|
UvrABC
UvrABC endonuclease is a multienzyme complex in bacteria involved in DNA repair by nucleotide excision repair, and it is, therefore, sometimes called an excinuclease. This UvrABC repair process, sometimes called the short-patch process, involves the removal of twelve nucleotides where a genetic mutation has occurred followed by a DNA polymerase, replacing these aberrant nucleotides with the correct nucleotides and completing the DNA repair. The subunits for this enzyme are encoded in the ''uvrA'', ''uvrB'', and ''uvrC'' genes. This enzyme complex is able to repair many different types of damage, including cyclobutyl dimer formation. Mechanism # Two UvrA proteins form a dimer and they both have ATPase/GTPase activity. # The UvrA dimer binds with a UvrB dimer and forms a complex that is able to detect DNA damage. The UvrA dimer functions as the unit responsible for the detection of DNA damage, probably through a mechanism of detecting distortions in the DNA double helix. # Upon bi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide Excision Repair
Nucleotide excision repair is a DNA repair mechanism. DNA damage occurs constantly because of chemicals (e.g. intercalating agents), radiation and other mutagens. Three excision repair pathways exist to repair single stranded DNA damage: Nucleotide excision repair (NER), base excision repair (BER), and DNA mismatch repair (MMR). While the BER pathway can recognize specific non-bulky lesions in DNA, it can correct only damaged bases that are removed by specific glycosylases. Similarly, the MMR pathway only targets mismatched Watson-Crick base pairs. Nucleotide excision repair (NER) is a particularly important excision mechanism that removes DNA damage induced by ultraviolet light (UV). UV DNA damage results in bulky DNA adducts - these adducts are mostly thymine dimers and 6,4-photoproducts. Recognition of the damage leads to removal of a short single-stranded DNA segment that contains the lesion. The undamaged single-stranded DNA remains and DNA polymerase uses it as a templa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide Excision Repair
Nucleotide excision repair is a DNA repair mechanism. DNA damage occurs constantly because of chemicals (e.g. intercalating agents), radiation and other mutagens. Three excision repair pathways exist to repair single stranded DNA damage: Nucleotide excision repair (NER), base excision repair (BER), and DNA mismatch repair (MMR). While the BER pathway can recognize specific non-bulky lesions in DNA, it can correct only damaged bases that are removed by specific glycosylases. Similarly, the MMR pathway only targets mismatched Watson-Crick base pairs. Nucleotide excision repair (NER) is a particularly important excision mechanism that removes DNA damage induced by ultraviolet light (UV). UV DNA damage results in bulky DNA adducts - these adducts are mostly thymine dimers and 6,4-photoproducts. Recognition of the damage leads to removal of a short single-stranded DNA segment that contains the lesion. The undamaged single-stranded DNA remains and DNA polymerase uses it as a templa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Repair
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages (interstrand crosslinks or ICLs). This can eventually lead to malignant ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Damage
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages (interstrand crosslinks or ICLs). This can eventually lead to maligna ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separating two hybridized nucleic acid strands (hence '' helic- + -ase''), using energy from ATP hydrolysis. There are many helicases, representing the great variety of processes in which strand separation must be catalyzed. Approximately 1% of eukaryotic genes code for helicases. The human genome codes for 95 non-redundant helicases: 64 RNA helicases and 31 DNA helicases. Many cellular processes, such as DNA replication, transcription, translation, recombination, DNA repair, and ribosome biogenesis involve the separation of nucleic acid strands that necessitates the use of helicases. Some specialized helicases are also involved in sensing of viral nucleic acids during infection and fulfill a immunological function. Function Helicases are of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
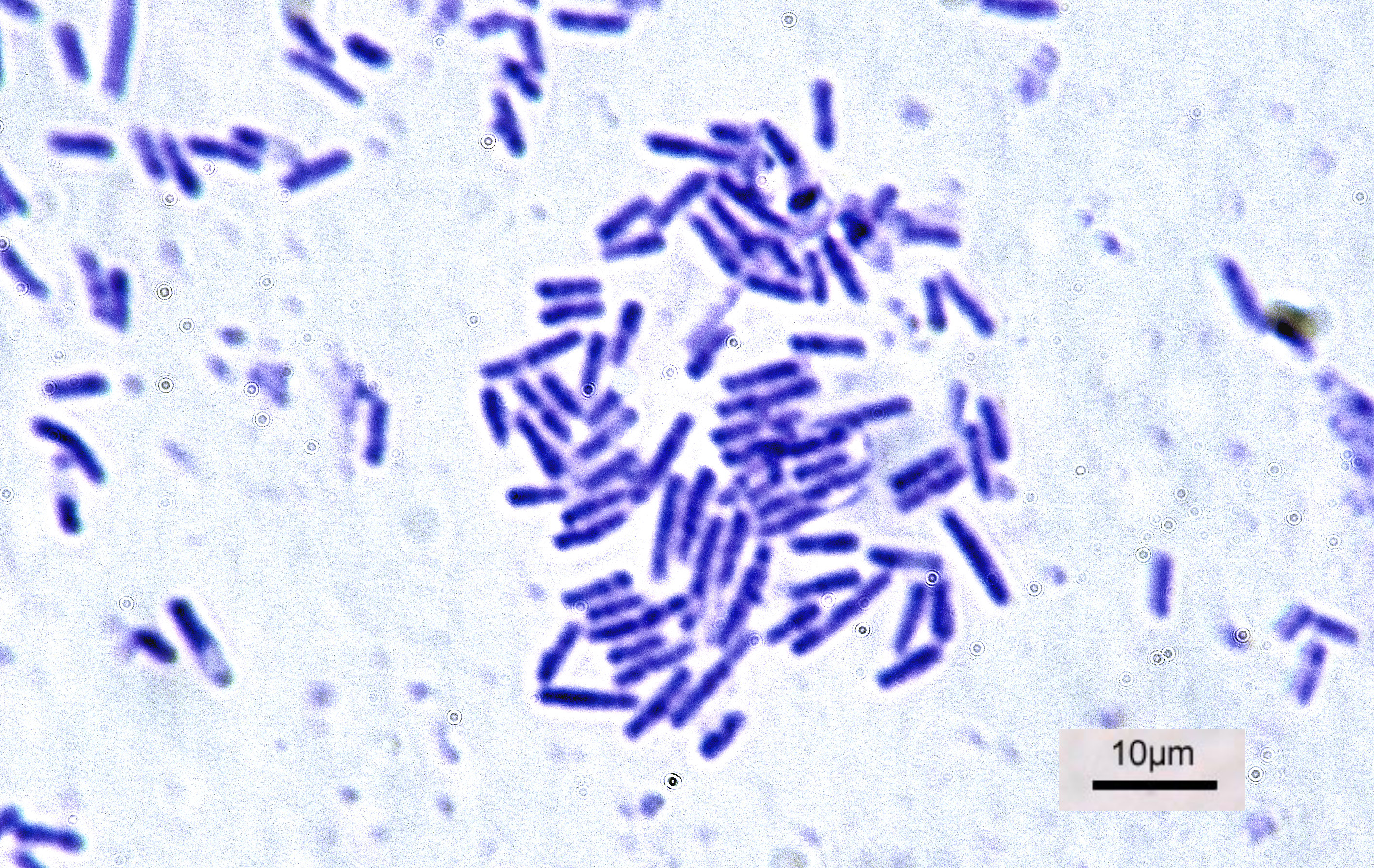
Bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit soil, water, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria are vital in many stages of the nutrient cycle by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. Bacteria also live in symbiotic and parasitic relationsh ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Excinuclease
Excision endonuclease, also known as excinuclease or UV-specific endonuclease, is a nuclease (enzyme) which excises a fragment of nucleotides during DNA repair. The excinuclease cuts out a fragment by hydrolyzing two phosphodiester bonds, one on either side of the lesion in the DNA. This process is part of "nucleotide excision repair", a mechanism that can fix specific types of damage to the DNA in the G1 phase of the eukaryotic cell cycle. Such damage may include thymine dimers created by UV rays as well as the bulky distortions in DNA caused by oxidized benzopyrenes from sources such as cigarette smoke. A deficiency of excinuclease occurs in a rare autosomal recessive disease called xeroderma pigmentosum. This disease can cause light-skin, extreme freckling and facial lesions, as well as preventing the repair of pyrimidine dimers. Diagnosis of this disease is done by measuring the enzyme's level in white blood cells in a blood sample. Symptoms in children include extreme UV s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cyclobutyl Dimer
Pyrimidine dimers are molecular lesions formed from thymine or cytosine bases in DNA via photochemical reactions, commonly associated with direct DNA damage. Ultraviolet light (UV; particularly UVB) induces the formation of covalent linkages between consecutive bases along the nucleotide chain in the vicinity of their carbon–carbon double bonds. The dimerization reaction can also occur among pyrimidine bases in dsRNA (double-stranded RNA)—uracil or cytosine. Two common UV products are cyclobutane pyrimidine dimers (CPDs) and 6–4 photoproducts. These premutagenic lesions alter the structure and possibly the base-pairing. Up to 50–100 such reactions per second might occur in a skin cell during exposure to sunlight, but are usually corrected within seconds by photolyase reactivation or nucleotide excision repair. Uncorrected lesions can inhibit polymerases, cause misreading during transcription or replication, or lead to arrest of replication. It causes sunburn and it trigger ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Dimer
In biochemistry, a protein dimer is a macromolecular complex formed by two protein monomers, or single proteins, which are usually non-covalently bound. Many macromolecules, such as proteins or nucleic acids, form dimers. The word ''dimer'' has roots meaning "two parts", '' di-'' + '' -mer''. A protein dimer is a type of protein quaternary structure. A protein homodimer is formed by two identical proteins. A protein heterodimer is formed by two different proteins. Most protein dimers in biochemistry are not connected by covalent bonds. An example of a non-covalent heterodimer is the enzyme reverse transcriptase, which is composed of two different amino acid chains. An exception is dimers that are linked by disulfide bridges such as the homodimeric protein NEMO. Some proteins contain specialized domains to ensure dimerization (dimerization domains) and specificity. The G protein-coupled cannabinoid receptors have the ability to form both homo- and heterodimers with several ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bond Cleavage
In chemistry, bond cleavage, or bond fission, is the splitting of chemical bonds. This can be generally referred to as dissociation when a molecule is cleaved into two or more fragments. In general, there are two classifications for bond cleavage: ''homo''lytic and ''hetero''lytic, depending on the nature of the process. The triplet and singlet excitation energies of a sigma bond can be used to determine if a bond will follow the homolytic or heterolytic pathway. A metal−metal sigma bond is an exception because the bond's excitation energy is extremely high, thus cannot be used for observation purposes. In some cases, bond cleavage requires catalysts. Due to the high bond-dissociation energy of C-H bonds, around , a large amount of energy is required to cleave the hydrogen atom from the carbon and bond a different atom to the carbon. Homolytic cleavage In homolytic cleavage, or homolysis, the two electrons in a cleaved covalent bond are divided equally between th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphodiester Bond
In chemistry, a phosphodiester bond occurs when exactly two of the hydroxyl groups () in phosphoric acid react with hydroxyl groups on other molecules to form two ester bonds. The "bond" involves this linkage . Discussion of phosphodiesters is dominated by their prevalence in DNA and RNA, but phosphodiesters occur in other biomolecules, e.g. acyl carrier proteins. Phosphodiester bonds make up the backbones of DNA and RNA. The phosphate is attached to the 5' carbon. The 3' carbon of one sugar is bonded to the 5' phosphate of the adjacent sugar. Specifically, the phosphodiester bond links the 3' carbon atom of one sugar molecule and the 5' carbon atom of another (hence the name, 3', 5' phosphodiester linkage). These saccharide groups are derived from deoxyribose in DNA and ribose in RNA. Phosphodiesters are negatively charged at pH 7. Repulsion between these negative charges influences the conformation of the polynucleic acids. The negative charge attracts histones, metal c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |