|
Intrinsic Termination
Intrinsic, or rho-independent termination, is a process in prokaryotes to signal the end of transcription and release the newly constructed RNA molecule. In prokaryotes such as E. coli, transcription is terminated either by a rho-dependent process or rho-independent process. In the Rho-dependent process, the rho-protein locates and binds the signal sequence in the mRNA and signals for cleavage. Contrarily, intrinsic termination does not require a special protein to signal for termination and is controlled by the specific sequences of RNA. When the termination process begins, the transcribed mRNA forms a stable secondary structure hairpin loop, also known as a Stem-loop. This RNA hairpin is followed by multiple uracil nucleotides. The bonds between uracil and adenine are very weak. A protein bound to RNA polymerase (nusA) binds to the stem-loop structure tightly enough to cause the polymerase to temporarily stall. This pausing of the polymerase coincides with transcription of the po ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
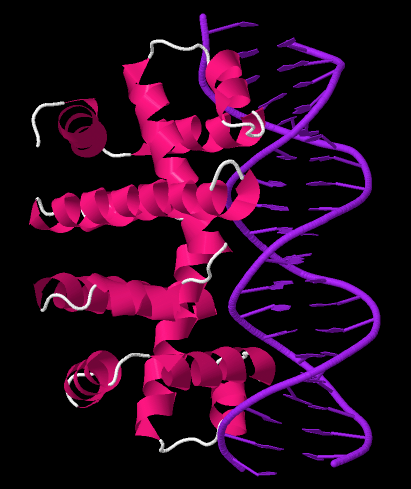
Intrinsic Termination Structure
In science and engineering, an intrinsic property is a property of a specified subject that exists itself or within the subject. An extrinsic property is not essential or inherent to the subject that is being characterized. For example, mass is an intrinsic property of any physical object, whereas weight is an extrinsic property that depends on the strength of the gravitational field in which the object is placed. Applications in science and engineering In materials science, an intrinsic property is independent of how much of a material is present and is independent of the form of the material, e.g., one large piece or a collection of small particles. Intrinsic properties are dependent mainly on the fundamental chemical composition and structure of the material. Extrinsic properties are differentiated as being dependent on the presence of avoidable chemical contaminants or structural defects. In biology, intrinsic effects originate from inside an organism or cell, such as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cytosine
Cytosine () ( symbol C or Cyt) is one of the four nucleobases found in DNA and RNA, along with adenine, guanine, and thymine (uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached (an amine group at position 4 and a keto group at position 2). The nucleoside of cytosine is cytidine. In Watson-Crick base pairing, it forms three hydrogen bonds with guanine. History Cytosine was discovered and named by Albrecht Kossel and Albert Neumann in 1894 when it was hydrolyzed from calf thymus tissues. A structure was proposed in 1903, and was synthesized (and thus confirmed) in the laboratory in the same year. In 1998, cytosine was used in an early demonstration of quantum information processing when Oxford University researchers implemented the Deutsch-Jozsa algorithm on a two qubit nuclear magnetic resonance quantum computer (NMRQC). In March 2015, NASA scientists reported the formation of cytosine, along with uracil and thym ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the central dogma of molecular biology first formulated by Francis Crick in 1958, further developed in his 1970 article, and expanded by the subsequent discoveries of reverse transcription and RNA replication. The process of gene expression is used by all known life—eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenotype, '' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Trp Operon
The ''trp'' operon is a group of genes that are transcribed together, encoding the enzymes that produce the amino acid tryptophan in bacteria. The ''trp'' operon was first characterized in ''Escherichia coli,'' and it has since been discovered in many other bacteria. The operon is regulated so that, when tryptophan is present in the environment, the genes for tryptophan synthesis are repressed. The ''trp'' operon contains five structural genes: ''trpE'', ''trpD'', ''trpC'', ''trpB'', and ''trpA'', which encode the enzymes needed to synthesize tryptophan. It also contains a repressive regulator gene called trpR. When tryptophan is present, the trpR protein binds to the operator, blocking transcription of the ''trp'' operon by RNA polymerase. This operon is an example of repressible negative regulation of gene expression. The repressor protein binds to the operator in the presence of tryptophan (repressing transcription) and is released from the operon when tryptophan is absent ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
WebGeSTer
WebGeSTer DB is a database of intrinsic transcription terminators See also * Intrinsic termination References External links * http://pallab.serc.iisc.ernet.in/gester Biological databases Gene expression Molecular biology {{Biodatabase-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Thermococcus Kodakarensis
''Thermococcus kodakarensis'' is a species of thermophilic archaea. The type strain ''T. kodakarensis'' KOD1 is one of the best-studied members of the genus. History ''T. kodakarensis'' was isolated from a solfatara near the shore of Kodakara Island, Kagoshima, Japan. The isolate was originally named ''Pyrococcus kodakarensis'' KOD1, but reclassified as a species of ''Thermococcus'', based on 16S rRNA sequence. Early research with ''T. kodakarensis'' was directed mostly at its thermostable enzymes, but its relative ease of handling and genetic manipulation facilitated by natural competence has made it an attractive system for the study of several biological processes. Properties ''T. kodakarensis'' cells are irregular cocci 1–2 μm in diameter, often occurring in pairs, and are highly motile by means of lophotrichous archaella. The cell wall consists of a layer of diether and tetraether lipids, and an outer glycoprotein coat. ''T. kodakarensis'' is an obligate ana ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TATA Box
In molecular biology, the TATA box (also called the Goldberg–Hogness box) is a sequence of DNA found in the core promoter region of genes in archaea and eukaryotes. The bacterial homolog of the TATA box is called the Pribnow box which has a shorter consensus sequence. The TATA box is considered a non-coding DNA sequence (also known as a cis-regulatory element). It was termed the "TATA box" as it contains a consensus sequence characterized by repeating T and A base pairs. How the term "box" originated is unclear. In the 1980s, while investigating nucleotide sequences in mouse genome loci, the Hogness box sequence was found and "boxed in" at the -31 position. When consensus nucleotides and alternative ones were compared, homologous regions were "boxed" by the researchers. The boxing in of sequences sheds light on the origin of the term "box". The TATA box was first identified in 1978 as a component of eukaryotic promoters. Transcription is initiated at the TATA box in TAT ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Archaea
Archaea ( ; singular archaeon ) is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotes. Archaea were initially classified as bacteria, receiving the name archaebacteria (in the Archaebacteria kingdom), but this term has fallen out of use. Archaeal cells have unique properties separating them from the other two domains, Bacteria and Eukaryota. Archaea are further divided into multiple recognized phyla. Classification is difficult because most have not been isolated in a laboratory and have been detected only by their gene sequences in environmental samples. Archaea and bacteria are generally similar in size and shape, although a few archaea have very different shapes, such as the flat, square cells of ''Haloquadratum walsbyi''. Despite this morphological similarity to bacteria, archaea possess genes and several metabolic pathways that are more closely related to those of eukaryotes, notably for the enzymes involved ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleation
In thermodynamics, nucleation is the first step in the formation of either a new thermodynamic phase or structure via self-assembly or self-organization within a substance or mixture. Nucleation is typically defined to be the process that determines how long an observer has to wait before the new phase or self-organized structure appears. For example, if a volume of water is cooled (at atmospheric pressure) below 0°C, it will tend to freeze into ice, but volumes of water cooled only a few degrees below 0°C often stay completely free of ice for long periods (supercooling). At these conditions, nucleation of ice is either slow or does not occur at all. However, at lower temperatures nucleation is fast, and ice crystals appear after little or no delay. Nucleation is a common mechanism which generates first-order phase transitions, and it is the start of the process of forming a new thermodynamic phase. In contrast, new phases at continuous phase transitions start to form immedi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids can be manufactured as single-stranded molecules with any user-specified sequence, and so are vital for artificial gene synthesis, polymerase chain reaction (PCR), DNA sequencing, molecular cloning and as molecular probes. In nature, oligonucleotides are usually found as small RNA molecules that function in the regulation of gene expression (e.g. microRNA), or are degradation intermediates derived from the breakdown of larger nucleic acid molecules. Oligonucleotides are characterized by the sequence of nucleotide residues that make up the entire molecule. The length of the oligonucleotide is usually denoted by " -mer" (from Greek ''meros'', "part"). For example, an oligonucleotide of six nucleotides (nt) is a hexamer, while one of 25 nt wou ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hydrogen Bond
In chemistry, a hydrogen bond (or H-bond) is a primarily electrostatic force of attraction between a hydrogen (H) atom which is covalently bound to a more electronegative "donor" atom or group (Dn), and another electronegative atom bearing a lone pair of electrons—the hydrogen bond acceptor (Ac). Such an interacting system is generally denoted , where the solid line denotes a polar covalent bond, and the dotted or dashed line indicates the hydrogen bond. The most frequent donor and acceptor atoms are the second-row elements nitrogen (N), oxygen (O), and fluorine (F). Hydrogen bonds can be intermolecular (occurring between separate molecules) or intramolecular (occurring among parts of the same molecule). The energy of a hydrogen bond depends on the geometry, the environment, and the nature of the specific donor and acceptor atoms and can vary between 1 and 40 kcal/mol. This makes them somewhat stronger than a van der Waals interaction, and weaker than fully covalent ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |



.jpg)

