|
SnRNA
Small nuclear RNA (snRNA) is a class of small RNA molecules that are found within the splicing speckles and Cajal bodies of the cell nucleus in eukaryotic cells. The length of an average snRNA is approximately 150 nucleotides. They are transcribed by either RNA polymerase II or RNA polymerase III. Their primary function is in the processing of pre- messenger RNA ( hnRNA) in the nucleus. They have also been shown to aid in the regulation of transcription factors ( 7SK RNA) or RNA polymerase II (B2 RNA), and maintaining the telomeres. snRNA are always associated with a set of specific proteins, and the complexes are referred to as small nuclear ribonucleoproteins (snRNP, often pronounced "snurps"). Each snRNP particle is composed of a snRNA component and several snRNP-specific proteins (including Sm proteins, a family of nuclear proteins). The most common human snRNA components of these complexes are known, respectively, as: U1 spliceosomal RNA, U2 spliceosomal RNA, U4 splic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U4 Spliceosomal RNA
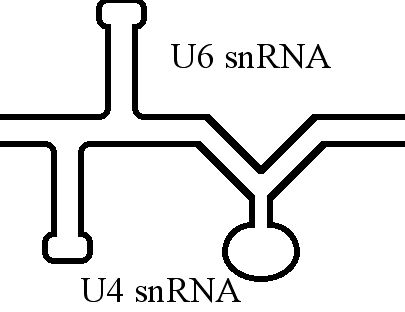
The U4 small nuclear Ribo-Nucleic Acid (U4 snRNA) is a non-coding RNA component of the major or U2-dependent spliceosome – a eukaryotic molecular machine involved in the splicing of pre-messenger RNA (pre-mRNA). It forms a duplex with U6, and with each splicing round, it is displaced from the U6 snRNA (and the spliceosome) in an ATP-dependent manner, allowing U6 to re-fold and create the active site for splicing catalysis. A recycling process involving protein Brr2 releases U4 from U6, while protein Prp24 re-anneals U4 and U6. The crystal structure of a 5′ stem-loop of U4 in complex with a binding protein has been solved. Biological role The U4 snRNA has been shown to exist in a number of different formats including: bound to proteins as a small nuclear Ribo-Nuclear Protein snRNP, involved with the U6 snRNA in the di-snRNP, as well as involved with both the U6 snRNA and the U5 snRNA in the tri-snRNP. The different formats have been proposed to coincide with different te ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SnRNP
snRNPs (pronounced "snurps"), or small nuclear ribonucleoproteins, are RNA-protein complexes that combine with unmodified pre-mRNA and various other proteins to form a spliceosome, a large RNA-protein molecular complex upon which splicing of pre-mRNA occurs. The action of snRNPs is essential to the removal of introns from pre-mRNA, a critical aspect of post-transcriptional modification of RNA, occurring only in the nucleus of eukaryotic cells. Additionally, '' U7 snRNP'' is not involved in splicing at all, as U7 snRNP is responsible for processing the 3′ stem-loop of histone pre-mRNA. The two essential components of snRNPs are protein molecules and RNA. The RNA found within each snRNP particle is known as ''small nuclear RNA'', or snRNA, and is usually about 150 nucleotides in length. The snRNA component of the snRNP gives specificity to individual introns by " recognizing" the sequences of critical splicing signals at the 5' and 3' ends and branch site of introns. The ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U7 Small Nuclear RNA
The U7 small nuclear RNA (U7 snRNA) is an RNA molecule and a component of the small nuclear ribonucleoprotein complex (U7 snRNP). The U7 snRNA is required for histone pre-mRNA processing. The 5' end of the U7 snRNA binds the HDE (histone downstream element), a conserved purine-rich region, located 15 nucleotides downstream the histone mRNA cleavage site. The binding of the HDE region by the U7 snRNA, through complementary base-pairing, is an important step for the future recruitment of cleavage factors during histone pre-mRNA processing. See also *Duchenne muscular dystrophy *Histone 3' UTR stem-loop In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wr ... * LSM10 References Further reading External ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U6 Spliceosomal RNA
U6 snRNA is the non-coding small nuclear RNA (snRNA) component of U6 snRNP (''small nuclear ribonucleoprotein''), an RNA-protein complex that combines with other snRNPs, unmodified pre-mRNA, and various other proteins to assemble a spliceosome, a large RNA-protein molecular complex that catalyzes the excision of introns from pre-mRNA. Splicing, or the removal of introns, is a major aspect of post-transcriptional modification and takes place only in the nucleus of eukaryotes. The RNA sequence of U6 is the most highly conserved across species of all five of the snRNAs involved in the spliceosome, suggesting that the function of the U6 snRNA has remained both crucial and unchanged through evolution. It is common in vertebrate genomes to find many copies of the U6 snRNA gene or U6-derived pseudogenes. This prevalence of "back-ups" of the U6 snRNA gene in vertebrates further implies its evolutionary importance to organism viability. The U6 snRNA gene has been isolated in many organis ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U2 Spliceosomal RNA
U2 spliceosomal snRNAs are a species of small nuclear RNA ( snRNA) molecules found in the major spliceosomal (Sm) machinery of virtually all eukaryotic organisms. ''In vivo'', U2 snRNA along with its associated polypeptides assemble to produce the U2 small nuclear ribonucleoprotein (snRNP), an essential component of the major spliceosomal complex. The major spliceosomal-splicing pathway is occasionally referred to as U2 dependent, based on a class of Sm intron—found in mRNA primary transcripts—that are recognized exclusively by the U2 snRNP during early stages of spliceosomal assembly. In addition to U2 dependent intron recognition, U2 snRNA has been theorized to serve a catalytic role in the chemistry of pre-RNA splicing as well. Similar to ribosomal RNAs ( rRNAs), Sm snRNAs must mediate both RNA:RNA and RNA:protein contacts and hence have evolved specialized, highly conserved, primary and secondary structural elements to facilitate these types of interactions. Shortly after ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Spliceosome
A spliceosome is a large ribonucleoprotein (RNP) complex found primarily within the nucleus of eukaryotic cells. The spliceosome is assembled from small nuclear RNAs ( snRNA) and numerous proteins. Small nuclear RNA (snRNA) molecules bind to specific proteins to form a small nuclear ribonucleoprotein complex (snRNP, pronounced “snurps”), which in turn combines with other snRNPs to form a large ribonucleoprotein complex called a spliceosome. The spliceosome removes introns from a transcribed pre-mRNA, a type of primary transcript. This process is generally referred to as splicing. An analogy is a film editor, who selectively cuts out irrelevant or incorrect material (equivalent to the introns) from the initial film and sends the cleaned-up version to the director for the final cut. However, sometimes the RNA within the intron acts as a ribozyme, splicing itself without the use of a spliceosome or protein enzymes. History In 1977, work by the Sharp and Roberts labs ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U11 Spliceosomal RNA
The U11 snRNA ( small nuclear ribonucleic acid) is an important non-coding RNA in the minor spliceosome protein complex, which activates the alternative splicing mechanism. The minor spliceosome is associated with similar protein components as the major spliceosome. It uses U11 snRNA to recognize the 5' splice site (functionally equivalent to U1 snRNA) while U12 snRNA binds to the branchpoint to recognize the 3' splice site (functionally equivalent to U2 snRNA). Secondary structure U11 snRNA has a stem-loop structure with a 5' end as splice site sequence (5' ss) and contains four stem loops structures (I-IV). A structural comparison of U11 snRNA between plants, vertebrates and insects shows that it is folded into a structure with a four-way junction at the 5' site and in a stem loop structure at the 3' site. Binding site during assembly pathway The 5' splice site region possesses sequence complementarity with the 5' splice site of the eukaryotic U12 type pre-mRNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U4atac Minor Spliceosomal RNA
U4atac minor spliceosomal RNA is a ncRNA which is an essential component of the minor U12-type spliceosome complex. The U12-type spliceosome is required for removal of the rarer class of eukaryotic introns (AT-AC, U12-type). U4atac snRNA is proposed to form a base-paired complex with another spliceosomal RNA U6atac via two stem loop regions. These interacting stem loops have been shown to be required for in vivo splicing. U4atac also contains a 3' Sm protein binding site which has been shown to be essential for splicing activity. U4atac is the functional analog of U4 spliceosomal RNA in the major U2-type spliceosomal complex. The Drosophila ''Drosophila'' () is a genus of flies, belonging to the family Drosophilidae, whose members are often called "small fruit flies" or (less frequently) pomace flies, vinegar flies, or wine flies, a reference to the characteristic of many species ... U4atac snRNA has an additional predicted 3' stem loop terminal to the Sm binding site ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U1 Spliceosomal RNA
U1 spliceosomal RNA is the small nuclear RNA (snRNA) component of U1 snRNP (''small nuclear ribonucleoprotein''), an RNA-protein complex that combines with other snRNPs, unmodified pre-mRNA, and various other proteins to assemble a spliceosome, a large RNA-protein molecular complex upon which splicing of pre-mRNA occurs. Splicing, or the removal of introns, is a major aspect of post-transcriptional modification, and takes place only in the nucleus of eukaryotes. Structure and function In humans, the U1 spliceosomal RNA is 164 bases long, forms four stem-loops, and possesses a 5'-trimethylguanosine five-prime cap. Bases 3 to 10 are a conserved sequence that base-pairs with the 5' splice site of introns during RNA splicing, and bases 126 to 133 form the Sm site, around which the Sm ring is assembled. Stem-loop I binds to the U1-70K protein, stem-loop II binds to the U1 A protein, stem-loops III and IV bind to the core RNP domain, a heteroheptameric Sm ring consisting of SmB/B', ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U5 Spliceosomal RNA
U5 snRNA is a small nuclear RNA (snRNA) that participates in RNA splicing as a component of the spliceosome. It forms the U5 snRNP (''small nuclear ribonucleoprotein'') by associating with several proteins including Prp8 - the largest and most conserved protein in the spliceosome, Brr2 - a helicase Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separat ... required for spliceosome activation, Snu114, and the 7 Sm proteins. U5 snRNA forms a coaxially-stacked series of helices that project into the active site of the spliceosome. Loop 1, which caps this series of helices, forms 4-5 base pairs with the 5'-exon during the two chemical reactions of splicing. This interaction appears to be especially important during step two of splicing, exon ligation. References Further reading * * ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Polymerase III
In eukaryote cells, RNA polymerase III (also called Pol III) is a protein that transcribes DNA to synthesize ribosomal 5S rRNA, tRNA and other small RNAs. The genes transcribed by RNA Pol III fall in the category of "housekeeping" genes whose expression is required in all cell types and most environmental conditions. Therefore, the regulation of Pol III transcription is primarily tied to the regulation of cell growth and the cell cycle, thus requiring fewer regulatory proteins than RNA polymerase II. Under stress conditions however, the protein Maf1 represses Pol III activity. Rapamycin is another Pol III inhibitor via its direct target TOR. Transcription The process of transcription (by any polymerase) involves three main stages: *Initiation, requiring construction of the RNA polymerase complex on the gene's promoter *Elongation, the synthesis of the RNA transcript *Termination, the finishing of RNA transcription and disassembly of the RNA polymerase complex Initiati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
U12 Minor Spliceosomal RNA
U12 minor spliceosomal RNA is formed from U12 small nuclear (snRNA), together with U4atac/ U6atac, U5, and U11 snRNAs and associated proteins, forms a spliceosome that cleaves a divergent class of low-abundance pre-mRNA introns. Although the U12 sequence is very divergent from that of U2, the two are functionally analogous. Structure The predicted secondary structure of U12 RNA is published,. However, the alternative single hairpin in the 3' end shown here seems to better match the alignment of divergent ''Drosophila melanogaster'' and ''Arabidopsis thaliana'' sequences. The sequences U12 introns that are spliced out are collected in a biological database called the U12 intron database U12 Intron Database (U12DB) is a biological database of containing the sequence of eukaryotic introns that are spliced out by a specialised minor spliceosome that contains U12 minor spliceosomal RNA U12 minor spliceosomal RNA is formed from U12 sm .... References External links * S ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |