|
Sequence Assembly
In bioinformatics, sequence assembly refers to aligning and merging fragments from a longer DNA sequence in order to reconstruct the original sequence. This is needed as DNA sequencing technology might not be able to 'read' whole genomes in one go, but rather reads small pieces of between 20 and 30,000 bases, depending on the technology used. Typically, the short fragments (reads) result from shotgun sequencing genomic DNA, or gene transcript ( ESTs). The problem of sequence assembly can be compared to taking many copies of a book, passing each of them through a shredder with a different cutter, and piecing the text of the book back together just by looking at the shredded pieces. Besides the obvious difficulty of this task, there are some extra practical issues: the original may have many repeated paragraphs, and some shreds may be modified during shredding to have typos. Excerpts from another book may also be added in, and some shreds may be completely unrecognizable. Gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bioinformatics
Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combines biology, chemistry, physics, computer science, information engineering, mathematics and statistics to analyze and interpret the biological data. Bioinformatics has been used for '' in silico'' analyses of biological queries using computational and statistical techniques. Bioinformatics includes biological studies that use computer programming as part of their methodology, as well as specific analysis "pipelines" that are repeatedly used, particularly in the field of genomics. Common uses of bioinformatics include the identification of candidates genes and single nucleotide polymorphisms ( SNPs). Often, such identification is made with the aim to better understand the genetic basis of disease, unique adaptations, desirable propertie ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Open-source Software
Open-source software (OSS) is computer software that is released under a license in which the copyright holder grants users the rights to use, study, change, and distribute the software and its source code to anyone and for any purpose. Open-source software may be developed in a collaborative public manner. Open-source software is a prominent example of open collaboration, meaning any capable user is able to participate online in development, making the number of possible contributors indefinite. The ability to examine the code facilitates public trust in the software. Open-source software development can bring in diverse perspectives beyond those of a single company. A 2008 report by the Standish Group stated that adoption of open-source software models has resulted in savings of about $60 billion per year for consumers. Open source code can be used for studying and allows capable end users to adapt software to their personal needs in a similar way user scripts ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
K-mer
In bioinformatics, ''k''-mers are substrings of length k contained within a biological sequence. Primarily used within the context of computational genomics and sequence analysis, in which ''k''-mers are composed of nucleotides (''i.e''. A, T, G, and C), ''k''-mers are capitalized upon to assemble DNA sequences, improve heterologous gene expression, identify species in metagenomic samples, and create attenuated vaccines. Usually, the term ''k''-mer refers to all of a sequence's subsequences of length k, such that the sequence AGAT would have four monomers (A, G, A, and T), three 2-mers (AG, GA, AT), two 3-mers (AGA and GAT) and one 4-mer (AGAT). More generally, a sequence of length L will have L - k + 1 ''k''-mers and n^ total possible ''k''-mers, where n is number of possible monomers (e.g. four in the case of DNA). Introduction ''k''-mers are simply length k subsequences. For example, all the possible ''k''-mers of a DNA sequence are shown below: A method of visual ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
De Novo Sequence Assemblers
De novo sequence assemblers are a type of program that assembles short nucleotide sequences into longer ones without the use of a reference genome. These are most commonly used in bioinformatic studies to assemble genomes or transcriptomes. Two common types of de novo assemblers are greedy algorithm assemblers and De Bruijn graph assemblers. Types of de novo assemblers There are two types of algorithms that are commonly utilized by these assemblers: greedy, which aim for local optima, and graph method algorithms, which aim for global optima. Different assemblers are tailored for particular needs, such as the assembly of (small) bacterial genomes, (large) eukaryotic genomes, or transcriptomes. Greedy algorithm assemblers are assemblers that find local optima in alignments of smaller reads. Greedy algorithm assemblers typically feature several steps: 1) pairwise distance calculation of reads, 2) clustering of reads with greatest overlap, 3) assembly of overlapping reads into la ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Types Of Sequencing Assembly
Type may refer to: Science and technology Computing * Typing, producing text via a keyboard, typewriter, etc. * Data type, collection of values used for computations. * File type * TYPE (DOS command), a command to display contents of a file. * Type (Unix), a command in POSIX shells that gives information about commands. * Type safety, the extent to which a programming language discourages or prevents type errors. * Type system, defines a programming language's response to data types. Mathematics * Type (model theory) * Type theory, basis for the study of type systems * Arity or type, the number of operands a function takes * Type, any proposition or set in the intuitionistic type theory * Type, of an entire function ** Exponential type Biology * Type (biology), which fixes a scientific name to a taxon * Dog type, categorization by use or function of domestic dogs Lettering * Type is a design concept for lettering used in typography which helped bring about modern textual printin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
De Novo Transcriptome Assembly
''De novo'' transcriptome assembly is the de novo sequence assembly method of creating a transcriptome without the aid of a reference genome. Introduction As a result of the development of novel sequencing technologies, the years between 2008 and 2012 saw a large drop in the cost of sequencing. Per megabase and genome, the cost dropped to 1/100,000th and 1/10,000th of the price, respectively. Prior to this, only transcriptomes of organisms that were of broad interest and utility to scientific research were sequenced; however, these developed in 2010s high-throughput sequencing (also called next-generation sequencing) technologies are both cost- and labor- effective, and the range of organisms studied via these methods is expanding. Transcriptomes have subsequently been created for chickpea, planarians, '' Parhyale hawaiensis'', as well as the brains of the Nile crocodile, the corn snake, the bearded dragon, and the red-eared slider, to name just a few. Examining non-model organ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA-Seq
RNA-Seq (named as an abbreviation of RNA sequencing) is a sequencing technique which uses next-generation sequencing (NGS) to reveal the presence and quantity of RNA in a biological sample at a given moment, analyzing the continuously changing cellular transcriptome. Specifically, RNA-Seq facilitates the ability to look at alternative gene spliced transcripts, post-transcriptional modifications, gene fusion, mutations/ SNPs and changes in gene expression over time, or differences in gene expression in different groups or treatments. In addition to mRNA transcripts, RNA-Seq can look at different populations of RNA to include total RNA, small RNA, such as miRNA, tRNA, and ribosomal profiling. RNA-Seq can also be used to determine exon/intron boundaries and verify or amend previously annotated 5' and 3' gene boundaries. Recent advances in RNA-Seq include single cell sequencing, in situ sequencing of fixed tissue, and native RNA molecule sequencing with single-molecule real- ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Post-transcriptional Modification
Transcriptional modification or co-transcriptional modification is a set of biological processes common to most eukaryotic cells by which an RNA primary transcript is chemically altered following transcription from a gene to produce a mature, functional RNA molecule that can then leave the nucleus and perform any of a variety of different functions in the cell. There are many types of post-transcriptional modifications achieved through a diverse class of molecular mechanisms. One example is the conversion of precursor messenger RNA transcripts into mature messenger RNA that is subsequently capable of being translated into protein. This process includes three major steps that significantly modify the chemical structure of the RNA molecule: the addition of a 5' cap, the addition of a 3' polyadenylated tail, and RNA splicing. Such processing is vital for the correct translation of eukaryotic genomes because the initial precursor mRNA produced by transcription often contains bo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single-nucleotide Polymorphism
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently large fraction of the population (e.g. 1% or more), many publications do not apply such a frequency threshold. For example, at a specific base position in the human genome, the G nucleotide may appear in most individuals, but in a minority of individuals, the position is occupied by an A. This means that there is a SNP at this specific position, and the two possible nucleotide variations – G or A – are said to be the alleles for this specific position. SNPs pinpoint differences in our susceptibility to a wide range of diseases, for example age-related macular degeneration (a common SNP in the CFH gene is associated with increased risk of the disease) or nonalcoholic fatty liver disease (a SNP in the PNPLA3 gene is associated with incr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Trans-splicing
''Trans''-splicing is a special form of RNA processing where exons from two different primary RNA transcripts are joined end to end and ligated. It is usually found in eukaryotes and mediated by the spliceosome, although some bacteria and archaea also have "half-genes" for tRNAs. Genic ''trans''-splicing Whereas "normal" (''cis''-)splicing processes a single molecule, ''trans''-splicing generates a single RNA transcript from multiple separate pre-mRNAs. This phenomenon can be exploited for molecular therapy to address mutated gene products. Genic trans-splicing allows variability in RNA diversity and increases proteome complexity. Oncogenesis While some fusion transcripts occur via ''trans''-splicing in normal human cells, ''trans''-splicing can also be the mechanism behind certain oncogenic fusion transcripts. SL ''trans''-splicing Spliced leader (SL) ''trans''-splicing is used by certain microorganisms, notably protists of the Kinetoplastae class to express genes. In ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
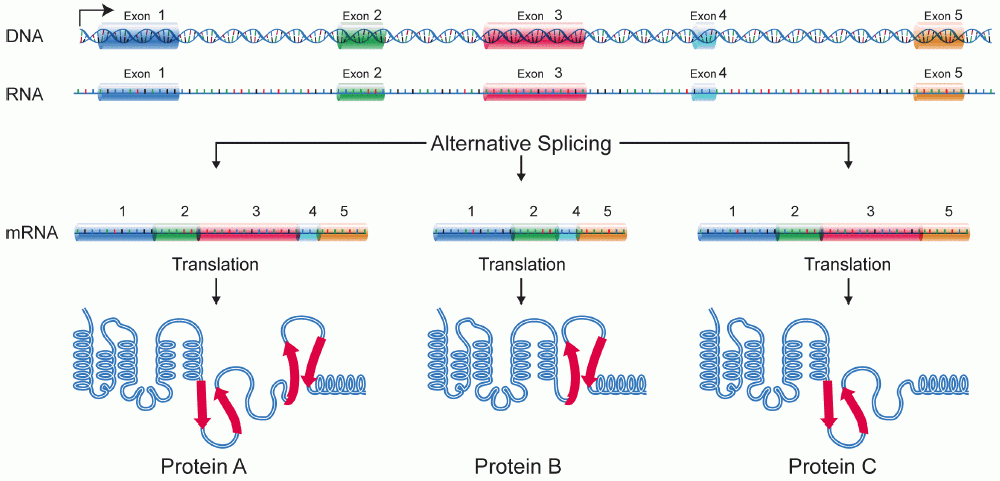
Alternative Splicing
Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Housekeeping Gene
In molecular biology, housekeeping genes are typically constitutive genes that are required for the maintenance of basic cellular function, and are expressed in all cells of an organism under normal and patho-physiological conditions. Although some housekeeping genes are expressed at relatively constant rates in most non-pathological situations, the expression of other housekeeping genes may vary depending on experimental conditions. The origin of the term "housekeeping gene" remains obscure. Literature from 1976 used the term to describe specifically tRNA and rRNA. For experimental purposes, the expression of one or multiple housekeeping genes is used as a reference point for the analysis of expression levels of other genes. The key criterion for the use of a housekeeping gene in this manner is that the chosen housekeeping gene is uniformly expressed with low variance under both control and experimental conditions. Validation of housekeeping genes should be performed before the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |