in many Amerindians and some East Asians and Siberians
I – 10% frequency in Northern, Eastern Europe
S – Some Australian aborigines
W – Some Eastern Europeans, South Asians, and southern East Asians
X – Some Amerindians, Southern Siberians, Southwest Asians, and Southern Europeans
Y – Most Nivkhs and people of Nias; many Ainus, Tungusic people, and Austronesians; also found with low frequency in some other populations of Siberia, East Asia, and Central Asia
R – Large group found within the N type. Populations contained therein can be divided geographically into West Eurasia and East Eurasia. Almost all European populations and a large number of Middle-Eastern population today are contained within this branch. A smaller percentage is contained in other N type groups (See above). Below are subclades of R:
B – Some Chinese, Tibetans, Mongolians, Central Asians, Koreans, Amerindians, South Siberians, Japanese, Austronesians
F – Mainly found in southeastern Asia, especially Vietnam; 8.3% in Hvar Island in Croatia.
R0 – Found in Arabia and among Ethiopians and Somalis; branch HV (branch H; branch V) – Europe, Western Asia, North Africa;
Pre-JT – Arose in the Levant (modern Lebanon area), found in 25% frequency in Bedouin populations; branch JT (branch J; branch T) – North, Eastern Europe, Indus, Mediterranean
U – High frequency in West Eurasia, Indian sub-continent, and Algeria, found from India to the Mediterranean and to the rest of Europe; U5 in particular shows high frequency in Scandinavia and Baltic countries with the highest frequency in the
Haplogroups can be used to define genetic populations and are often geographically oriented. For example, the following are common divisions for mtDNA haplogroups:
* African: L0, L1, L2, L3, L4, L5, L6
* West Eurasian: H, T, U, V, X, K, I, J, W (all listed West Eurasian haplogroups are derived from macro-haplogroup N)
* East Eurasian: A, B, C, D, E, F, G, Y, Z (note: C, D, E, G, and Z belong to macro-haplogroup M)
* Native American: A, B, C, D, X
* Australo-Melanesian: P, Q, S
The mitochondrial haplogroups are divided into three main groups, which are designated by the sequential letters L, M, N.
Humanity first split within the L group between L0 and L1-6. L1-6 gave rise to other L groups, one of which, L3, split into the M and N group.
The M group comprises the first wave of human migration which is thought to have evolved outside of Africa, following an eastward route along southern coastal areas. Descendant lineages of haplogroup M are now found throughout Asia, the Americas, and Melanesia, as well as in parts of the Horn of Africa and North Africa; almost none have been found in Europe. The N haplogroup may represent another macrolineage that evolved outside of Africa, heading northward instead of eastward. Shortly after the migration, the large R group split off from the N.
Haplogroup R consists of two subgroups defined on the basis of their geographical distributions, one found in southeastern Asia and Oceania and the other containing almost all of the modern European populations. Haplogroup N(xR), i.e. mtDNA that belongs to the N group but not to its R subgroup, is typical of Australian aboriginal populations, while also being present at low frequencies among many populations of Eurasia and the Americas.
The L type consists of nearly all Africans.
The M type consists of:
M1 – Ethiopian, Somali and Indian populations. Likely due to much gene flow between the Horn of Africa and the Arabian Peninsula (Saudi Arabia, Yemen, Oman), separated only by a narrow strait between the Red Sea and the Gulf of Aden.
CZ – Many Siberians; branch C – Some Amerindian; branch Z – Many Saami, some Korean, some North Chinese, some Central Asian populations.
D – Some Amerindians, many Siberians and northern East Asians
E – Malay, Borneo, Philippines, Taiwanese aborigines, Papua New Guinea
G – Many Northeast Siberians, northern East Asians, and Central Asians
Q – Melanesian, Polynesian, New Guinean populations
The N type consists of:
A – Found in many Amerindians and some East Asians and Siberians
I – 10% frequency in Northern, Eastern Europe
S – Some Australian aborigines
W – Some Eastern Europeans, South Asians, and southern East Asians
X – Some Amerindians, Southern Siberians, Southwest Asians, and Southern Europeans
Y – Most Nivkhs and people of Nias; many Ainus, Tungusic people, and Austronesians; also found with low frequency in some other populations of Siberia, East Asia, and Central Asia
R – Large group found within the N type. Populations contained therein can be divided geographically into West Eurasia and East Eurasia. Almost all European populations and a large number of Middle-Eastern population today are contained within this branch. A smaller percentage is contained in other N type groups (See above). Below are subclades of R:
B – Some Chinese, Tibetans, Mongolians, Central Asians, Koreans, Amerindians, South Siberians, Japanese, Austronesians
F – Mainly found in southeastern Asia, especially Vietnam; 8.3% in Hvar Island in Croatia.
R0 – Found in Arabia and among Ethiopians and Somalis; branch HV (branch H; branch V) – Europe, Western Asia, North Africa;
Pre-JT – Arose in the Levant (modern Lebanon area), found in 25% frequency in Bedouin populations; branch JT (branch J; branch T) – North, Eastern Europe, Indus, Mediterranean
U – High frequency in West Eurasia, Indian sub-continent, and Algeria, found from India to the Mediterranean and to the rest of Europe; U5 in particular shows high frequency in Scandinavia and Baltic countries with the highest frequency in the Sami people
Acronyms
* SAMI, ''Synchronized Accessible Media Interchange'', a closed-captioning format developed by Microsoft
* Saudi Arabian Military Industries, a government-owned defence company
* South African Malaria Initiative, a virtual expertise net ...
.
Y-chromosome and MtDNA geographic haplogroup assignation
Here is a list of Y-chromosome and MtDNA geographic haplogroup assignation proposed by Bekada et al. 2013.Y-chromosome
According to SNPS haplogroups which are the age of the first extinction event tend to be around 45–50 kya. Haplogroups of the second extinction event seemed to diverge 32–35 kya according to Mal'ta. The ground zero extinction event appears to be Toba during which haplogroup CDEF* appeared to diverge into C, DE and F. C and F have almost nothing in common while D and E have plenty in common. Extinction event #1 according to current estimates occurred after Toba, although older ancient DNA could push the ground zero extinction event to long before Toba, and push the first extinction event here back to Toba. Haplogroups with extinction event notes by them have a dubious origin and this is because extinction events lead to severe bottlenecks, so all notes by these groups are just guesses. Note that the SNP counting of ancient DNA can be highly variable meaning that even though all these groups diverged around the same time no one knows when.mtDNA
See also
* International HapMap Project * Molecular evolution * Molecular phylogenetics * Human evolutionary genetics * Race (biology) / Race (human categorization) *Genetic genealogy
Genetic genealogy is the use of genealogical DNA tests, i.e., DNA profiling and DNA testing, in combination with traditional genealogical methods, to infer genetic relationships between individuals. This application of genetics came to be used b ...
* Genealogical DNA test
* List of genetic genealogy topics
* List of haplogroups of notable people
This is a list of haplogroups of historic people. Haplogroups can be determined from the remains of historical figures, or derived from genealogical DNA tests of people who trace their direct maternal or paternal ancestry to a noted historical fi ...
References
External links
General
The Genographic Project
all DNA haplogroups
Y-Chromosome *https://web.archive.org/web/20040728005528/http://www.scs.uiuc.edu/~mcdonald/WorldHaplogroupsMaps.pdfY chromosome DNA haplogroups
Y Chromosome Consortium
PhyloTree's Y-tree
A minimal reference phylogeny for the human Y-chromosome
* The Y Chromosome Consortium (2002)
A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups
Genome Research, Vol. 12(2), 339–48, February 2002. (Detailed hierarchica
chart
has conversions from previous naming schemes) * Semino et al. (2000)
The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans
Science, Vol 290 (paper which introduced the "Eu" haplogroups).
Y-DNA Ethnographic and Genographic Atlas and Open-Source Data Compilation
Mitochondrial DNA haplogroups
* PhyloTree – The phylogenetic tree of global human mitochondrial DNA variationPhyloD3 – D3.js-based phylogenetic tree based on PhyloTree
MitoTool – a web server for the analysis and retrieval of human mitochondrial DNA sequence variations
HaploGrep – automatic classification of mitochondrial DNA haplogroups based on PhyloTree
{{Webarchive, url=https://web.archive.org/web/20160612075145/http://haplogrep.uibk.ac.at/ , date=2016-06-12
HaploFind – fast automatic haplogroup assignment pipeline for human mitochondrial DNA
graphical mtDNA haplogroup skeleton
The Making of the African mtDNA Landscape
Do the Four Clades of the mtDNA Haplogroup L2 Evolve at Different Rates?
Software
Y-DNA Haplogroup Browser
DNA Human evolution Phylogenetics Population genetics Classical genetics