|
Retrotransposon
Retrotransposons (also called Class I transposable elements) are mobile elements which move in the host genome by converting their transcribed RNA into DNA through reverse transcription. Thus, they differ from Class II transposable elements, or DNA transposons, in utilizing an RNA intermediate for the transposition and leaving the transposition donor site unchanged. Through reverse transcription, retrotransposons amplify themselves quickly to become abundant in eukaryotic genomes such as maize (49–78%) and humans (42%). They are only present in eukaryotes but share features with retroviruses such as HIV, for example, discontinuous reverse transcriptase-mediated extrachromosomal recombination. There are two main types of retrotransposons, long terminal repeats (LTRs) and non-long terminal repeats (non-LTRs). Retrotransposons are classified based on sequence and method of transposition. Most retrotransposons in the maize genome are LTR, whereas in humans they are mostly non-L ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Retrotransposons
Retrotransposons (also called Class I transposable elements) are transposable element, mobile elements which move in the host genome by converting their transcribed RNA into DNA through reverse transcription. Thus, they differ from Class II transposable elements, or DNA transposons, in utilizing an RNA intermediate for the transposition and leaving the transposition donor site unchanged. Through reverse transcription, retrotransposons amplify themselves quickly to become abundant in eukaryote, eukaryotic genomes such as maize (49–78%) and humans (42%). They are only present in eukaryotes but share features with retroviruses such as HIV, for example, discontinuous reverse transcriptase-mediated extrachromosomal recombination. There are two main types of retrotransposons, long terminal repeats (LTRs) and non-long terminal repeats (non-LTRs). Retrotransposons are classified based on sequence and method of transposition. Most retrotransposons in the maize genome are LTR, whereas in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Retrotransposon Marker
Retrotransposon markers are components of DNA which are used as cladistic markers. They assist in determining the common ancestry, or not, of related taxa. The "presence" of a given retrotransposon in related taxa suggests their orthologous integration, a derived condition acquired via a common ancestry, while the "absence" of particular elements indicates the plesiomorphic condition prior to integration in more distant taxa. The use of presence/absence analyses to reconstruct the systematic biology of mammals depends on the availability of retrotransposons that were actively integrating before the divergence of a particular species. Details The analysis of SINEs – Short INterspersed Elements – LINEs – Long INterspersed Elements – or truncated LTRs – Long Terminal Repeats – as molecular cladistic markers represents a particularly interesting complement to DNA sequence and morphological data. The reason for this is that retrotransposons are assumed to represent pow ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Long Interspersed Nuclear Element
Long interspersed nuclear elements (LINEs) (also known as long interspersed nucleotide elements or long interspersed elements) are a group of non-LTR (long terminal repeat) retrotransposons that are widespread in the genome of many eukaryotes. LINEs contain an internal RNA polymerase, Pol II promoter to initiate Transcription (biology), transcription into Messenger RNA, mRNA, and encode one or two proteins, ORF1 and ORF2. The functional domains present within ORF1 vary greatly among LINEs, but often exhibit RNA/DNA binding activity. ORF2 is essential to successful retrotransposition, and encodes a protein with both reverse transcriptase and endonuclease activity. LINEs are the most abundant transposable element within the human genome, with approximately 20.7% of the sequences identified as being derived from LINEs. The only active lineage of LINE found within humans belongs to the LINE1, LINE-1 class, and is referred to as L1Hs. The human genome contains an estimated 100,000 tru ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transposable Element
A transposable element (TE), also transposon, or jumping gene, is a type of mobile genetic element, a nucleic acid sequence in DNA that can change its position within a genome. The discovery of mobile genetic elements earned Barbara McClintock a Nobel Prize in 1983. There are at least two classes of TEs: Class I TEs or retrotransposons generally function via reverse transcription, while Class II TEs or DNA transposons encode the protein transposase, which they require for insertion and excision, and some of these TEs also encode other proteins. Discovery by Barbara McClintock Barbara McClintock discovered the first TEs in maize (''Zea mays'') at the Cold Spring Harbor Laboratory in New York. McClintock was experimenting with maize plants that had broken chromosomes. In the winter of 1944–1945, McClintock planted corn kernels that were self-pollinated, meaning that the silk (style) of the flower received pollen from its own anther. These kernels came from a long line ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Long Terminal Repeat
A long terminal repeat (LTR) is a pair of identical sequences of DNA, several hundred base pairs long, which occur in eukaryotic genomes on either end of a series of genes or pseudogenes that form a retrotransposon or an endogenous retrovirus or a retroviral provirus. All retroviral genomes are flanked by LTRs, while there are some retrotransposons without LTRs. Typically, an element flanked by a pair of LTRs will encode a reverse transcriptase and an integrase, allowing the element to be copied and inserted at a different location of the genome. Copies of such an LTR-flanked element can often be found hundreds or thousands of times in a genome. LTR retrotransposons comprise about 8% of the human genome. The first LTR sequences were found by A.P. Czernilofsky and J. Shine in 1977 and 1980. Transcription The LTR-flanked sequences are partially transcribed into an RNA intermediate, followed by reverse transcription into complementary DNA (cDNA) and ultimately dsDNA (doubl ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alu Element
An Alu element is a short stretch of DNA originally characterized by the action of the ''Arthrobacter luteus (Alu)'' restriction endonuclease. ''Alu'' elements are the most abundant transposable elements in the human genome, present in excess of one million copies. Most ''Alu'' elements are thought to be selfish or parasitic DNA. However, it has been suggested that at least some are likely to play a role in evolution and have been used as genetic markers. They are derived from the small cytoplasmic 7SL RNA, a component of the signal recognition particle. ''Alu'' elements are not highly conserved within primate genomes, as only a minority have retained activity, and originated in the genome of an ancestor of Supraprimates. ''Alu'' insertions have been implicated in several inherited human diseases and in various forms of cancer. The study of Alu elements has also been important in elucidating human population genetics and the evolution of primates, including the evolution of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Metaviridae
''Metaviridae'' is a family of viruses which exist as Ty3-gypsy LTR retrotransposons in a eukaryotic host's genome. They are closely related to retroviruses: members of the family ''Metaviridae'' share many genomic elements with retroviruses, including length, organization, and genes themselves. This includes genes that encode reverse transcriptase, integrase, and capsid proteins. The reverse transcriptase and integrase proteins are needed for the retrotransposon activity of the virus. In some cases, virus-like particles can be formed from capsid proteins. Some assembled virus-like particles of members of the family ''Metaviridae'' can penetrate and infect previously uninfected cells. An example of this is the gypsy, a retroelement found in the ''Drosophila melanogaster'' genome. The ability to infect other cells is determined by the presence of the retroviral ''env'' genes which encode coat proteins. Metaviridae retrotransposons are found in all eukaryotes known and studied ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribonuclease H
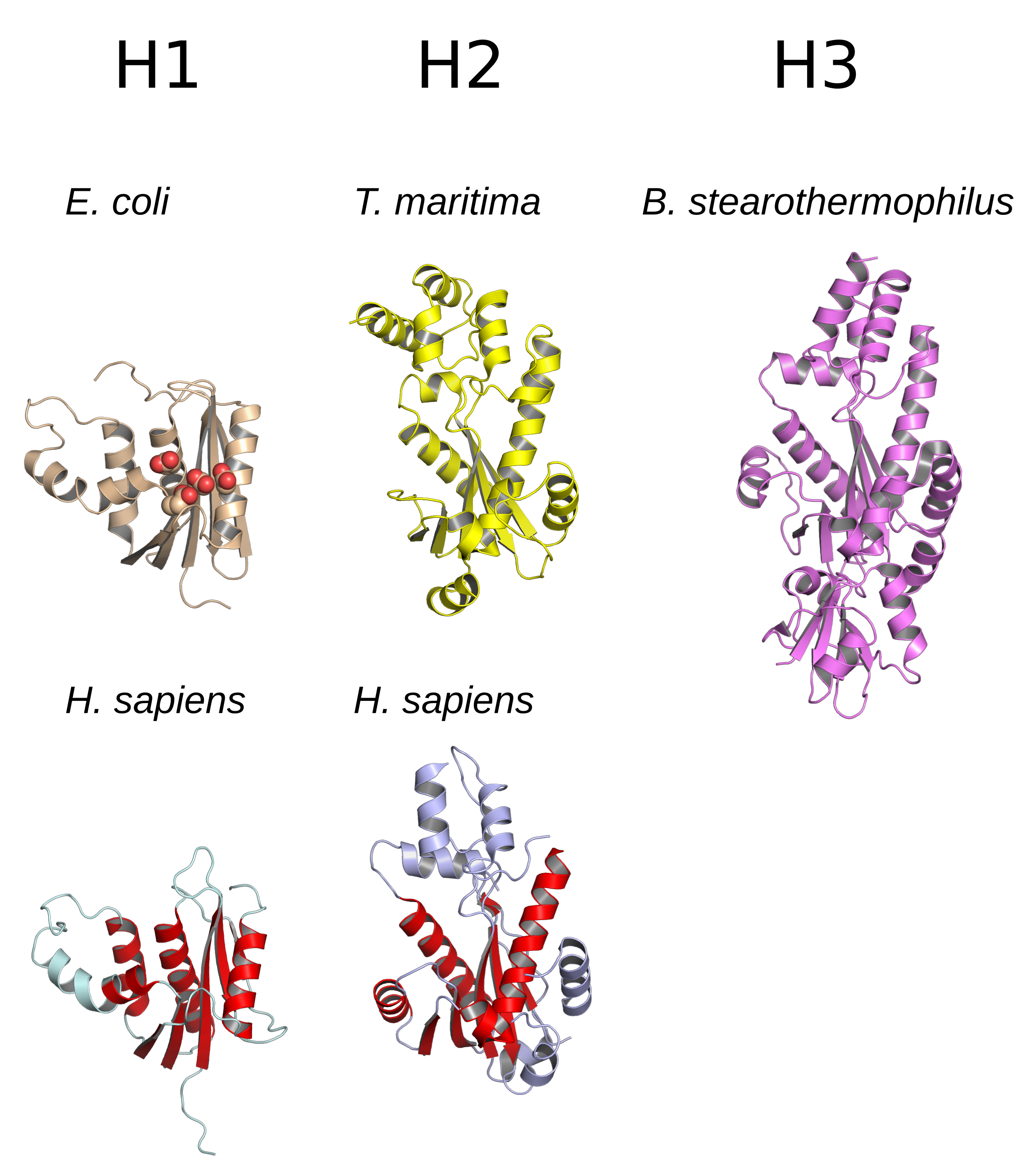
Ribonuclease H (abbreviated RNase H or RNH) is a family of non-sequence-specific endonuclease enzymes that catalyze the cleavage of RNA in an RNA/DNA substrate via a hydrolytic mechanism. Members of the RNase H family can be found in nearly all organisms, from bacteria to archaea to eukaryotes. The family is divided into evolutionarily related groups with slightly different substrate preferences, broadly designated ribonuclease H1 and H2. The human genome encodes both H1 and H2. Human ribonuclease H2 is a heterotrimeric complex composed of three subunits, mutations in any of which are among the genetic causes of a rare disease known as Aicardi–Goutières syndrome. A third type, closely related to H2, is found only in a few prokaryotes, whereas H1 and H2 occur in all domains of life. Additionally, RNase H1-like retroviral ribonuclease H domains occur in multidomain reverse transcriptase proteins, which are encoded by retroviruses such as HIV and are required for viral repl ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Reverse Transcription
A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B virus, hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryote, eukaryotic cells to extend the telomeres at the ends of their Chromosome#Eukaryotes, linear chromosomes. The process does not violate the flows of genetic information as described by the classical central dogma of molecular biology, central dogma, but rather expands it to include transfers of information from RNA to DNA. Retroviral RT has three sequential biochemical activities: RNA-dependent DNA polymerase activity, ribonuclease H (RNase H), and DNA-dependent DNA polymerase activity. Collectively, these activities enable the enzyme to convert single-stranded RNA into double-stranded cDNA. In retroviruses and retrotransposons, this cDNA can then inte ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RetrOryza
RetrOryza is a database of Long terminal repeat-retrotransposons for the rice genome. See also * Long terminal repeat * Retrotransposon * Rice Rice is a cereal grain and in its Domestication, domesticated form is the staple food of over half of the world's population, particularly in Asia and Africa. Rice is the seed of the grass species ''Oryza sativa'' (Asian rice)—or, much l ... References External links * http://retroryza.fr/ Biological databases Molecular genetics Mobile genetic elements Rice {{Biodatabase-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Reverse Transcriptase
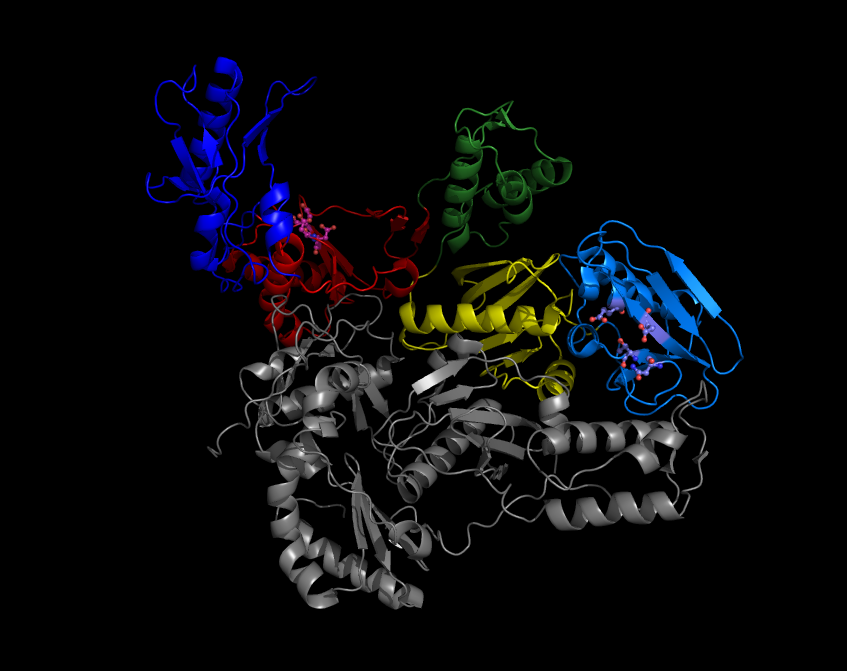
A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. The process does not violate the flows of genetic information as described by the classical central dogma, but rather expands it to include transfers of information from RNA to DNA. Retroviral RT has three sequential biochemical activities: RNA-dependent DNA polymerase activity, ribonuclease H (RNase H), and DNA-dependent DNA polymerase activity. Collectively, these activities enable the enzyme to convert single-stranded RNA into double-stranded cDNA. In retroviruses and retrotransposons, this cDNA can then integrate into the host genome, from which new RNA copies can be made via host-cell ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pseudoviridae
''Pseudoviridae'' is a family of viruses, which includes three genera. Viruses of the family are actually LTR retrotransposons of the Ty1-copia family. They replicate via structures called virus-like particles (VLPs). VLPs are not infectious like normal virions, but they nevertheless make up an essential part of the pseudoviral lifecycle. Taxonomy ''Pseudoviridae'' is unofficially classified under group VI RNA Reverse Transcribing Viruses and infect fungi and invertebrates. ''Pseudoviridae'' comprises highly divergent members and most ''Pseudoviridae'' encode Gag and Pol on a single open reading frame. ''Pseudoviridae'' is included in the order ''Ortervirales'' along with families ''Belpaoviridae'', ''Metaviridae'', ''Retroviridae'', and ''Caulimoviridae''. The family includes the following genera: * ''Hemivirus'' * ''Pseudovirus (genus), Pseudovirus'' * ''Sirevirus'' Further ''Pseudoviridae'' species not classified into a genus are: * Penicillium camemberti virus – GP1NCB ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |