|
Messenger RNP
Messenger RNP (messenger ribonucleoprotein) is mRNA with bound proteins. mRNA does not exist "naked" ''in vivo'' but is always bound by various proteins while being synthesized, spliced, exported, and translated in the cytoplasm. Messenger RNPs were first discovered in Alexander S. Spirin's laboratory in Moscow, Russia in 1964. The discovery was based in their study of fish embryo cytoplasm extracts, where they found these mRNPs. This finding was discovered after the mRNA of the fish embryo was centrifuged. The mRNA liquid separated into two parts, having the scientists question what is separate of the mRNA from the ribosomes. Spirin and his collaborators analyzed the mRNA against CsCl density gradients and discovered that parts of the mRNA were coated in proteins. The weight ratio of mRNPs was found to be 1:3, mRNA to protein. mRNPs were thus denoted as ''informosomes'' by the lab. There are three major informosomes found in mammalian cells: nuclear ribonucleoproteins, cytoplasmic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme ( RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nonsense Mediated Decay
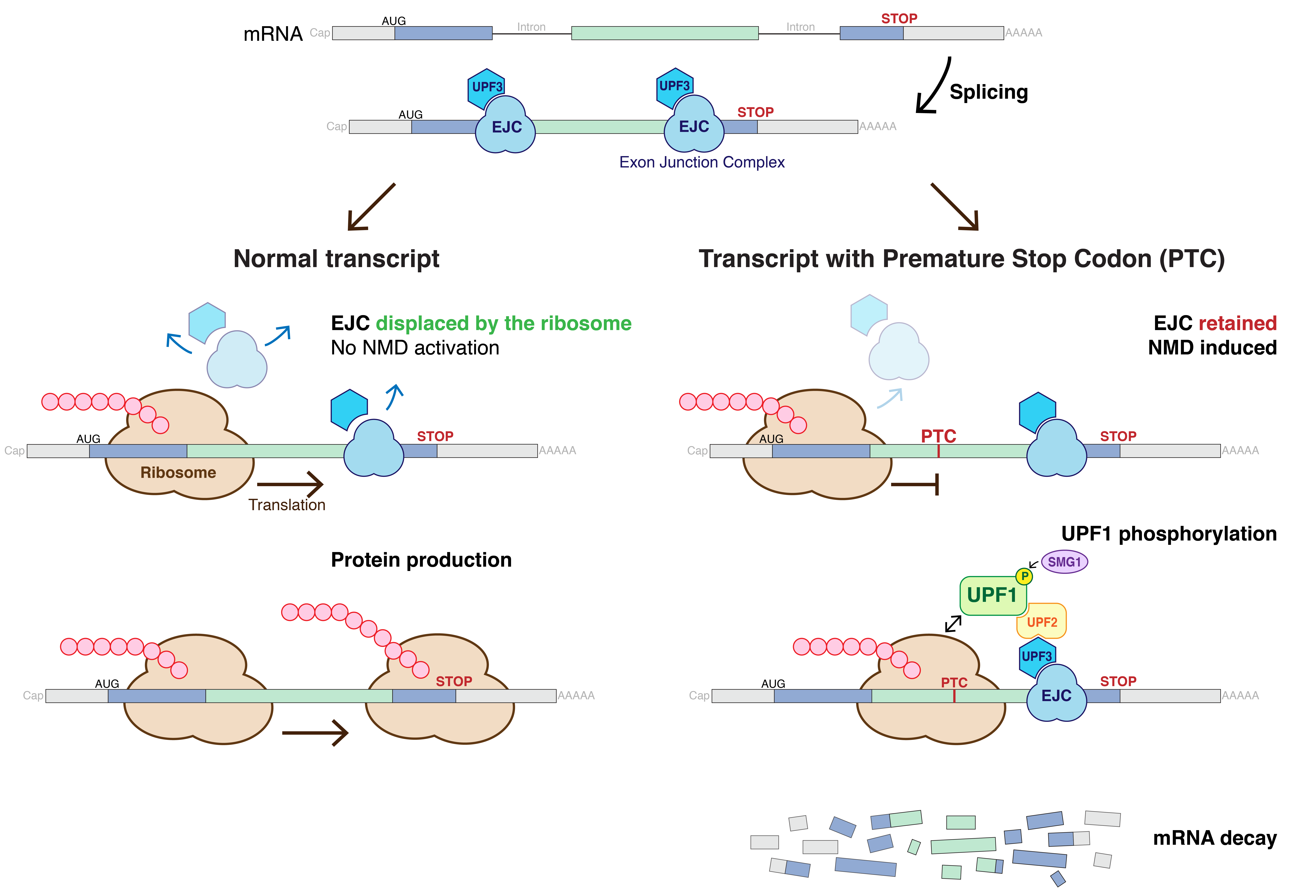
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all eukaryotes. Its main function is to reduce errors in gene expression by eliminating mRNA transcripts that contain premature stop codons. Translation of these aberrant mRNAs could, in some cases, lead to deleterious gain-of-function or dominant-negative activity of the resulting proteins. NMD was first described in human cells and in yeast almost simultaneously in 1979. This suggested broad phylogenetic conservation and an important biological role of this intriguing mechanism. NMD was discovered when it was realized that cells often contain unexpectedly low concentrations of mRNAs that are transcribed from alleles carrying nonsense mutations. Nonsense mutations code for a premature stop codon which causes the protein to be shortened. The truncated protein may or may not be functional, depending on the severity of what is not translated. In human genetics, NMD has the possibility to not only limit the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genet ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Heterogeneous Ribonucleoprotein Particle
Heterogeneous nuclear ribonucleoproteins (hnRNPs) are complexes of RNA and protein present in the cell nucleus during gene transcription and subsequent post-transcriptional modification of the newly synthesized RNA (pre-mRNA). The presence of the proteins bound to a pre-mRNA molecule serves as a signal that the pre-mRNA is not yet fully processed and therefore not ready for export to the cytoplasm. Since most mature RNA is exported from the nucleus relatively quickly, most RNA-binding protein in the nucleus exist as heterogeneous ribonucleoprotein particles. After splicing has occurred, the proteins remain bound to spliced introns and target them for degradation. hnRNPs are also integral to the 40s subunit of the ribosome and therefore important for the translation of mRNA in the cytoplasm. However, hnRNPs also have their own nuclear localization sequences (NLS) and are therefore found mainly in the nucleus. Though it is known that a few hnRNPs shuttle between the cytoplasm and nucle ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neurodegenerative Disease
A neurodegenerative disease is caused by the progressive loss of structure or function of neurons, in the process known as neurodegeneration. Such neuronal damage may ultimately involve cell death. Neurodegenerative diseases include amyotrophic lateral sclerosis, multiple sclerosis, Parkinson's disease, Alzheimer's disease, Huntington's disease, multiple system atrophy, and prion diseases. Neurodegeneration can be found in the brain at many different levels of neuronal circuitry, ranging from molecular to systemic. Because there is no known way to reverse the progressive degeneration of neurons, these diseases are considered to be incurable; however research has shown that the two major contributing factors to neurodegeneration are oxidative stress and inflammation. Biomedical research has revealed many similarities between these diseases at the subcellular level, including atypical protein assemblies (like proteinopathy) and induced cell death. These similarities suggest th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Demethylation
For molecular biology in mammals, DNA demethylation causes replacement of 5-methylcytosine (5mC) in a DNA sequence by cytosine (C) (see figure of 5mC and C). DNA demethylation can occur by an active process at the site of a 5mC in a DNA sequence or, in replicating cells, by preventing addition of methyl groups to DNA so that the replicated DNA will largely have cytosine in the DNA sequence (5mC will be diluted out). Methylated cytosine is frequently present in the linear DNA sequence where a cytosine is followed by a guanine in a 5' → 3' direction (a CpG site). In mammals, DNA methyltransferases (which add methyl groups to DNA bases) exhibit a strong sequence preference for cytosines at CpG sites. There appear to be more than 20 million CpG dinucleotides in the human genome (see genomic distribution). In mammals, on average, 70% to 80% of CpG cytosines are methylated, though the level of methylation varies with different tissues. Methylated cytosines often occur in groups ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Methylation
DNA methylation is a biological process by which methyl groups are added to the DNA molecule. Methylation can change the activity of a DNA segment without changing the sequence. When located in a gene promoter, DNA methylation typically acts to repress gene transcription. In mammals, DNA methylation is essential for normal development and is associated with a number of key processes including genomic imprinting, X-chromosome inactivation, repression of transposable elements, aging, and carcinogenesis. As of 2016, two nucleobases have been found on which natural, enzymatic DNA methylation takes place: adenine and cytosine. The modified bases are N6-methyladenineD. B. Dunn, J. D. Smith: ''The occurrence of 6-methylaminopurine in deoxyribonucleic acids.'' In: ''Biochem J.'' 68(4), Apr 1958, S. 627–636. PMID 13522672. ., 5-methylcytosineB. F. Vanyushin, S. G. Tkacheva, A. N. Belozersky: ''Rare bases in animal DNA.'' In: ''Nature.'' 225, 1970, S. 948–949. PMID 4391887. a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Fear Conditioning
Pavlovian fear conditioning is a behavioral paradigm in which organisms learn to predict aversive events. It is a form of learning in which an aversive stimulus (e.g. an electrical shock) is associated with a particular neutral context (e.g., a room) or neutral stimulus (e.g., a tone), resulting in the expression of fear responses to the originally neutral stimulus or context. This can be done by pairing the neutral stimulus with an aversive stimulus (e.g., an electric shock, loud noise, or unpleasant odor). Eventually, the neutral stimulus alone can elicit the state of fear. In the vocabulary of classical conditioning, the neutral stimulus or context is the "conditional stimulus" (CS), the aversive stimulus is the "unconditional stimulus" (US), and the fear is the "conditional response" (CR). Fear conditioning has been studied in numerous species, from snails to humans. In humans, conditioned fear is often measured with verbal report and galvanic skin response. In other animals ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exon Junction Complex
An exon junction complex (EJC) is a protein complex which forms on a pre-messenger RNA strand at the junction of two exons which have been joined together during RNA splicing. The EJC has major influences on translation, surveillance and localization of the spliced mRNA. It is first deposited onto mRNA during splicing and is then transported into the cytoplasm. There it plays a major role in post-transcriptional regulation of mRNA. It is believed that exon junction complexes provide a position-specific memory of the splicing event. The EJC consists of a stable heterotetramer core, which serves as a binding platform for other factors necessary for the mRNA pathway. The core of the EJC contains the protein eukaryotic initiation factor 4A-III ( eIF4A-III; a DEAD-box RNA helicase) bound to an adenosine triphosphate (ATP) analog, as well as the additional proteins Magoh and Y14. The binding of these proteins to nuclear speckled domains has been measured recently and it may be regula ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Proteins
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residues in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Christine Guthrie
Christine Guthrie (1945-2022) was an American yeast geneticist and American Cancer Society Research Professor of Genetics at University of California San Francisco. She showed that yeast have small nuclear RNAs (snRNAs) involved in splicing pre-messenger RNA into messenger RNA in eukaryotic cells. Guthrie cloned and sequenced the genes for yeast snRNA and established the role of base pairing between the snRNAs and their target sequences at each step in the removal of an intron. She also identified proteins that formed part of the spliceosome complex with the snRNAs. Elected to the National Academy of Sciences in 1993, Guthrie edited ''Guide to Yeast Genetics and Molecular Biology'', an influential methods series for many years. Early life and education Christine Guthrie was born in Brooklyn, New York. She received a BS in Zoology from University of Michigan and a PhD in genetics from University of Wisconsin. Her PhD advisor was Masayasu Nomura. She was the daughter of Bro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nuclear RNA
Small nuclear RNA (snRNA) is a class of small RNA molecules that are found within the splicing speckles and Cajal bodies of the cell nucleus in eukaryotic cells. The length of an average snRNA is approximately 150 nucleotides. They are transcribed by either RNA polymerase II or RNA polymerase III. Their primary function is in the processing of pre- messenger RNA ( hnRNA) in the nucleus. They have also been shown to aid in the regulation of transcription factors ( 7SK RNA) or RNA polymerase II (B2 RNA), and maintaining the telomeres. snRNA are always associated with a set of specific proteins, and the complexes are referred to as small nuclear ribonucleoproteins (snRNP, often pronounced "snurps"). Each snRNP particle is composed of a snRNA component and several snRNP-specific proteins (including Sm proteins, a family of nuclear proteins). The most common human snRNA components of these complexes are known, respectively, as: U1 spliceosomal RNA, U2 spliceosomal RNA, U4 splic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |