|
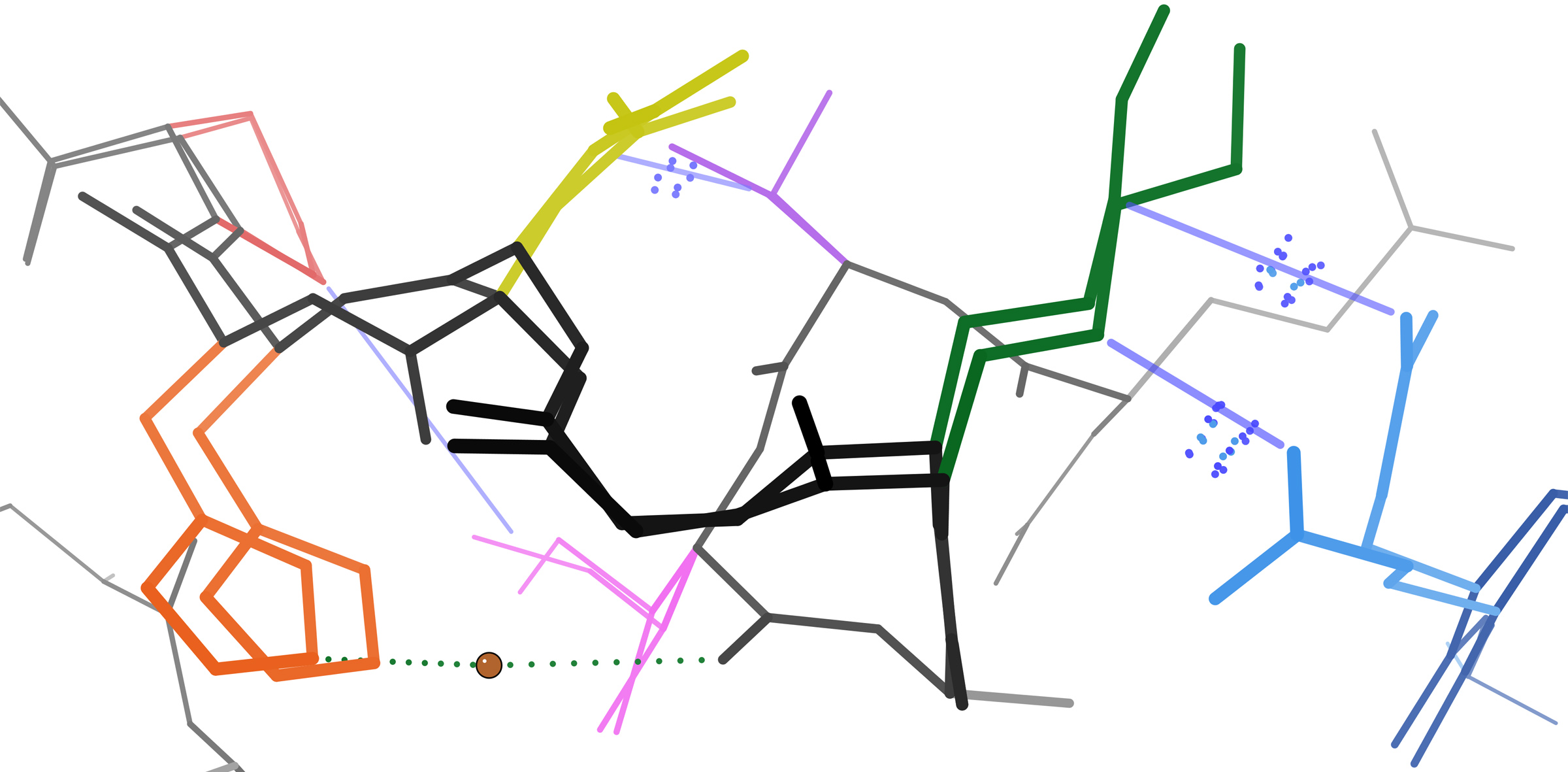
Omega Loop
The omega loop is a non-regular protein structural motif, consisting of a loop of six or more amino acid residues and any amino acid sequence. The defining characteristic is that residues that make up the beginning and end of the loop are close together in space with no intervening lengths of regular secondary structural motifs. It is named after its shape, which resembles the upper-case Greek letter Omega (Ω). Structure Omega loops, being non-regular, non-repeating secondary structural units, have a variety of three-dimensional shapes. Omega loop shapes are analyzed to identify recurring patterns in dihedral angles and overall loop shape to help identify potential roles in protein folding and function. Since loops are almost always at the protein surface, it is often assumed that these structures are flexible; however, different omega loops exhibit ranges of flexibility across different time scales of protein motion and have been identified as playing a role in the folding of so ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ligand (biochemistry)
In biochemistry and pharmacology, a ligand is a substance that forms a complex with a biomolecule to serve a biological purpose. The etymology stems from ''ligare'', which means 'to bind'. In protein-ligand binding, the ligand is usually a molecule which produces a signal by binding to a site on a target protein. The binding typically results in a change of conformational isomerism (conformation) of the target protein. In DNA-ligand binding studies, the ligand can be a small molecule, ion, or protein which binds to the DNA double helix. The relationship between ligand and binding partner is a function of charge, hydrophobicity, and molecular structure. Binding occurs by intermolecular forces, such as ionic bonds, hydrogen bonds and Van der Waals forces. The association or docking is actually reversible through dissociation. Measurably irreversible covalent bonding between a ligand and target molecule is atypical in biological systems. In contrast to the definition of lig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Dynamics
Proteins are generally thought to adopt unique structures determined by their amino acid sequences. However, proteins are not strictly static objects, but rather populate ensembles of (sometimes similar) conformations. Transitions between these states occur on a variety of length scales (tenths of Å to nm) and time scales (ns to s), and have been linked to functionally relevant phenomena such as allosteric signaling and enzyme catalysis. The study of protein dynamics is most directly concerned with the transitions between these states, but can also involve the nature and equilibrium populations of the states themselves. These two perspectives—kinetics and thermodynamics, respectively—can be conceptually synthesized in an "energy landscape" paradigm: highly populated states and the kinetics of transitions between them can be described by the depths of energy wells and the heights of energy barriers, respectively. Local flexibility: atoms and residues Portions of protein s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta-lactamase
Beta-lactamases, (β-lactamases) are enzymes () produced by bacteria that provide multi-resistance to beta-lactam antibiotics such as penicillins, cephalosporins, cephamycins, monobactams and carbapenems (ertapenem), although carbapenems are relatively resistant to beta-lactamase. Beta-lactamase provides antibiotic resistance by breaking the antibiotics' structure. These antibiotics all have a common element in their molecular structure: a four-atom ring known as a beta-lactam (β-lactam) ring. Through hydrolysis, the enzyme lactamase breaks the β-lactam ring open, deactivating the molecule's antibacterial properties. Beta-lactam antibiotics are typically used to target a broad spectrum of gram-positive and gram-negative bacteria. Beta-lactamases produced by gram-negative bacteria are usually secreted, especially when antibiotics are present in the environment. Structure The structure of a '' Streptomyces'' serine β-lactamase (SBLs) is given by . The alpha-beta fold ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein C
Protein C, also known as autoprothrombin IIA and blood coagulation factor XIX, is a zymogen, that is, an inactive enzyme. The activated form plays an important role in regulating anticoagulation, inflammation, and apoptosis, cell death and maintaining the vascular permeability, permeability of blood vessel walls in humans and other animals. Activated protein C (APC) performs these operations primarily by proteolysis, proteolytically inactivating proteins Factor V, Factor Va and Factor VIII, Factor VIIIa. APC is classified as a serine protease since it contains a residue (chemistry), residue of serine in its active site. In humans, protein C is encoded by the ''PROC'' gene, which is found on chromosome 2 (human), chromosome 2. The zymogenic form of protein C is a vitamin K-dependent glycoprotein that circulates in blood plasma. Its structure is that of a two-chain polypeptide consisting of a light chain and a heavy chain connected by a disulfide bond. The protein C zymog ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mitosis, or meiosis or other types of damage to DNA (such as pyrimidine dimers caused by exposure to ultraviolet radiation), which then may undergo error-prone repair (especially microhomology-mediated end joining), cause an error during other forms of repair, or cause an error during replication (translesion synthesis). Mutations may also result from insertion or deletion of segments of DNA due to mobile genetic elements. Mutations may or may not produce detectable changes in the observable characteristics (phenotype) of an organism. Mutations play a part in both normal and abnormal biological processes including: evolution, cancer, and the development of the immune system, including junctional diversity. Mutation is the ultimate source o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Coagulation Disorder
Coagulopathy (also called a bleeding disorder) is a condition in which the blood's ability to coagulate (form clots) is impaired. This condition can cause a tendency toward prolonged or excessive bleeding (bleeding diathesis), which may occur spontaneously or following an injury or medical and dental procedures. Coagulopathies are sometimes erroneously referred to as "clotting disorders", but a clotting disorder is the opposite, defined as a predisposition to excessive clot formation (thrombus), also known as a hypercoagulable state or thrombophilia. Signs and symptoms Coagulopathy may cause uncontrolled internal or external bleeding. Left untreated, uncontrolled bleeding may cause damage to joints, muscles, or internal organs and may be life-threatening. People should seek immediate medical care for serious symptoms, including heavy external bleeding, blood in the urine or stool, double vision, severe head or neck pain, repeated vomiting, difficulty walking, convulsions, or se ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Heritable
Heredity, also called inheritance or biological inheritance, is the passing on of traits from parents to their offspring; either through asexual reproduction or sexual reproduction, the offspring cells or organisms acquire the genetic information of their parents. Through heredity, variations between individuals can accumulate and cause species to evolve by natural selection. The study of heredity in biology is genetics. Overview In humans, eye color is an example of an inherited characteristic: an individual might inherit the "brown-eye trait" from one of the parents. Inherited traits are controlled by genes and the complete set of genes within an organism's genome is called its genotype. The complete set of observable traits of the structure and behavior of an organism is called its phenotype. These traits arise from the interaction of its genotype with the environment. As a result, many aspects of an organism's phenotype are not inherited. For example, suntanned skin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Triosephosphateisomerase
Triose-phosphate isomerase (TPI or TIM) is an enzyme () that catalyzes the reversible interconversion of the triose phosphate isomers dihydroxyacetone phosphate and D-glyceraldehyde 3-phosphate. TPI plays an important role in glycolysis and is essential for efficient energy production. TPI has been found in nearly every organism searched for the enzyme, including animals such as mammals and insects as well as in fungi, plants, and bacteria. However, some bacteria that do not perform glycolysis, like ureaplasmas, lack TPI. In humans, deficiencies in TPI are associated with a progressive, severe neurological disorder called triose phosphate isomerase deficiency. Triose phosphate isomerase deficiency is characterized by chronic hemolytic anemia. While there are various mutations that cause this disease, most include the replacement of glutamic acid at position 104 with an aspartic acid. Triose phosphate isomerase is a highly efficient enzyme, performing the reaction billions ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. Almost all metabolic processes in the cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts are catalytic RNA molecules, called ribozymes. Enzymes' specificity comes from their unique three-dimensional structures. Like all catalysts, enzymes increase the reaction ra ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Structural Motif
In a polymer, chain-like biological molecule, such as a protein or nucleic acid, a structural motif is a common Biomolecular structure#Tertiary structure, three-dimensional structure that appears in a variety of different, evolutionarily unrelated molecules. A structural motif does not have to be associated with a sequence motif; it can be represented by different and completely unrelated sequences in different proteins or RNA. In nucleic acids Depending upon the sequence and other conditions, nucleic acids can form a variety of structural motifs which is thought to have biological significance. ;Stem-loop: Stem-loop intramolecular base pairing is a pattern that can occur in single-stranded DNA or, more commonly, in RNA. The structure is also known as a hairpin or hairpin loop. It occurs when two regions of the same strand, usually complementary in nucleotide sequence when read in opposite directions, base-pair to form a double helix that ends in an unpaired loop. The re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |