|
Variome
The variome is the whole set of genetic variations found in populations of species that have gone through a relatively short evolution change. For example, among humans, about 1 in every 1,200 nucleotide bases differ. The size of human variome in terms of effective population size is claimed to be about 10,000 individuals. This variation rate is comparatively small compared to other species. For example, the effective population size of tigers which perhaps has the whole population size less than 10,000 in the wild is not much smaller than the human species indicating a much higher level of genetic diversity although they are close to extinction in the wild. In practice, the variome can be the sum of the single nucleotide polymorphisms (SNPs), indels, and structural variation (SV) of a population or speciesThe Human Variome Projectseeks to compile this genetic variation data worldwide. Variomics is the study of variome and a branch of bioinformatics. Ethnic variomes The human vari ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Human Variome Project
The Human Variome Project (HVP) is the global initiative to collect and curate all human genetic variation affecting human health. Its mission is to improve health outcomes by facilitating the unification of data on human genetic variation and its impact on human health. Inception The HVP concept was conceived by Richard Cotton, a leader in the field of human genetic variation. His group, the Genomic Disorders Research Centre, based at the University of Melbourne and St. Vincent's Hospital, has established a consortium that covers genomic variation and its health implications in a comprehensive form. This consortium has encouraged the creation and supported many of the 571 gene specific variation databases currently available on the internet. However, these databases are of varying completeness and individualistic, so the Human Variome Project was born to establish a central project to encourage the collection and sourcing of this data, verifying it and ultimately using it for ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
KoVariome
KoVariome is the variome The variome is the whole set of genetic variations found in populations of species that have gone through a relatively short evolution change. For example, among humans, about 1 in every 1,200 nucleotide bases differ. The size of human variome in ... of Korean ethnic groups. It was initiated in 2010 when the Genome Research Foundation in Korea was established. KoVariome has produced around 100 Korean genome diversity data on 4 April 2018 in Scientific Reports and 1,094 Korean genome variation information on 27 May 2020. References {{Korea-stub Ethnic groups in Korea ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single Nucleotide Polymorphisms
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently large fraction of the population (e.g. 1% or more), many publications do not apply such a frequency threshold. For example, at a specific base position in the human genome, the Guanine, G nucleotide may appear in most individuals, but in a minority of individuals, the position is occupied by an Adenine, A. This means that there is a SNP at this specific position, and the two possible nucleotide variations – G or A – are said to be the alleles for this specific position. SNPs pinpoint differences in our susceptibility to a wide range of diseases, for example age-related macular degeneration (a common SNP in the Factor H, CFH gene is associated with increased risk of the disease) or nonalcoholic fatty liver disease (a SNP in the PNPLA3 gene ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bioinformatics
Bioinformatics () is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combines biology, chemistry, physics, computer science, information engineering, mathematics and statistics to analyze and interpret the biological data. Bioinformatics has been used for '' in silico'' analyses of biological queries using computational and statistical techniques. Bioinformatics includes biological studies that use computer programming as part of their methodology, as well as specific analysis "pipelines" that are repeatedly used, particularly in the field of genomics. Common uses of bioinformatics include the identification of candidates genes and single nucleotide polymorphisms (SNPs). Often, such identification is made with the aim to better understand the genetic basis of disease, unique adaptations, desirable properties (e ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
International HapMap Project
The International HapMap Project was an organization that aimed to develop a haplotype map (HapMap) of the human genome, to describe the common patterns of human genetic variation. HapMap is used to find genetic variants affecting health, disease and responses to drugs and environmental factors. The information produced by the project is made freely available for research. The International HapMap Project is a collaboration among researchers at academic centers, non-profit biomedical research groups and private companies in Canada, China (including Hong Kong), Japan, Nigeria, the United Kingdom, and the United States. It officially started with a meeting on October 27 to 29, 2002, and was expected to take about three years. It comprises two phases; the complete data obtained in Phase I were published on 27 October 2005. The analysis of the Phase II dataset was published in October 2007. The Phase III dataset was released in spring 2009 and the publication presenting the final resul ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single Nucleotide Polymorphism
In genetics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently large fraction of the population (e.g. 1% or more), many publications do not apply such a frequency threshold. For example, at a specific base position in the human genome, the G nucleotide may appear in most individuals, but in a minority of individuals, the position is occupied by an A. This means that there is a SNP at this specific position, and the two possible nucleotide variations – G or A – are said to be the alleles for this specific position. SNPs pinpoint differences in our susceptibility to a wide range of diseases, for example age-related macular degeneration (a common SNP in the CFH gene is associated with increased risk of the disease) or nonalcoholic fatty liver disease (a SNP in the PNPLA3 gene is associated with incr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Variation
Genetic variation is the difference in DNA among individuals or the differences between populations. The multiple sources of genetic variation include mutation and genetic recombination. Mutations are the ultimate sources of genetic variation, but other mechanisms, such as genetic drift, contribute to it, as well. Among individuals within a population Genetic variation can be identified at many levels. Identifying genetic variation is possible from observations of phenotypic variation in either quantitative traits (traits that vary continuously and are coded for by many genes (e.g., leg length in dogs)) or discrete traits (traits that fall into discrete categories and are coded for by one or a few genes (e.g., white, pink, or red petal color in certain flowers)). Genetic variation can also be identified by examining variation at the level of enzymes using the process of protein electrophoresis. Polymorphic genes have more than one allele at each locus. Half of the genes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Species
In biology, a species is the basic unit of classification and a taxonomic rank of an organism, as well as a unit of biodiversity. A species is often defined as the largest group of organisms in which any two individuals of the appropriate sexes or mating types can produce fertile offspring, typically by sexual reproduction. Other ways of defining species include their karyotype, DNA sequence, morphology, behaviour or ecological niche. In addition, paleontologists use the concept of the chronospecies since fossil reproduction cannot be examined. The most recent rigorous estimate for the total number of species of eukaryotes is between 8 and 8.7 million. However, only about 14% of these had been described by 2011. All species (except viruses) are given a two-part name, a "binomial". The first part of a binomial is the genus to which the species belongs. The second part is called the specific name or the specific epithet (in botanical nomenclature, also sometimes i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Evolution
Evolution is change in the heritable characteristics of biological populations over successive generations. These characteristics are the expressions of genes, which are passed on from parent to offspring during reproduction. Variation tends to exist within any given population as a result of genetic mutation and recombination. Evolution occurs when evolutionary processes such as natural selection (including sexual selection) and genetic drift act on this variation, resulting in certain characteristics becoming more common or more rare within a population. The evolutionary pressures that determine whether a characteristic is common or rare within a population constantly change, resulting in a change in heritable characteristics arising over successive generations. It is this process of evolution that has given rise to biodiversity at every level of biological organisation, including the levels of species, individual organisms, and molecules. The theory of evolution by ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide Base
Nucleobases, also known as ''nitrogenous bases'' or often simply ''bases'', are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic building blocks of nucleic acids. The ability of nucleobases to form base pairs and to stack one upon another leads directly to long-chain helical structures such as ribonucleic acid (RNA) and deoxyribonucleic acid (DNA). Five nucleobases—adenine (A), cytosine (C), guanine (G), thymine (T), and uracil (U)—are called ''primary'' or ''canonical''. They function as the fundamental units of the genetic code, with the bases A, G, C, and T being found in DNA while A, G, C, and U are found in RNA. Thymine and uracil are distinguished by merely the presence or absence of a methyl group on the fifth carbon (C5) of these heterocyclic six-membered rings. In addition, some viruses have aminoadenine (Z) instead of adenine. It differs in having an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Sequence
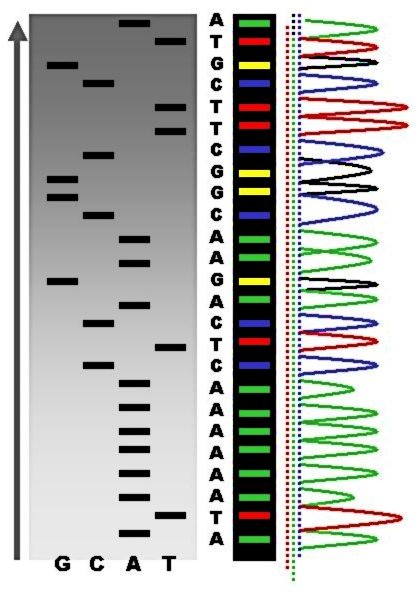
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, guanine, cytosine, and thymine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid speed of sequencing attained with modern D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
OMIM
Online Mendelian Inheritance in Man (OMIM) is a continuously updated catalog of human genes and genetic disorders and traits, with a particular focus on the gene-phenotype relationship. , approximately 9,000 of the over 25,000 entries in OMIM represented phenotypes; the rest represented genes, many of which were related to known phenotypes. Versions and history OMIM is the online continuation of Dr. Victor A. McKusick's ''Mendelian Inheritance in Man'' (MIM), which was published in 12 editions between 1966 and 1998.McKusick, V. A. ''Mendelian Inheritance in Man. Catalogs of Autosomal Dominant, Autosomal Recessive and X-Linked Phenotypes.'' Baltimore, MD: Johns Hopkins University Press, 1st ed, 1996; 2nd ed, 1969; 3rd ed, 1971; 4th ed, 1975; 5th ed, 1978; 6th ed, 1983; 7th ed, 1986; 8th ed, 1988; 9th ed, 1990; 10th ed, 1992. Nearly all of the 1,486 entries in the first edition of MIM discussed phenotypes. MIM/OMIM is produced and curated at the Johns Hopkins School of Medicine ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |