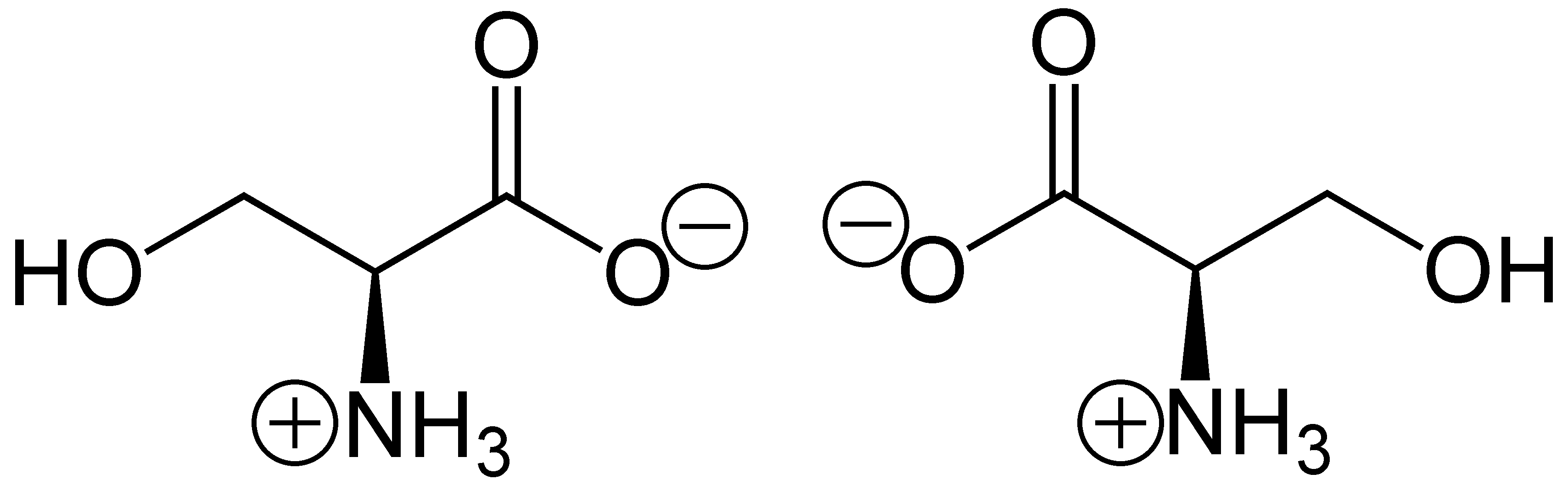|
N CAP
The term N cap (N-cap, Ncap) describes an amino acid in a particular position within a protein or polypeptide.{{cite journal, last=Leader, first=DP, author2=Milner-White EJ , title=The structure of the ends of helices in globular proteins, journal=Proteins, year=2011, volume=79, issue=3, pages=1010–1019, doi= 10.1002/prot.22942, pmid=21287629, s2cid=22240314 The N cap residue of an alpha helix is the first amino acid residue at the N terminus of the helix. More precisely, it is defined as the first residue (i) whose CO group is hydrogen-bonded to the NH group of residue i+4 (or sometimes residue i+3). Because of this it is sometimes also described as the residue prior to the helix. Capping motifs are those often found at the N cap. Asx turns, ST turns, and asx motifs are often found at such situations, with the asx or serine or threonine residue at the N cap. The C cap The term C cap (C-cap, Ccap) describes an amino acid in a particular position within a protein or polypeptide.{ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Amino Acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha amino acids appear in the genetic code. Amino acids can be classified according to the locations of the core structural functional groups, as Alpha and beta carbon, alpha- , beta- , gamma- or delta- amino acids; other categories relate to Chemical polarity, polarity, ionization, and side chain group type (aliphatic, Open-chain compound, acyclic, aromatic, containing hydroxyl or sulfur, etc.). In the form of proteins, amino acid '' residues'' form the second-largest component (water being the largest) of human muscles and other tissues. Beyond their role as residues in proteins, amino acids participate in a number of processes such as neurotransmitter transport and biosynthesis. It is thought that they played a key role in enabling life ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polypeptide
Peptides (, ) are short chains of amino acids linked by peptide bonds. Long chains of amino acids are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides. A polypeptide is a longer, continuous, unbranched peptide chain. Hence, peptides fall under the broad chemical classes of biological polymers and oligomers, alongside nucleic acids, oligosaccharides, polysaccharides, and others. A polypeptide that contains more than approximately 50 amino acids is known as a protein. Proteins consist of one or more polypeptides arranged in a biologically functional way, often bound to ligands such as coenzymes and cofactors, or to another protein or other macromolecule such as DNA or RNA, or to complex macromolecular assemblies. Amino acids that have been incorporated into peptides are termed residues. A water molecule is released during formation of each amide bond.. All peptides except cyclic peptides ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helix
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence. The alpha helix is also called a classic Pauling–Corey–Branson α-helix. The name 3.613-helix is also used for this type of helix, denoting the average number of residues per helical turn, with 13 atoms being involved in the ring formed by the hydrogen bond. Among types of local structure in proteins, the α-helix is the most extreme and the most predictable from sequence, as well as the most prevalent. Discovery In the early 1930s, William Astbury showed that there were drastic changes in the X-ray fiber diffraction of moist wool or hair fibers upon significant stretching. The data suggested that the unstretched fibers had a coiled molecular structure with a characteristic repeat of ≈. Astb ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
N Terminus
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein. Chemistry Each amino acid has an amine group and a carboxylic group. Amino acids link to one another by peptide bonds which form through a dehydration reaction that jo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hydrogen-bond
In chemistry, a hydrogen bond (or H-bond) is a primarily electrostatic force of attraction between a hydrogen (H) atom which is covalently bound to a more electronegative "donor" atom or group (Dn), and another electronegative atom bearing a lone pair of electrons—the hydrogen bond acceptor (Ac). Such an interacting system is generally denoted , where the solid line denotes a polar covalent bond, and the dotted or dashed line indicates the hydrogen bond. The most frequent donor and acceptor atoms are the second-row elements nitrogen (N), oxygen (O), and fluorine (F). Hydrogen bonds can be intermolecular (occurring between separate molecules) or intramolecular (occurring among parts of the same molecule). The energy of a hydrogen bond depends on the geometry, the environment, and the nature of the specific donor and acceptor atoms and can vary between 1 and 40 kcal/mol. This makes them somewhat stronger than a van der Waals interaction, and weaker than fully covalent or ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Asx Turn
The Asx turn is a structural feature in proteins and polypeptides. It consists of three amino acid residues (labeled i, i+1 and i+2) in which residue i is an aspartate (Asp) or asparagine (Asn) that forms a hydrogen bond from its sidechain CO group to the mainchain NH group of residue i+2. About 14% of Asx residues present in proteins belong to Asx turns. The name "Asx" is used here to represent either of the amino acids aspartate (Asp) or asparagine (Asn). Types Four types of Asx turn can be distinguished: types I, I’, II and II’. These categories correspond to those of the better-known hydrogen-bonded beta turns, which have four residues and a hydrogen bond between the CO of residue i and the NH of residue i+3. Asx turns and beta turns have structurally similar hydrogen-bonded loops and exhibit sidechain-mainchain mimicry in the sense that the sidechain of residue i of the Asx turn mimics the mainchain of residue i of the beta turn. Regarding their occurrence in proteins, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ST Turn
The ST turn is a structural feature in proteins and polypeptides. Each consists of three amino acid residues (labeled ''i'', ''i'' + 1 and ''i'' + 2) in which residue ''i'' is a serine (S) or threonine (T) that forms a hydrogen bond from its sidechain oxygen group to the mainchain NH group of residue ''i'' + 2. Similar Structural motif, motifs occur with aspartate or asparagine as residue ''i'', called asx turn. Four types of asx turn and ST turn can be distinguished: types I, I’, II and II’. These categories correspond (via sidechain-mainchain mimicry of residue i) to those of the more abundant hydrogen-bonded beta turns, which have four residues and a hydrogen bond between the CO of residue ''i'' and the NH of residue ''i'' + 3. Regarding their occurrence in proteins, they differ in that type I is the commonest of the four beta turns while type II’ is the commonest of the ST and asx turns. Asx and ST turns both occur frequently at th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Asx Motif
The Asx motif is a commonly occurring feature in proteins and polypeptides. It consists of four or five amino acid residues with either aspartate or asparagine as the first residue (residue i). It is defined by two internal hydrogen bonds. One is between the side chain oxygen of residue i and the main chain NH of residue i+2 or i+3; the other is between the main chain oxygen of residue i and the main chain NH of residue i+3 or i+4. Asx motifs occur commonly in proteins and polypeptides. When one of the hydrogen bonds is between the main chain oxygen of residue i and the side chain NH of residue i+3 the motif incorporates a beta turn. When one of the hydrogen bonds is between the side chain oxygen of residue i and the main chain NH of residue i+2 the motif incorporates an Asx turn. As with Asx turns, a significant proportion of Asx motifs occur at the N-terminus of an alpha helix with the Asx as the N cap residue. Asx motifs have thus often been described as helix capping fea ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine
Serine (symbol Ser or S) is an α-amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated − form under biological conditions), a carboxyl group (which is in the deprotonated − form under biological conditions), and a side chain consisting of a hydroxymethyl group, classifying it as a polar amino acid. It can be synthesized in the human body under normal physiological circumstances, making it a nonessential amino acid. It is encoded by the codons UCU, UCC, UCA, UCG, AGU and AGC. Occurrence This compound is one of the naturally occurring proteinogenic amino acids. Only the L-stereoisomer appears naturally in proteins. It is not essential to the human diet, since it is synthesized in the body from other metabolites, including glycine. Serine was first obtained from silk protein, a particularly rich source, in 1865 by Emil Cramer. Its name is derived from the Latin for silk, ''sericum''. Serine's structure was estab ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Threonine
Threonine (symbol Thr or T) is an amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated −NH form under biological conditions), a carboxyl group (which is in the deprotonated −COO− form under biological conditions), and a side chain containing a hydroxyl group, making it a polar, uncharged amino acid. It is essential in humans, meaning the body cannot synthesize it: it must be obtained from the diet. Threonine is synthesized from aspartate in bacteria such as ''E. coli''. It is encoded by all the codons starting AC (ACU, ACC, ACA, and ACG). Threonine sidechains are often hydrogen bonded; the most common small motifs formed are based on interactions with serine: ST turns, ST motifs (often at the beginning of alpha helices) and ST staples (usually at the middle of alpha helices). Modifications The threonine residue is susceptible to numerous posttranslational modifications. The hydroxyl side-chain can unde ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C Cap
The term C cap (C-cap, Ccap) describes an amino acid in a particular position within a protein or polypeptide.{{cite journal, last=Leader, first=DP, author2=Milner-White EJ , title=The structure of the ends of helices in globular proteins, journal=Proteins, year=2011, volume=79, issue=3, pages=1010–1019, doi= 10.1002/prot.22942, pmid=21287629, s2cid=22240314 The C cap residue of an alpha helix is the last amino acid residue at the C terminus of the helix. More precisely, it is defined as the last residue (i) whose NH group is hydrogen-bonded to the CO group of residue i-4 (or sometimes residue i-3). Because of this it is sometimes also described as the residue following the helix. Certain motifs occur commonly at or around the C cap, notably the Schellman loop and the niche (protein structural motif). The N cap The term N cap (N-cap, Ncap) describes an amino acid in a particular position within a protein or polypeptide.{{cite journal, last=Leader, first=DP, author2=Milner-White ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |