Bacteria on:
[Wikipedia]
[Google]
[Amazon]
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one
 The word ''bacteria'' is the plural of the
The word ''bacteria'' is the plural of the
 The ancestors of bacteria were unicellular microorganisms that were the first forms of life to appear on Earth, about 4 billion years ago. For about 3 billion years, most organisms were microscopic, and bacteria and archaea were the dominant forms of life. Although bacterial
The ancestors of bacteria were unicellular microorganisms that were the first forms of life to appear on Earth, about 4 billion years ago. For about 3 billion years, most organisms were microscopic, and bacteria and archaea were the dominant forms of life. Although bacterial
 Size. Bacteria display a wide diversity of shapes and sizes. Bacterial cells are about one-tenth the size of eukaryotic cells and are typically 0.5–5.0
Size. Bacteria display a wide diversity of shapes and sizes. Bacterial cells are about one-tenth the size of eukaryotic cells and are typically 0.5–5.0  Multicellularity. Most bacterial species exist as single cells; others associate in characteristic patterns: ''
Multicellularity. Most bacterial species exist as single cells; others associate in characteristic patterns: ''

 Bacteria do not have a membrane-bound nucleus, and their
Bacteria do not have a membrane-bound nucleus, and their
 Flagella are rigid protein structures, about 20 nanometres in diameter and up to 20 micrometres in length, that are used for
Flagella are rigid protein structures, about 20 nanometres in diameter and up to 20 micrometres in length, that are used for
 Some genera of Gram-positive bacteria, such as ''
Some genera of Gram-positive bacteria, such as ''

 Unlike in multicellular organisms, increases in cell size (cell growth) and reproduction by
Unlike in multicellular organisms, increases in cell size (cell growth) and reproduction by
 Most bacteria have a single circular
Most bacteria have a single circular
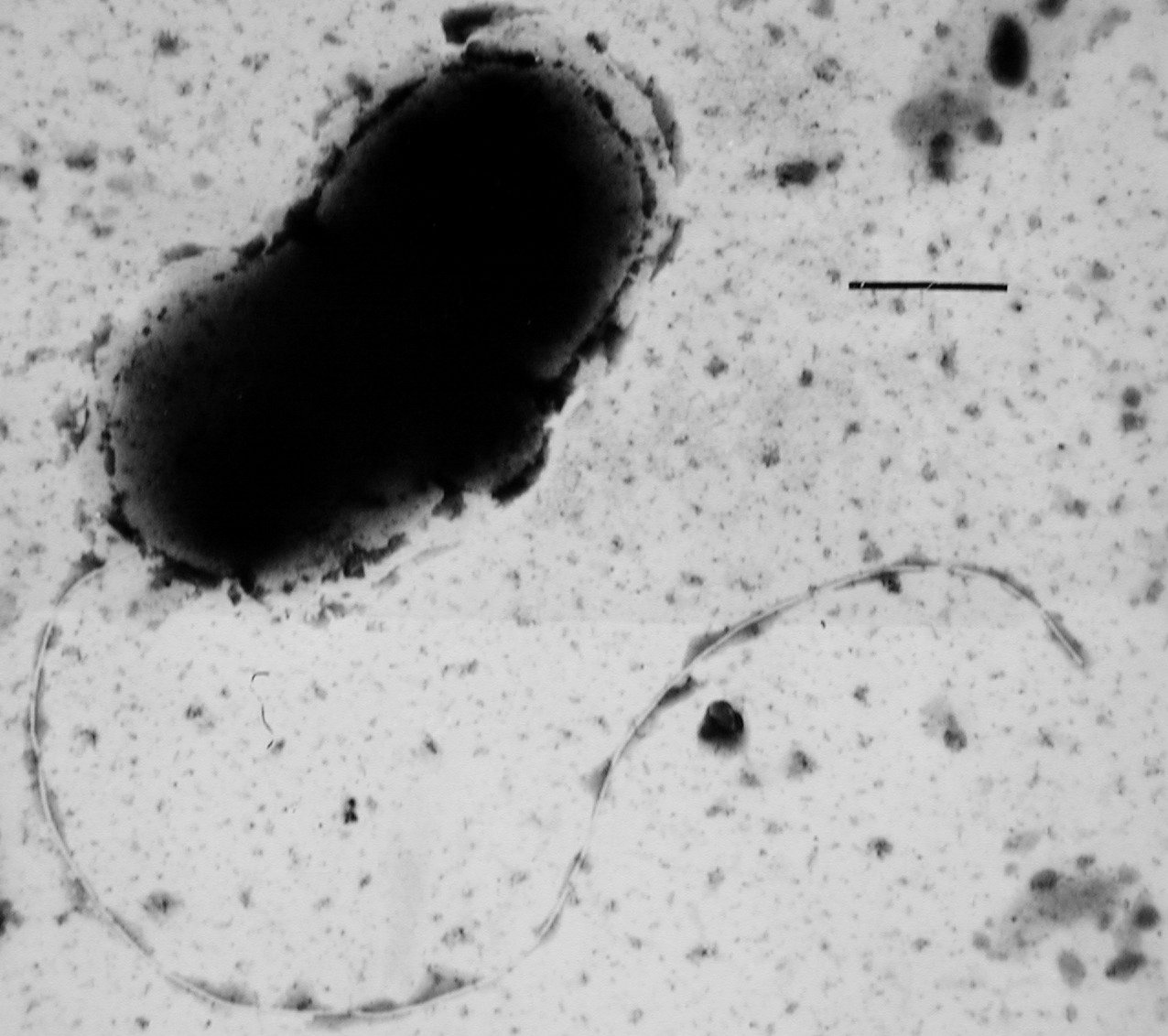 Many bacteria are Motility, motile (able to move themselves) and do so using a variety of mechanisms. The best studied of these are Flagellum, flagella, long filaments that are turned by a motor at the base to generate propeller-like movement. The bacterial flagellum is made of about 20 proteins, with approximately another 30 proteins required for its regulation and assembly. The flagellum is a rotating structure driven by a reversible motor at the base that uses the
Many bacteria are Motility, motile (able to move themselves) and do so using a variety of mechanisms. The best studied of these are Flagellum, flagella, long filaments that are turned by a motor at the base to generate propeller-like movement. The bacterial flagellum is made of about 20 proteins, with approximately another 30 proteins required for its regulation and assembly. The flagellum is a rotating structure driven by a reversible motor at the base that uses the  Bacteria can use flagella in different ways to generate different kinds of movement. Many bacteria (such as ''Escherichia coli, E. coli'') have two distinct modes of movement: forward movement (swimming) and tumbling. The tumbling allows them to reorient and makes their movement a three-dimensional random walk. Bacterial species differ in the number and arrangement of flagella on their surface; some have a single flagellum (''Flagellum#Flagellar arrangement schemes, monotrichous''), a flagellum at each end (''Flagellum#Flagellar arrangement schemes, amphitrichous''), clusters of flagella at the poles of the cell (''Flagellum#Flagellar arrangement schemes, lophotrichous''), while others have flagella distributed over the entire surface of the cell (''Flagellum#Flagellar arrangement schemes, peritrichous''). The flagella of a unique group of bacteria, the spirochaetes, are found between two membranes in the periplasmic space. They have a distinctive helix, helical body that twists about as it moves.
Two other types of bacterial motion are called twitching motility that relies on a structure called the pilus#Type IV pili, type IV pilus, and Bacterial gliding, gliding motility, that uses other mechanisms. In twitching motility, the rod-like pilus extends out from the cell, binds some substrate, and then retracts, pulling the cell forward.
Motile bacteria are attracted or repelled by certain stimulus (physiology), stimuli in behaviours called ''taxis, taxes'': these include chemotaxis, phototaxis, taxis, energy taxis, and magnetotaxis. In one peculiar group, the myxobacteria, individual bacteria move together to form waves of cells that then differentiate to form fruiting bodies containing spores. The myxobacteria move only when on solid surfaces, unlike ''E. coli'', which is motile in liquid or solid media.
Several ''Listeria'' and ''Shigella'' species move inside host cells by usurping the
Bacteria can use flagella in different ways to generate different kinds of movement. Many bacteria (such as ''Escherichia coli, E. coli'') have two distinct modes of movement: forward movement (swimming) and tumbling. The tumbling allows them to reorient and makes their movement a three-dimensional random walk. Bacterial species differ in the number and arrangement of flagella on their surface; some have a single flagellum (''Flagellum#Flagellar arrangement schemes, monotrichous''), a flagellum at each end (''Flagellum#Flagellar arrangement schemes, amphitrichous''), clusters of flagella at the poles of the cell (''Flagellum#Flagellar arrangement schemes, lophotrichous''), while others have flagella distributed over the entire surface of the cell (''Flagellum#Flagellar arrangement schemes, peritrichous''). The flagella of a unique group of bacteria, the spirochaetes, are found between two membranes in the periplasmic space. They have a distinctive helix, helical body that twists about as it moves.
Two other types of bacterial motion are called twitching motility that relies on a structure called the pilus#Type IV pili, type IV pilus, and Bacterial gliding, gliding motility, that uses other mechanisms. In twitching motility, the rod-like pilus extends out from the cell, binds some substrate, and then retracts, pulling the cell forward.
Motile bacteria are attracted or repelled by certain stimulus (physiology), stimuli in behaviours called ''taxis, taxes'': these include chemotaxis, phototaxis, taxis, energy taxis, and magnetotaxis. In one peculiar group, the myxobacteria, individual bacteria move together to form waves of cells that then differentiate to form fruiting bodies containing spores. The myxobacteria move only when on solid surfaces, unlike ''E. coli'', which is motile in liquid or solid media.
Several ''Listeria'' and ''Shigella'' species move inside host cells by usurping the
 Scientific classification, Classification seeks to describe the diversity of bacterial species by naming and grouping organisms based on similarities. Bacteria can be classified on the basis of cell structure, Cell metabolism, cellular metabolism or on differences in cell components, such as DNA, fatty acids, pigments,
Scientific classification, Classification seeks to describe the diversity of bacterial species by naming and grouping organisms based on similarities. Bacteria can be classified on the basis of cell structure, Cell metabolism, cellular metabolism or on differences in cell components, such as DNA, fatty acids, pigments,
 Despite their apparent simplicity, bacteria can form complex associations with other organisms. These symbiosis, symbiotic associations can be divided into parasitism, Mutualism (biology), mutualism and commensalism.
Despite their apparent simplicity, bacteria can form complex associations with other organisms. These symbiosis, symbiotic associations can be divided into parasitism, Mutualism (biology), mutualism and commensalism.
 The body is continually exposed to many species of bacteria, including beneficial commensals, which grow on the skin and mucous membranes, and saprophytes, which grow mainly in the soil and in decomposition, decaying matter. The blood and tissue fluids contain nutrients sufficient to sustain the growth of many bacteria. The body has defence mechanisms that enable it to resist microbial invasion of its tissues and give it a natural immune system, immunity or innate immunity, innate resistance against many microorganisms. Unlike some
The body is continually exposed to many species of bacteria, including beneficial commensals, which grow on the skin and mucous membranes, and saprophytes, which grow mainly in the soil and in decomposition, decaying matter. The blood and tissue fluids contain nutrients sufficient to sustain the growth of many bacteria. The body has defence mechanisms that enable it to resist microbial invasion of its tissues and give it a natural immune system, immunity or innate immunity, innate resistance against many microorganisms. Unlike some  Each species of pathogen has a characteristic spectrum of interactions with its human host (biology), hosts. Some organisms, such as ''Staphylococcus'' or ''Streptococcus'', can cause skin infections, pneumonia, meningitis and sepsis, a systemic Inflammation, inflammatory response producing shock (circulatory), shock, massive vasodilator, vasodilation and death. Yet these organisms are also part of the normal human flora and usually exist on the skin or in the nose without causing any disease at all. Other organisms invariably cause disease in humans, such as ''Rickettsia'', which are obligate intracellular parasites able to grow and reproduce only within the cells of other organisms. One species of ''Rickettsia'' causes typhus, while another causes Rocky Mountain spotted fever. ''Chlamydia (bacterium), Chlamydia'', another phylum of obligate intracellular parasites, contains species that can cause pneumonia or urinary tract infection and may be involved in coronary heart disease. Some species, such as ''
Each species of pathogen has a characteristic spectrum of interactions with its human host (biology), hosts. Some organisms, such as ''Staphylococcus'' or ''Streptococcus'', can cause skin infections, pneumonia, meningitis and sepsis, a systemic Inflammation, inflammatory response producing shock (circulatory), shock, massive vasodilator, vasodilation and death. Yet these organisms are also part of the normal human flora and usually exist on the skin or in the nose without causing any disease at all. Other organisms invariably cause disease in humans, such as ''Rickettsia'', which are obligate intracellular parasites able to grow and reproduce only within the cells of other organisms. One species of ''Rickettsia'' causes typhus, while another causes Rocky Mountain spotted fever. ''Chlamydia (bacterium), Chlamydia'', another phylum of obligate intracellular parasites, contains species that can cause pneumonia or urinary tract infection and may be involved in coronary heart disease. Some species, such as ''
On-line text book on bacteriology (2015)
{{Authority control Bacteria, Bacteriology Domains (biology)
biological cell
The cell is the basic structural and functional unit of life forms. Every cell consists of a cytoplasm enclosed within a membrane, and contains many biomolecules such as proteins, DNA and RNA, as well as many small molecules of nutrients an ...
. They constitute a large domain of prokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
microorganism
A microorganism, or microbe,, ''mikros'', "small") and ''organism'' from the el, ὀργανισμός, ''organismós'', "organism"). It is usually written as a single word but is sometimes hyphenated (''micro-organism''), especially in olde ...
s. Typically a few micrometre
The micrometre ( international spelling as used by the International Bureau of Weights and Measures; SI symbol: μm) or micrometer (American spelling), also commonly known as a micron, is a unit of length in the International System of Unit ...
s in length, bacteria were among the first life forms to appear on Earth
Earth is the third planet from the Sun and the only astronomical object known to harbor life. While large volumes of water can be found throughout the Solar System, only Earth sustains liquid surface water. About 71% of Earth's surfa ...
, and are present in most of its habitat
In ecology, the term habitat summarises the array of resources, physical and biotic factors that are present in an area, such as to support the survival and reproduction of a particular species. A species habitat can be seen as the physical ...
s. Bacteria inhabit soil, water, acidic hot springs, radioactive waste
Radioactive waste is a type of hazardous waste that contains radioactive material. Radioactive waste is a result of many activities, including nuclear medicine, nuclear research, nuclear power generation, rare-earth mining, and nuclear weapons r ...
, and the deep biosphere
The deep biosphere is the part of the biosphere that resides below the first few meters of the surface. It extends down at least 5 kilometers below the continental surface and 10.5 kilometers below the sea surface, at temperatures that ...
of Earth's crust. Bacteria are vital in many stages of the nutrient cycle
A nutrient cycle (or ecological recycling) is the movement and exchange of inorganic and organic matter back into the production of matter. Energy flow is a unidirectional and noncyclic pathway, whereas the movement of mineral nutrients is cyc ...
by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition
Decomposition or rot is the process by which dead organic substances are broken down into simpler organic or inorganic matter such as carbon dioxide, water, simple sugars and mineral salts. The process is a part of the nutrient cycle and is e ...
of dead bodies; bacteria are responsible for the putrefaction
Putrefaction is the fifth stage of death, following pallor mortis, algor mortis, rigor mortis, and livor mortis. This process references the breaking down of a body of an animal, such as a human, post-mortem. In broad terms, it can be view ...
stage in this process. In the biological communities surrounding hydrothermal vents and cold seep
A cold seep (sometimes called a cold vent) is an area of the ocean floor where hydrogen sulfide, methane and other hydrocarbon-rich fluid seepage occurs, often in the form of a brine pool. ''Cold'' does not mean that the temperature of the see ...
s, extremophile
An extremophile (from Latin ' meaning "extreme" and Greek ' () meaning "love") is an organism that is able to live (or in some cases thrive) in extreme environments, i.e. environments that make survival challenging such as due to extreme temper ...
bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane
Methane ( , ) is a chemical compound with the chemical formula (one carbon atom bonded to four hydrogen atoms). It is a group-14 hydride, the simplest alkane, and the main constituent of natural gas. The relative abundance of methane on Ea ...
, to energy. Bacteria also live in symbiotic and parasitic
Parasitism is a close relationship between species, where one organism, the parasite, lives on or inside another organism, the host, causing it some harm, and is adapted structurally to this way of life. The entomologist E. O. Wilson ha ...
relationships with plants and animals. Most bacteria have not been characterised and there are many species that cannot be grown in the laboratory. The study of bacteria is known as bacteriology Bacteriology is the branch and specialty of biology that studies the morphology, ecology, genetics and biochemistry of bacteria as well as many other aspects related to them. This subdivision of microbiology involves the identification, classificat ...
, a branch of microbiology.
Humans and most other animals carry millions of bacteria. Most are in the gut, and there are many on the skin. Most of the bacteria in and on the body are harmless or rendered so by the protective effects of the immune system
The immune system is a network of biological processes that protects an organism from diseases. It detects and responds to a wide variety of pathogens, from viruses to parasitic worms, as well as cancer cells and objects such as wood splint ...
, and many are beneficial, particularly the ones in the gut. However, several species of bacteria are pathogenic and cause infectious disease
An infection is the invasion of tissues by pathogens, their multiplication, and the reaction of host tissues to the infectious agent and the toxins they produce. An infectious disease, also known as a transmissible disease or communicable di ...
s, including cholera, syphilis, anthrax, leprosy
Leprosy, also known as Hansen's disease (HD), is a long-term infection by the bacteria ''Mycobacterium leprae'' or ''Mycobacterium lepromatosis''. Infection can lead to damage of the nerves, respiratory tract, skin, and eyes. This nerve damag ...
, tuberculosis
Tuberculosis (TB) is an infectious disease usually caused by '' Mycobacterium tuberculosis'' (MTB) bacteria. Tuberculosis generally affects the lungs, but it can also affect other parts of the body. Most infections show no symptoms, i ...
, tetanus
Tetanus, also known as lockjaw, is a bacterial infection caused by ''Clostridium tetani'', and is characterized by muscle spasms. In the most common type, the spasms begin in the jaw and then progress to the rest of the body. Each spasm usually ...
and bubonic plague. The most common fatal bacterial diseases are respiratory infection
Respiratory tract infections (RTIs) are infectious diseases involving the respiratory tract. An infection of this type usually is further classified as an upper respiratory tract infection (URI or URTI) or a lower respiratory tract infection (LRI ...
s. Antibiotics are used to treat bacterial infections
Pathogenic bacteria are bacteria that can cause disease. This article focuses on the bacteria that are pathogenic to humans. Most species of bacteria are harmless and are often beneficial but others can cause infectious diseases. The number of t ...
and are also used in farming, making antibiotic resistance a growing problem. Bacteria are important in sewage treatment
Sewage treatment (or domestic wastewater treatment, municipal wastewater treatment) is a type of wastewater treatment which aims to remove contaminants from sewage to produce an effluent that is suitable for discharge to the surrounding e ...
and the breakdown of oil spills, the production of cheese and yogurt
Yogurt (; , from tr, yoğurt, also spelled yoghurt, yogourt or yoghourt) is a food produced by bacterial fermentation of milk. The bacteria used to make yogurt are known as ''yogurt cultures''. Fermentation of sugars in the milk by these bac ...
through fermentation, the recovery of gold, palladium, copper and other metals in the mining sector, as well as in biotechnology
Biotechnology is the integration of natural sciences and engineering sciences in order to achieve the application of organisms, cells, parts thereof and molecular analogues for products and services. The term ''biotechnology'' was first used ...
, and the manufacture of antibiotics and other chemicals.
Once regarded as plant
Plants are predominantly photosynthetic eukaryotes of the kingdom Plantae. Historically, the plant kingdom encompassed all living things that were not animals, and included algae and fungi; however, all current definitions of Plantae exclu ...
s constituting the class ''Schizomycetes'' ("fission fungi"), bacteria are now classified as prokaryotes. Unlike cells of animals and other eukaryotes, bacterial cells do not contain a nucleus
Nucleus ( : nuclei) is a Latin word for the seed inside a fruit. It most often refers to:
*Atomic nucleus, the very dense central region of an atom
* Cell nucleus, a central organelle of a eukaryotic cell, containing most of the cell's DNA
Nucl ...
and rarely harbour membrane-bound organelles. Although the term ''bacteria'' traditionally included all prokaryotes, the scientific classification
Taxonomy is the practice and science of categorization or classification.
A taxonomy (or taxonomical classification) is a scheme of classification, especially a hierarchical classification, in which things are organized into groups or types. ...
changed after the discovery in the 1990s that prokaryotes consist of two very different groups of organisms that evolved
Evolution is change in the heritable characteristics of biological populations over successive generations. These characteristics are the expressions of genes, which are passed on from parent to offspring during reproduction. Variation ...
from an ancient common ancestor. These evolutionary domains are called Bacteria and Archaea.
Etymology
 The word ''bacteria'' is the plural of the
The word ''bacteria'' is the plural of the New Latin
New Latin (also called Neo-Latin or Modern Latin) is the revival of Literary Latin used in original, scholarly, and scientific works since about 1500. Modern scholarly and technical nomenclature, such as in zoological and botanical taxonomy ...
', which is the latinisation of the Ancient Greek
Ancient Greek includes the forms of the Greek language used in ancient Greece and the ancient world from around 1500 BC to 300 BC. It is often roughly divided into the following periods: Mycenaean Greek (), Dark Ages (), the Archaic p ...
('), the diminutive of ('), meaning "staff, cane", because the first ones to be discovered were rod-shaped
A bacillus (), also called a bacilliform bacterium or often just a rod (when the context makes the sense clear), is a rod-shaped bacterium or archaeon. Bacilli are found in many different taxonomic groups of bacteria. However, the name '' Baci ...
.
Origin and early evolution
fossil
A fossil (from Classical Latin , ) is any preserved remains, impression, or trace of any once-living thing from a past geological age. Examples include bones, shells, exoskeletons, stone imprints of animals or microbes, objects preserved ...
s exist, such as stromatolite
Stromatolites () or stromatoliths () are layered sedimentary formations ( microbialite) that are created mainly by photosynthetic microorganisms such as cyanobacteria, sulfate-reducing bacteria, and Pseudomonadota (formerly proteobacteria). T ...
s, their lack of distinctive morphology
Morphology, from the Greek and meaning "study of shape", may refer to:
Disciplines
* Morphology (archaeology), study of the shapes or forms of artifacts
* Morphology (astronomy), study of the shape of astronomical objects such as nebulae, galaxies ...
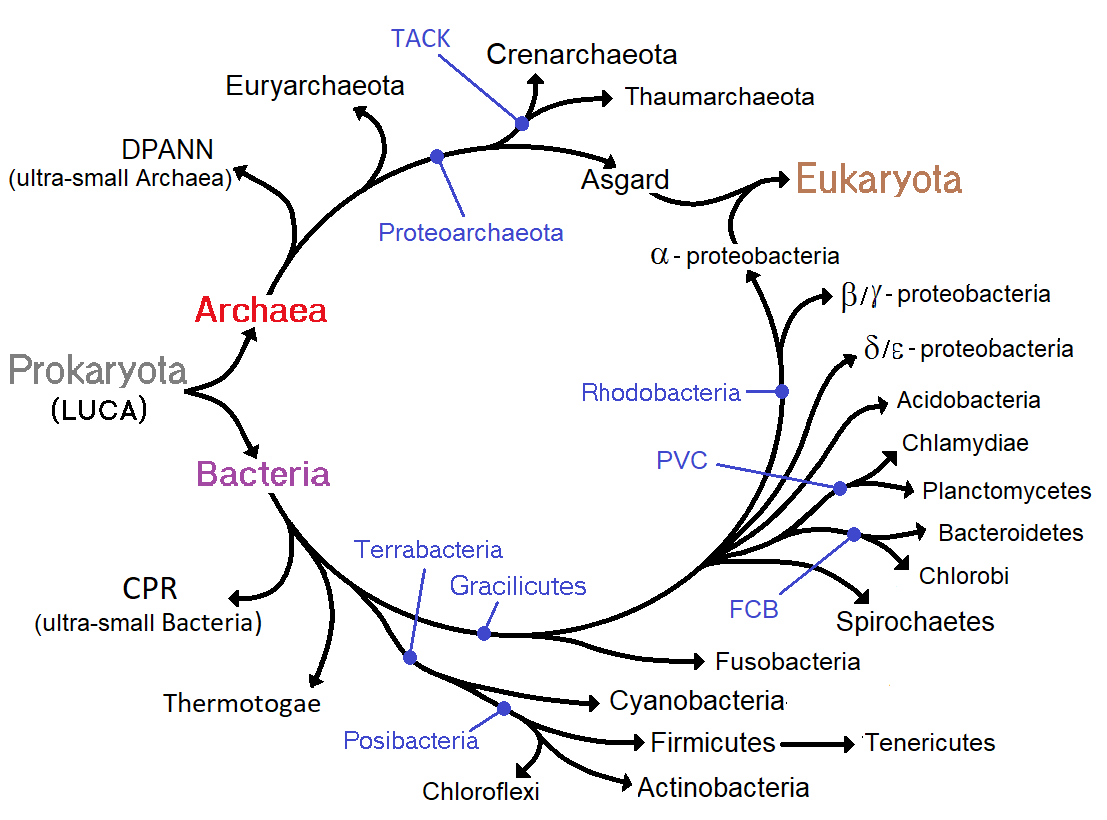
prevents them from being used to examine the history of bacterial evolution, or to date the time of origin of a particular bacterial species. However, gene sequences can be used to reconstruct the bacterial phylogeny
A phylogenetic tree (also phylogeny or evolutionary tree Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA.) is a branching diagram or a tree showing the evolutionary relationships among various biological spe ...
, and these studies indicate that bacteria diverged first from the archaeal/eukaryotic lineage. The most recent common ancestor
In biology and genetic genealogy, the most recent common ancestor (MRCA), also known as the last common ancestor (LCA) or concestor, of a set of organisms is the most recent individual from which all the organisms of the set are descended. The ...
of bacteria and archaea was probably a hyperthermophile
A hyperthermophile is an organism that thrives in extremely hot environments—from 60 °C (140 °F) upwards. An optimal temperature for the existence of hyperthermophiles is often above 80 °C (176 °F). Hyperthermophiles are often within the doma ...
that lived about 2.5 billion–3.2 billion years ago. The earliest life on land may have been bacteria some 3.22 billion years ago.
Bacteria were also involved in the second great evolutionary divergence, that of the archaea and eukaryotes. Here, eukaryotes resulted from the entering of ancient bacteria into endosymbiotic
An ''endosymbiont'' or ''endobiont'' is any organism that lives within the body or cells of another organism most often, though not always, in a mutualistic relationship.
(The term endosymbiosis is from the Greek: ἔνδον ''endon'' "within ...
associations with the ancestors of eukaryotic cells, which were themselves possibly related to the Archaea. This involved the engulfment by proto-eukaryotic cells of alphaproteobacteria
Alphaproteobacteria is a class of bacteria in the phylum Pseudomonadota (formerly Proteobacteria). The Magnetococcales and Mariprofundales are considered basal or sister to the Alphaproteobacteria. The Alphaproteobacteria are highly diverse and ...
l symbionts to form either mitochondria or hydrogenosome
A hydrogenosome is a membrane-enclosed organelle found in some anaerobic ciliates, flagellates, and fungi. Hydrogenosomes are highly variable organelles that have presumably evolved from protomitochondria to produce molecular hydrogen and ATP i ...
s, which are still found in all known Eukarya (sometimes in highly reduced form In statistics, and particularly in econometrics, the reduced form of a system of equations is the result of solving the system for the endogenous variables. This gives the latter as functions of the exogenous variables, if any. In econometrics, the ...
, e.g. in ancient "amitochondrial" protozoa). Later, some eukaryotes that already contained mitochondria also engulfed cyanobacteria-like organisms, leading to the formation of chloroplasts in algae and plants. This is known as primary endosymbiosis
Symbiogenesis (endosymbiotic theory, or serial endosymbiotic theory,) is the leading evolutionary theory of the origin of eukaryotic cells from prokaryotic organisms. The theory holds that mitochondria, plastids such as chloroplasts, and possi ...
.
Habitat
Bacteria are ubiquitous, living in every possible habitat on the planet including soil, underwater, deep in Earth's crust and even such extreme environments as acidic hot springs and radioactive waste. There are approximately 2×1030 bacteria on Earth, forming a biomass that is only exceeded by plants. They are abundant in lakes and oceans, in arctic ice, andgeothermal springs
A hot spring, hydrothermal spring, or geothermal spring is a spring produced by the emergence of geothermally heated groundwater onto the surface of the Earth. The groundwater is heated either by shallow bodies of magma (molten rock) or by cir ...
where they provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. They live on and in plants and animals. Most do not cause diseases, are beneficial to their environments, and are essential for life. The soil is a rich source of bacteria and a few grams contain around a thousand million of them. They are all essential to soil ecology, breaking down toxic waste and recycling nutrients. They are even found in the atmosphere and one cubic metre of air holds around one hundred million bacterial cells. The oceans and seas harbour around 3 x 1026 bacteria which provide up to 50% of the oxygen humans breathe. Only around 2% of bacterial species have been fully studied.
Morphology
micrometre
The micrometre ( international spelling as used by the International Bureau of Weights and Measures; SI symbol: μm) or micrometer (American spelling), also commonly known as a micron, is a unit of length in the International System of Unit ...
s in length. However, a few species are visible to the unaided eye—for example, ''Thiomargarita namibiensis
''Thiomargarita namibiensis'' is a Gram-negative coccoid bacterium, found in the ocean sediments of the continental shelf of Namibia. It is the second largest bacterium ever discovered, as a rule in diameter, but sometimes attaining . Cells of ...
'' is up to half a millimetre long, ''Epulopiscium fishelsoni
"''Candidatus'' Epulonipiscium" is a genus of Gram-positive bacteria that have a symbiotic relationship with surgeonfish. These bacteria are known for their unusually large size, many ranging from 200–700 μm in length. Until the discovery of ...
'' reaches 0.7 mm, and ''Thiomargarita magnifica
''Thiomargarita magnifica'' is a species of sulfur-oxidizing gammaproteobacteria, found growing underwater on the detached leaves of red mangroves from the Guadeloupe archipelago in the Lesser Antilles. This filament-shaped bacteria is the large ...
'' can reach even 2 cm in length, which is 50 times larger than other known bacteria. Among the smallest bacteria are members of the genus '' Mycoplasma'', which measure only 0.3 micrometres, as small as the largest virus
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsk ...
es. Some bacteria may be even smaller, but these ultramicrobacteria
Ultramicrobacteria are bacteria that are smaller than 0.1 μm3 under all growth conditions. This term was coined in 1981, describing cocci in seawater that were less than 0.3 μm in diameter. Ultramicrobacteria have also been recovered from soil a ...
are not well-studied.
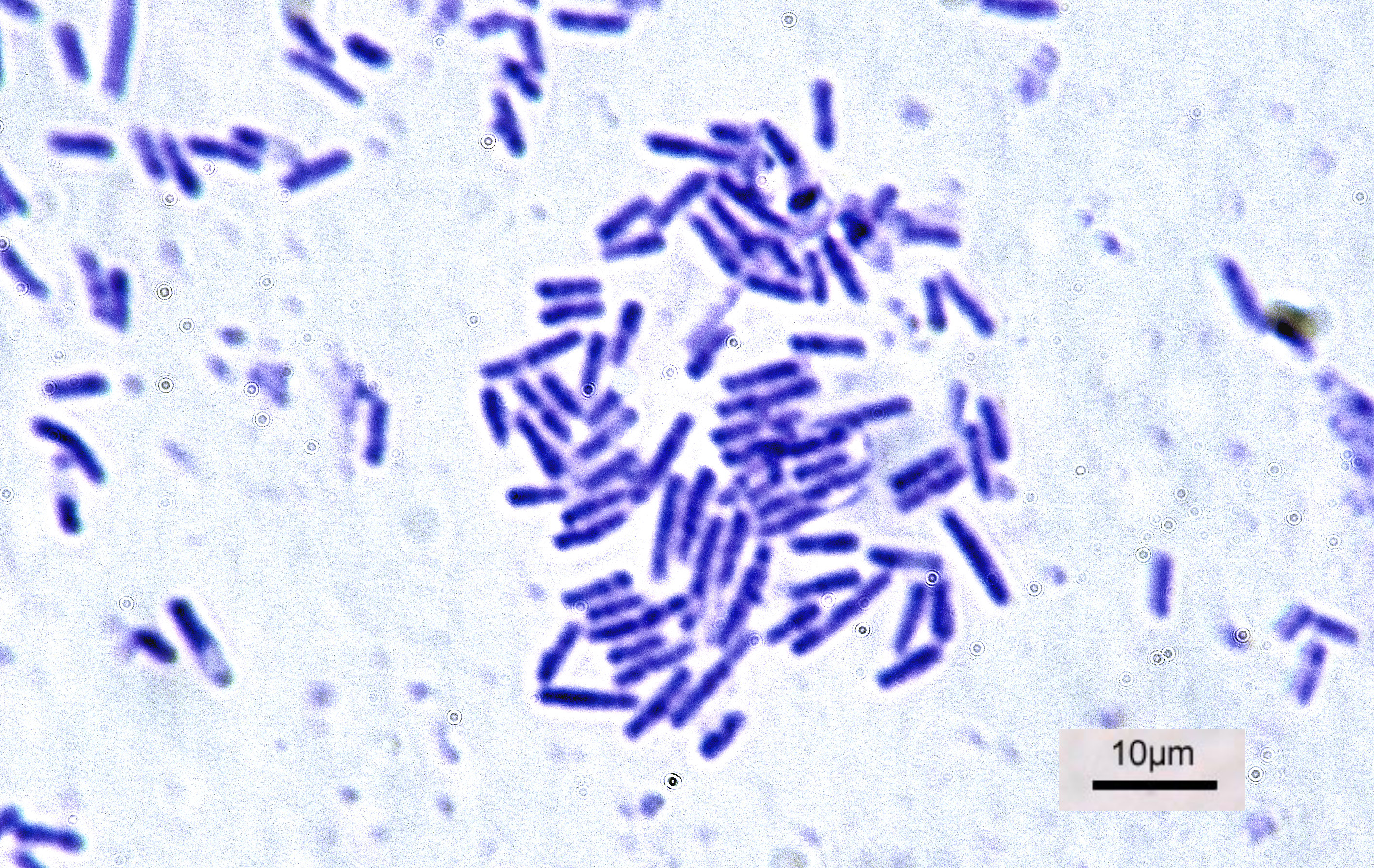
Shape. Most bacterial species are either spherical, called ''cocci
A coccus (plural cocci) is any bacterium or archaeon that has a spherical, ovoid, or generally round shape. Bacteria are categorized based on their shapes into three classes: cocci (spherical-shaped), bacillus (rod-shaped) and spiral ( of whi ...
'' (''singular coccus'', from Greek ''kókkos'', grain, seed), or rod shaped, called ''bacilli
Bacilli is a taxonomic class of bacteria that includes two orders, Bacillales and Lactobacillales, which contain several well-known pathogens such as ''Bacillus anthracis'' (the cause of anthrax). ''Bacilli'' are almost exclusively gram-positi ...
'' (''sing''. bacillus, from Latin
Latin (, or , ) is a classical language belonging to the Italic branch of the Indo-European languages. Latin was originally a dialect spoken in the lower Tiber area (then known as Latium) around present-day Rome, but through the power of the ...
''baculus'', stick). Some bacteria, called ''vibrio
''Vibrio'' is a genus of Gram-negative bacteria, possessing a curved-rod (comma) shape, several species of which can cause foodborne infection, usually associated with eating undercooked seafood. Being highly salt tolerant and unable to survive ...
'', are shaped like slightly curved rods or comma shaped; others can be spiral shaped, called '' spirilla'', or tightly coiled, called ''spirochaetes
A spirochaete () or spirochete is a member of the phylum Spirochaetota (), (synonym Spirochaetes) which contains distinctive diderm (double-membrane) gram-negative bacteria, most of which have long, helically coiled (corkscrew-shaped or s ...
''. A small number of other unusual shapes have been described, such as star-shaped bacteria. This wide variety of shapes is determined by the bacterial cell wall and cytoskeleton
The cytoskeleton is a complex, dynamic network of interlinking protein filaments present in the cytoplasm of all cells, including those of bacteria and archaea. In eukaryotes, it extends from the cell nucleus to the cell membrane and is com ...
, and is important because it can influence the ability of bacteria to acquire nutrients, attach to surfaces, swim through liquids and escape predators
Predation is a biological interaction where one organism, the predator, kills and eats another organism, its prey. It is one of a family of common feeding behaviours that includes parasitism and micropredation (which usually do not kill th ...
.
Neisseria
''Neisseria'' is a large genus of bacteria that colonize the mucosal surfaces of many animals. Of the 11 species that colonize humans, only two are pathogens, '' N. meningitidis'' and ''N. gonorrhoeae''.
''Neisseria'' species are Gram-negativ ...
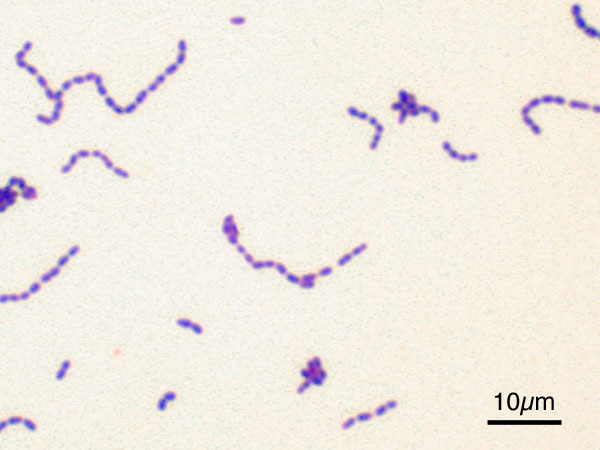
'' forms diploids (pairs), streptococci form chains, and staphylococci
''Staphylococcus'' is a genus of Gram-positive bacteria in the family Staphylococcaceae from the order Bacillales. Under the microscope, they appear spherical ( cocci), and form in grape-like clusters. ''Staphylococcus'' species are facultati ...
group together in "bunch of grapes" clusters. Bacteria can also group to form larger multicellular structures, such as the elongated filaments of ''Actinomycetota
The ''Actinomycetota'' (or ''Actinobacteria'') are a phylum of all gram-positive bacteria. They can be terrestrial or aquatic. They are of great economic importance to humans because agriculture and forests depend on their contributions to s ...
'' species, the aggregates of ''Myxobacteria
The myxobacteria ("slime bacteria") are a group of bacteria that predominantly live in the soil and feed on insoluble organic substances. The myxobacteria have very large genomes relative to other bacteria, e.g. 9–10 million nucleotides except ...
'' species, and the complex hyphae of '' Streptomyces'' species. These multicellular structures are often only seen in certain conditions. For example, when starved of amino acids, myxobacteria detect surrounding cells in a process known as quorum sensing
In biology, quorum sensing or quorum signalling (QS) is the ability to detect and respond to cell population density by gene regulation. As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at ...
, migrate towards each other, and aggregate to form fruiting bodies up to 500 micrometres long and containing approximately 100,000 bacterial cells. In these fruiting bodies, the bacteria perform separate tasks; for example, about one in ten cells migrate to the top of a fruiting body and differentiate into a specialised dormant state called a myxospore, which is more resistant to drying and other adverse environmental conditions.
Biofilms. Bacteria often attach to surfaces and form dense aggregations called biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
s, and larger formations known as microbial mat
A microbial mat is a multi-layered sheet of microorganisms, mainly bacteria and archaea, or bacteria alone. Microbial mats grow at interfaces between different types of material, mostly on submerged or moist surfaces, but a few survive in deserts ...
s. These biofilms and mats can range from a few micrometres in thickness to up to half a metre in depth, and may contain multiple species of bacteria, protist
A protist () is any eukaryotic organism (that is, an organism whose cells contain a cell nucleus) that is not an animal, plant, or fungus. While it is likely that protists share a common ancestor (the last eukaryotic common ancestor), the exc ...
s and archaea. Bacteria living in biofilms display a complex arrangement of cells and extracellular components, forming secondary structures, such as microcolonies, through which there are networks of channels to enable better diffusion of nutrients. In natural environments, such as soil or the surfaces of plants, the majority of bacteria are bound to surfaces in biofilms. Biofilms are also important in medicine, as these structures are often present during chronic bacterial infections or in infections of implanted medical device
A medical device is any device intended to be used for medical purposes. Significant potential for hazards are inherent when using a device for medical purposes and thus medical devices must be proved safe and effective with reasonable assura ...
s, and bacteria protected within biofilms are much harder to kill than individual isolated bacteria.
Cellular structure
Intracellular structures
The bacterial cell is surrounded by acell membrane
The cell membrane (also known as the plasma membrane (PM) or cytoplasmic membrane, and historically referred to as the plasmalemma) is a biological membrane that separates and protects the interior of all cells from the outside environment ( ...
, which is made primarily of phospholipids. This membrane encloses the contents of the cell and acts as a barrier to hold nutrients, protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, res ...
s and other essential components of the cytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
within the cell. Unlike eukaryotic cells
Eukaryotes () are organisms whose cells have a nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the three domains of life. Bact ...
, bacteria usually lack large membrane-bound structures in their cytoplasm such as a nucleus
Nucleus ( : nuclei) is a Latin word for the seed inside a fruit. It most often refers to:
*Atomic nucleus, the very dense central region of an atom
* Cell nucleus, a central organelle of a eukaryotic cell, containing most of the cell's DNA
Nucl ...
, mitochondria, chloroplasts and the other organelles present in eukaryotic cells. However, some bacteria have protein-bound organelles in the cytoplasm which compartmentalize aspects of bacterial metabolism, such as the carboxysome
Carboxysomes are bacterial microcompartments (BMCs) consisting of polyhedral protein shells filled with the enzymes ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO)—the predominant enzyme in carbon fixation and the rate limiting e ...
. Additionally, bacteria have a multi-component cytoskeleton
The cytoskeleton is a complex, dynamic network of interlinking protein filaments present in the cytoplasm of all cells, including those of bacteria and archaea. In eukaryotes, it extends from the cell nucleus to the cell membrane and is com ...
to control the localisation of proteins and nucleic acids within the cell, and to manage the process of cell division
Cell division is the process by which a parent cell divides into two daughter cells. Cell division usually occurs as part of a larger cell cycle in which the cell grows and replicates its chromosome(s) before dividing. In eukaryotes, there ar ...
.
Many important biochemical reactions, such as energy generation, occur due to concentration gradients across membranes, creating a potential
Potential generally refers to a currently unrealized ability. The term is used in a wide variety of fields, from physics to the social sciences to indicate things that are in a state where they are able to change in ways ranging from the simple r ...
difference analogous to a battery. The general lack of internal membranes in bacteria means these reactions, such as electron transport
An electron transport chain (ETC) is a series of protein complexes and other molecules that transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples thi ...
, occur across the cell membrane between the cytoplasm and the outside of the cell or periplasm
The periplasm is a concentrated gel-like matrix in the space between the inner cytoplasmic membrane and the bacterial outer membrane called the ''periplasmic space'' in gram-negative bacteria. Using cryo-electron microscopy it has been found that ...
. However, in many photosynthetic bacteria the plasma membrane is highly folded and fills most of the cell with layers of light-gathering membrane. These light-gathering complexes may even form lipid-enclosed structures called chlorosome
A chlorosome is a photosynthetic antenna complex found in green sulfur bacteria (GSB) and some green filamentous anoxygenic phototrophs (FAP) ( Chloroflexaceae, Oscillochloridaceae; both members of Chloroflexia). They differ from other antenna ...
s in green sulfur bacteria
The green sulfur bacteria are a phylum of obligately anaerobic photoautotrophic bacteria that metabolize sulfur.
Green sulfur bacteria are nonmotile (except ''Chloroherpeton thalassium'', which may glide) and capable of anoxygenic photosynthe ...
.
 Bacteria do not have a membrane-bound nucleus, and their
Bacteria do not have a membrane-bound nucleus, and their gene
In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b ...
tic material is typically a single circular bacterial chromosome
A circular chromosome is a chromosome in bacteria, archaea, mitochondria, and chloroplasts, in the form of a molecule of circular DNA, unlike the linear chromosome of most eukaryotes.
Most prokaryote chromosomes contain a circular DNA molecu ...
of DNA located in the cytoplasm in an irregularly shaped body called the nucleoid. The nucleoid contains the chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
with its associated proteins and RNA. Like all other organism
In biology, an organism () is any living system that functions as an individual entity. All organisms are composed of cells (cell theory). Organisms are classified by taxonomy into groups such as multicellular animals, plants, and ...
s, bacteria contain ribosomes for the production of proteins, but the structure of the bacterial ribosome is different from that of eukaryotes and archaea.
Some bacteria produce intracellular nutrient storage granules, such as glycogen, polyphosphate
Polyphosphates are salts or esters of polymeric oxyanions formed from tetrahedral PO4 (phosphate) structural units linked together by sharing oxygen atoms. Polyphosphates can adopt linear or a cyclic ring structures. In biology, the polyphosphate e ...
, sulfur or polyhydroxyalkanoates
Polyhydroxyalkanoates or PHAs are polyesters produced in nature by numerous microorganisms, including through bacterial fermentation of sugars or lipids. When produced by bacteria they serve as both a source of energy and as a carbon store. M ...
. Bacteria such as the photosynthetic cyanobacteria, produce internal gas vacuoles, which they use to regulate their buoyancy, allowing them to move up or down into water layers with different light intensities and nutrient levels.
Extracellular structures
Around the outside of the cell membrane is the cell wall. Bacterial cell walls are made ofpeptidoglycan
Peptidoglycan or murein is a unique large macromolecule, a polysaccharide, consisting of sugars and amino acids that forms a mesh-like peptidoglycan layer outside the plasma membrane, the rigid cell wall (murein sacculus) characteristic of most ba ...
(also called murein), which is made from polysaccharide chains cross-linked by peptide
Peptides (, ) are short chains of amino acids linked by peptide bonds. Long chains of amino acids are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides.
...
s containing D-amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha a ...
s. Bacterial cell walls are different from the cell walls of plant
Plants are predominantly photosynthetic eukaryotes of the kingdom Plantae. Historically, the plant kingdom encompassed all living things that were not animals, and included algae and fungi; however, all current definitions of Plantae exclu ...
s and fungi
A fungus ( : fungi or funguses) is any member of the group of eukaryotic organisms that includes microorganisms such as yeasts and molds, as well as the more familiar mushrooms. These organisms are classified as a kingdom, separately from ...
, which are made of cellulose
Cellulose is an organic compound with the formula , a polysaccharide consisting of a linear chain of several hundred to many thousands of β(1→4) linked D-glucose units. Cellulose is an important structural component of the primary cell w ...
and chitin, respectively. The cell wall of bacteria is also distinct from that of achaea, which do not contain peptidoglycan. The cell wall is essential to the survival of many bacteria, and the antibiotic penicillin (produced by a fungus called ''Penicillium
''Penicillium'' () is a genus of ascomycetous fungi that is part of the mycobiome of many species and is of major importance in the natural environment, in food spoilage, and in food and drug production.
Some members of the genus produce pe ...
'') is able to kill bacteria by inhibiting a step in the synthesis of peptidoglycan.
There are broadly speaking two different types of cell wall in bacteria, that classify bacteria into Gram-positive bacteria and Gram-negative bacteria
Gram-negative bacteria are bacteria that do not retain the crystal violet stain used in the Gram staining method of bacterial differentiation. They are characterized by their cell envelopes, which are composed of a thin peptidoglycan cell wall ...
. The names originate from the reaction of cells to the Gram stain
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish b ...
, a long-standing test for the classification of bacterial species.
Gram-positive bacteria possess a thick cell wall containing many layers of peptidoglycan and teichoic acid
Teichoic acids (''cf.'' Greek τεῖχος, ''teīkhos'', "wall", to be specific a fortification wall, as opposed to τοῖχος, ''toīkhos'', a regular wall) are bacterial copolymers of glycerol phosphate or ribitol phosphate and carbohydr ...
s. In contrast, Gram-negative bacteria have a relatively thin cell wall consisting of a few layers of peptidoglycan surrounded by a second lipid membrane
The lipid bilayer (or phospholipid bilayer) is a thin polar membrane made of two layers of lipid molecules. These membranes are flat sheets that form a continuous barrier around all cells. The cell membranes of almost all organisms and many vir ...
containing lipopolysaccharide
Lipopolysaccharides (LPS) are large molecules consisting of a lipid and a polysaccharide that are bacterial toxins. They are composed of an O-antigen, an outer core, and an inner core all joined by a covalent bond, and are found in the outer ...
s and lipoprotein
A lipoprotein is a biochemical assembly whose primary function is to transport hydrophobic lipid (also known as fat) molecules in water, as in blood plasma or other extracellular fluids. They consist of a triglyceride and cholesterol center, su ...
s. Most bacteria have the Gram-negative cell wall, and only members of the ''Bacillota
The Bacillota (synonym Firmicutes) are a phylum of bacteria, most of which have gram-positive cell wall structure. The renaming of phyla such as Firmicutes in 2021 remains controversial among microbiologists, many of whom continue to use the earl ...
'' group and actinomycetota
The ''Actinomycetota'' (or ''Actinobacteria'') are a phylum of all gram-positive bacteria. They can be terrestrial or aquatic. They are of great economic importance to humans because agriculture and forests depend on their contributions to s ...
(previously known as the low G+C and high G+C Gram-positive bacteria, respectively) have the alternative Gram-positive arrangement. These differences in structure can produce differences in antibiotic susceptibility; for instance, vancomycin
Vancomycin is a glycopeptide antibiotic medication used to treat a number of bacterial infections. It is recommended intravenously as a treatment for complicated skin infections, bloodstream infections, endocarditis, bone and joint infections, ...
can kill only Gram-positive bacteria and is ineffective against Gram-negative pathogen
In biology, a pathogen ( el, πάθος, "suffering", "passion" and , "producer of") in the oldest and broadest sense, is any organism or agent that can produce disease. A pathogen may also be referred to as an infectious agent, or simply a germ ...
s, such as ''Haemophilus influenzae
''Haemophilus influenzae'' (formerly called Pfeiffer's bacillus or ''Bacillus influenzae'') is a Gram-negative, non-motile, coccobacillary, facultatively anaerobic, capnophilic pathogenic bacterium of the family Pasteurellaceae. The bacter ...
'' or ''Pseudomonas aeruginosa
''Pseudomonas aeruginosa'' is a common encapsulated, gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, ''P. aerug ...
''. Some bacteria have cell wall structures that are neither classically Gram-positive or Gram-negative. This includes clinically important bacteria such as mycobacteria which have a thick peptidoglycan cell wall like a Gram-positive bacterium, but also a second outer layer of lipids.
In many bacteria, an S-layer An S-layer (surface layer) is a part of the cell envelope found in almost all archaea, as well as in many types of bacteria.
The S-layers of both archaea and bacteria consists of a monomolecular layer composed of only one (or, in a few cases, two) ...
of rigidly arrayed protein molecules covers the outside of the cell. This layer provides chemical and physical protection for the cell surface and can act as a macromolecular
A macromolecule is a very large molecule important to biophysical processes, such as a protein or nucleic acid. It is composed of thousands of covalently bonded atoms. Many macromolecules are polymers of smaller molecules called monomers. The ...
diffusion barrier. S-layers have diverse functions and are known to act as virulence factors in '' Campylobacter'' species and contain surface enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
s in '' Bacillus stearothermophilus''.
 Flagella are rigid protein structures, about 20 nanometres in diameter and up to 20 micrometres in length, that are used for
Flagella are rigid protein structures, about 20 nanometres in diameter and up to 20 micrometres in length, that are used for motility
Motility is the ability of an organism to move independently, using metabolic energy.
Definitions
Motility, the ability of an organism to move independently, using metabolic energy, can be contrasted with sessility, the state of organisms th ...
. Flagella are driven by the energy released by the transfer of ion
An ion () is an atom or molecule with a net electrical charge.
The charge of an electron is considered to be negative by convention and this charge is equal and opposite to the charge of a proton, which is considered to be positive by conve ...
s down an electrochemical gradient
An electrochemical gradient is a gradient of electrochemical potential, usually for an ion that can move across a membrane. The gradient consists of two parts, the chemical gradient, or difference in solute concentration across a membrane, and ...
across the cell membrane.
Fimbriae (sometimes called " attachment pili") are fine filaments of protein, usually 2–10 nanometres in diameter and up to several micrometres in length. They are distributed over the surface of the cell, and resemble fine hairs when seen under the electron microscope
An electron microscope is a microscope that uses a beam of accelerated electrons as a source of illumination. As the wavelength of an electron can be up to 100,000 times shorter than that of visible light photons, electron microscopes have a hi ...
. Fimbriae are believed to be involved in attachment to solid surfaces or to other cells, and are essential for the virulence of some bacterial pathogens. Pili (''sing''. pilus) are cellular appendages, slightly larger than fimbriae, that can transfer genetic material
Nucleic acids are biopolymers, macromolecules, essential to all known forms of life. They are composed of nucleotides, which are the monomers made of three components: a 5-carbon sugar, a phosphate group and a nitrogenous base. The two main clas ...
between bacterial cells in a process called conjugation
Conjugation or conjugate may refer to:
Linguistics
* Grammatical conjugation, the modification of a verb from its basic form
* Emotive conjugation or Russell's conjugation, the use of loaded language
Mathematics
* Complex conjugation, the chang ...
where they are called conjugation pili or sex pili (see bacterial genetics, below). They can also generate movement where they are called type IV pili
A pilus (Latin for 'hair'; plural: ''pili'') is a hair-like appendage found on the surface of many bacteria and archaea. The terms ''pilus'' and '' fimbria'' (Latin for 'fringe'; plural: ''fimbriae'') can be used interchangeably, although some r ...
.
Glycocalyx
The glycocalyx, also known as the pericellular matrix, is a glycoprotein and glycolipid covering that surrounds the cell membranes of bacteria, epithelial cells, and other cells. In 1970, Martinez-Palomo discovered the cell coating in animal c ...
is produced by many bacteria to surround their cells, and varies in structural complexity: ranging from a disorganised slime layer of extracellular polymeric substances to a highly structured capsule. These structures can protect cells from engulfment by eukaryotic cells such as macrophages (part of the human immune system
The immune system is a network of biological processes that protects an organism from diseases. It detects and responds to a wide variety of pathogens, from viruses to parasitic worms, as well as cancer cells and objects such as wood splint ...
). They can also act as antigen
In immunology, an antigen (Ag) is a molecule or molecular structure or any foreign particulate matter or a pollen grain that can bind to a specific antibody or T-cell receptor. The presence of antigens in the body may trigger an immune respons ...
s and be involved in cell recognition, as well as aiding attachment to surfaces and the formation of biofilms.
The assembly of these extracellular structures is dependent on bacterial secretion systems. These transfer proteins from the cytoplasm into the periplasm or into the environment around the cell. Many types of secretion systems are known and these structures are often essential for the virulence
Virulence is a pathogen's or microorganism's ability to cause damage to a host.
In most, especially in animal systems, virulence refers to the degree of damage caused by a microbe to its host. The pathogenicity of an organism—its ability to ...
of pathogens, so are intensively studied.
Endospores
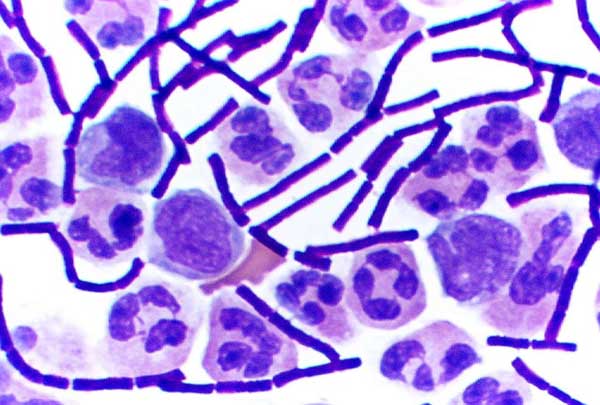
 Some genera of Gram-positive bacteria, such as ''
Some genera of Gram-positive bacteria, such as ''Bacillus
''Bacillus'' (Latin "stick") is a genus of Gram-positive, rod-shaped bacteria, a member of the phylum '' Bacillota'', with 266 named species. The term is also used to describe the shape (rod) of other so-shaped bacteria; and the plural ''Bacill ...
'', '' Clostridium'', ''Sporohalobacter
''Sporohalobacter'' are a genus of anaerobic bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. ...
'', '' Anaerobacter'', and '' Heliobacterium'', can form highly resistant, dormant structures called '' endospores''. Endospores develop within the cytoplasm of the cell; generally a single endospore develops in each cell. Each endospore contains a core of DNA and ribosomes surrounded by a cortex layer and protected by a multilayer rigid coat composed of peptidoglycan and a variety of proteins.
Endospores show no detectable metabolism
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of life-sustaining chemical reactions in organisms. The three main functions of metabolism are: the conversion of the energy in food to energy available to run c ...
and can survive extreme physical and chemical stresses, such as high levels of UV light
Ultraviolet (UV) is a form of electromagnetic radiation with wavelength from 10 nm (with a corresponding frequency around 30 PHz) to 400 nm (750 THz), shorter than that of visible light, but longer than X-rays. UV radiation i ...
, gamma radiation, detergents, disinfectants, heat, freezing, pressure, and desiccation. In this dormant state, these organisms may remain viable for millions of years, and endospores even allow bacteria to survive exposure to the vacuum
A vacuum is a space devoid of matter. The word is derived from the Latin adjective ''vacuus'' for "vacant" or " void". An approximation to such vacuum is a region with a gaseous pressure much less than atmospheric pressure. Physicists often di ...
and radiation in space, possibly bacteria could be distributed throughout the Universe
The universe is all of space and time and their contents, including planets, stars, galaxies, and all other forms of matter and energy. The Big Bang theory is the prevailing cosmological description of the development of the universe. ...
by space dust
Cosmic dust, also called extraterrestrial dust, star dust or space dust, is dust which exists in outer space, or has fallen on Earth. Most cosmic dust particles measure between a few molecules and 0.1 mm (100 micrometers). Larger particles are c ...
, meteoroid
A meteoroid () is a small rocky or metallic body in outer space.
Meteoroids are defined as objects significantly smaller than asteroids, ranging in size from grains to objects up to a meter wide. Objects smaller than this are classified as mi ...
s, asteroids
An asteroid is a minor planet of the inner Solar System. Sizes and shapes of asteroids vary significantly, ranging from 1-meter rocks to a dwarf planet almost 1000 km in diameter; they are rocky, metallic or icy bodies with no atmosphere. ...
, comets
A comet is an icy, small Solar System body that, when passing close to the Sun, warms and begins to release gases, a process that is called outgassing. This produces a visible atmosphere or coma, and sometimes also a tail. These phenomena ar ...
, planetoids
According to the International Astronomical Union (IAU), a minor planet is an astronomical object in direct orbit around the Sun that is exclusively classified as neither a planet nor a comet. Before 2006, the IAU officially used the term ''mino ...
or via directed panspermia
Directed panspermia is the deliberate transport of microorganisms into space to be used as introduced species on lifeless but habitable astronomical objects.
Historically, Shklovskii and Sagan (1966) and Crick and Orgel (1973) hypothesized that li ...
. Endospore-forming bacteria can also cause disease: for example, anthrax can be contracted by the inhalation of '' Bacillus anthracis'' endospores, and contamination of deep puncture wounds with ''Clostridium tetani
''Clostridium tetani'' is a common soil bacterium and the causative agent of tetanus. Vegetative cells of ''Clostridium tetani'' are usually rod-shaped and up to 2.5 μm long, but they become enlarged and tennis racket- or drumstick-shaped wh ...
'' endospores causes tetanus
Tetanus, also known as lockjaw, is a bacterial infection caused by ''Clostridium tetani'', and is characterized by muscle spasms. In the most common type, the spasms begin in the jaw and then progress to the rest of the body. Each spasm usually ...
, which like botulism
Botulism is a rare and potentially fatal illness caused by a toxin produced by the bacterium ''Clostridium botulinum''. The disease begins with weakness, blurred vision, feeling tired, and trouble speaking. This may then be followed by weakne ...
is caused by a toxin released by the bacteria that grow from the spores. Clostridioides difficile infection
''Clostridioides difficile'' infection
(CDI or C-diff), also known as ''Clostridium difficile'' infection, is a symptomatic infection due to the spore-forming bacterium '' Clostridioides difficile''. Symptoms include watery diarrhea, fever, n ...
, which is a problem in healthcare settings is also caused by spore-forming bacteria.
Metabolism
Bacteria exhibit an extremely wide variety of metabolic types. The distribution of metabolic traits within a group of bacteria has traditionally been used to define theirtaxonomy
Taxonomy is the practice and science of categorization or classification.
A taxonomy (or taxonomical classification) is a scheme of classification, especially a hierarchical classification, in which things are organized into groups or types. ...
, but these traits often do not correspond with modern genetic classifications. Bacterial metabolism is classified into nutritional groups on the basis of three major criteria: the source of energy
In physics, energy (from Ancient Greek: ἐνέργεια, ''enérgeia'', “activity”) is the quantitative property that is transferred to a body or to a physical system, recognizable in the performance of work and in the form of hea ...
, the electron donor
In chemistry, an electron donor is a chemical entity that donates electrons to another compound. It is a reducing agent that, by virtue of its donating electrons, is itself oxidized in the process.
Typical reducing agents undergo permanent chemi ...
s used, and the source of carbon
Carbon () is a chemical element with the symbol C and atomic number 6. It is nonmetallic and tetravalent—its atom making four electrons available to form covalent chemical bonds. It belongs to group 14 of the periodic table. Carbon mak ...
used for growth.
Bacteria either derive energy from light using photosynthesis
Photosynthesis is a process used by plants and other organisms to convert light energy into chemical energy that, through cellular respiration, can later be released to fuel the organism's activities. Some of this chemical energy is stored i ...
(called phototrophy
Phototrophs () are organisms that carry out photon capture to produce complex organic compounds (e.g. carbohydrates) and acquire energy. They use the energy from light to carry out various cellular metabolic processes. It is a common misconcep ...
), or by breaking down chemical compounds using oxidation
Redox (reduction–oxidation, , ) is a type of chemical reaction in which the oxidation states of substrate change. Oxidation is the loss of electrons or an increase in the oxidation state, while reduction is the gain of electrons or a ...
(called chemotroph
A Chemotroph is an organism that obtains energy by the oxidation of electron donors in their environments. These molecules can be organic (chemoorganotrophs) or inorganic ( chemolithotrophs). The chemotroph designation is in contrast to phototr ...
y). Chemotrophs use chemical compounds as a source of energy by transferring electrons from a given electron donor to a terminal electron acceptor
An electron acceptor is a chemical entity that accepts electrons transferred to it from another compound. It is an oxidizing agent that, by virtue of its accepting electrons, is itself reduced in the process. Electron acceptors are sometimes mista ...
in a redox reaction
Redox (reduction–oxidation, , ) is a type of chemical reaction in which the oxidation states of substrate change. Oxidation is the loss of electrons or an increase in the oxidation state, while reduction is the gain of electrons or a d ...
. This reaction releases energy that can be used to drive metabolism. Chemotrophs are further divided by the types of compounds they use to transfer electrons. Bacteria that use inorganic compounds such as hydrogen, carbon monoxide
Carbon monoxide (chemical formula CO) is a colorless, poisonous, odorless, tasteless, flammable gas that is slightly less dense than air. Carbon monoxide consists of one carbon atom and one oxygen atom connected by a triple bond. It is the simple ...
, or ammonia
Ammonia is an inorganic compound of nitrogen and hydrogen with the formula . A stable binary hydride, and the simplest pnictogen hydride, ammonia is a colourless gas with a distinct pungent smell. Biologically, it is a common nitrogenous wa ...
as sources of electrons are called lithotroph
Lithotrophs are a diverse group of organisms using an inorganic substrate (usually of mineral origin) to obtain reducing equivalents for use in biosynthesis (e.g., carbon dioxide fixation) or energy conservation (i.e., ATP production) via aerobi ...
s, while those that use organic compounds are called organotroph
An organotroph is an organism that obtains hydrogen or electrons from organic substrates. This term is used in microbiology to classify and describe organisms based on how they obtain electrons for their respiration processes. Some organotrophs su ...
s. The compounds used to receive electrons are also used to classify bacteria: aerobic organism
Aerobic means "requiring air," in which "air" usually means oxygen.
Aerobic may also refer to
* Aerobic exercise, prolonged exercise of moderate intensity
* Aerobics
Aerobics is a form of physical exercise that combines rhythmic aerobic exe ...
s use oxygen
Oxygen is the chemical element with the symbol O and atomic number 8. It is a member of the chalcogen group in the periodic table, a highly reactive nonmetal, and an oxidizing agent that readily forms oxides with most elements as ...
as the terminal electron acceptor, while anaerobic organisms use other compounds such as nitrate, sulfate
The sulfate or sulphate ion is a polyatomic anion with the empirical formula . Salts, acid derivatives, and peroxides of sulfate are widely used in industry. Sulfates occur widely in everyday life. Sulfates are salts of sulfuric acid and many ...
, or carbon dioxide.
Many bacteria get their carbon from other organic carbon
Total organic carbon (TOC) is the amount of carbon found in an organic compound and is often used as a non-specific indicator of water quality or cleanliness of pharmaceutical manufacturing equipment. TOC may also refer to the amount of organic c ...
, called heterotroph
A heterotroph (; ) is an organism that cannot produce its own food, instead taking nutrition from other sources of organic carbon, mainly plant or animal matter. In the food chain, heterotrophs are primary, secondary and tertiary consumers, but ...
y. Others such as cyanobacteria and some purple bacteria
Purple bacteria or purple photosynthetic bacteria are Gram-negative proteobacteria that are phototrophic, capable of producing their own food via photosynthesis. They are pigmented with bacteriochlorophyll ''a'' or ''b'', together with various ...
are autotroph
An autotroph or primary producer is an organism that produces complex organic compounds (such as carbohydrates, fats, and proteins) using carbon from simple substances such as carbon dioxide,Morris, J. et al. (2019). "Biology: How Life Wo ...
ic, meaning that they obtain cellular carbon by fixing
Fixing may refer to:
* The present participle of the verb "to fix", an action meaning maintenance, repair, and operations
* "fixing someone up" in the context of arranging or finding a social date for someone
* "Fixing", craving an addictive drug, ...
carbon dioxide
Carbon dioxide ( chemical formula ) is a chemical compound made up of molecules that each have one carbon atom covalently double bonded to two oxygen atoms. It is found in the gas state at room temperature. In the air, carbon dioxide is trans ...
. In unusual circumstances, the gas methane
Methane ( , ) is a chemical compound with the chemical formula (one carbon atom bonded to four hydrogen atoms). It is a group-14 hydride, the simplest alkane, and the main constituent of natural gas. The relative abundance of methane on Ea ...
can be used by methanotroph
Methanotrophs (sometimes called methanophiles) are prokaryotes that metabolize methane as their source of carbon and chemical energy. They are bacteria or archaea, can grow aerobically or anaerobically, and require single-carbon compounds to s ...
ic bacteria as both a source of electron
The electron ( or ) is a subatomic particle with a negative one elementary electric charge. Electrons belong to the first generation of the lepton particle family,
and are generally thought to be elementary particles because they have no ...
s and a substrate for carbon anabolism.
In many ways, bacterial metabolism provides traits that are useful for ecological stability and for human society. One example is that some bacteria called diazotroph Diazotrophs are bacteria and archaea that fix gaseous nitrogen in the atmosphere into a more usable form such as ammonia.
A diazotroph is a microorganism that is able to grow without external sources of fixed nitrogen. Examples of organisms that ...
s have the ability to fix nitrogen gas using the enzyme nitrogenase
Nitrogenases are enzymes () that are produced by certain bacteria, such as cyanobacteria (blue-green bacteria) and rhizobacteria. These enzymes are responsible for the reduction of nitrogen (N2) to ammonia (NH3). Nitrogenases are the only fa ...
. This environmentally important trait can be found in bacteria of most metabolic types listed above. This leads to the ecologically important processes of denitrification
Denitrification is a microbially facilitated process where nitrate (NO3−) is reduced and ultimately produces molecular nitrogen (N2) through a series of intermediate gaseous nitrogen oxide products. Facultative anaerobic bacteria perform denitr ...
, sulfate reduction, and acetogenesis Acetogenesis is a process through which acetate is produced either by the reduction of CO2 or by the reduction of organic acids, rather than by the oxidative breakdown of carbohydrates or ethanol, as with acetic acid bacteria.
The different bac ...
, respectively. Bacterial metabolic processes are also important in biological responses to pollution
Pollution is the introduction of contaminants into the natural environment that cause adverse change. Pollution can take the form of any substance (solid, liquid, or gas) or energy (such as radioactivity, heat, sound, or light). Pollutants, the ...
; for example, sulfate-reducing bacteria
Sulfate-reducing microorganisms (SRM) or sulfate-reducing prokaryotes (SRP) are a group composed of sulfate-reducing bacteria (SRB) and sulfate-reducing archaea (SRA), both of which can perform anaerobic respiration utilizing sulfate () as termina ...
are largely responsible for the production of the highly toxic forms of mercury ( methyl- and dimethylmercury) in the environment. Non-respiratory anaerobes use fermentation to generate energy and reducing power, secreting metabolic by-products (such as ethanol in brewing) as waste. Facultative anaerobes can switch between fermentation and different terminal electron acceptor
An electron acceptor is a chemical entity that accepts electrons transferred to it from another compound. It is an oxidizing agent that, by virtue of its accepting electrons, is itself reduced in the process. Electron acceptors are sometimes mista ...
s depending on the environmental conditions in which they find themselves.
Growth and reproduction
 Unlike in multicellular organisms, increases in cell size (cell growth) and reproduction by
Unlike in multicellular organisms, increases in cell size (cell growth) and reproduction by cell division
Cell division is the process by which a parent cell divides into two daughter cells. Cell division usually occurs as part of a larger cell cycle in which the cell grows and replicates its chromosome(s) before dividing. In eukaryotes, there ar ...
are tightly linked in unicellular organisms. Bacteria grow to a fixed size and then reproduce through binary fission, a form of asexual reproduction. Under optimal conditions, bacteria can grow and divide extremely rapidly, and some bacterial populations can double as quickly as every 17 minutes. In cell division, two identical clone (genetics), clone daughter cells are produced. Some bacteria, while still reproducing asexually, form more complex reproductive structures that help disperse the newly formed daughter cells. Examples include fruiting body formation by myxobacteria and aerial hyphae formation by '' Streptomyces'' species, or budding. Budding involves a cell forming a protrusion that breaks away and produces a daughter cell.
In the laboratory, bacteria are usually grown using solid or liquid media. Solid Growth medium, growth media, such as agar plates, are used to Isolation (microbiology), isolate pure cultures of a bacterial strain. However, liquid growth media are used when the measurement of growth or large volumes of cells are required. Growth in stirred liquid media occurs as an even cell suspension, making the cultures easy to divide and transfer, although isolating single bacteria from liquid media is difficult. The use of selective media (media with specific nutrients added or deficient, or with antibiotics added) can help identify specific organisms.
Most laboratory techniques for growing bacteria use high levels of nutrients to produce large amounts of cells cheaply and quickly. However, in natural environments, nutrients are limited, meaning that bacteria cannot continue to reproduce indefinitely. This nutrient limitation has led the evolution of different growth strategies (see r/K selection theory). Some organisms can grow extremely rapidly when nutrients become available, such as the formation of algal bloom, algal and cyanobacterial blooms that often occur in lakes during the summer. Other organisms have adaptations to harsh environments, such as the production of multiple antibiotics by streptomyces that inhibit the growth of competing microorganisms. In nature, many organisms live in communities (e.g., biofilm
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular ...
s) that may allow for increased supply of nutrients and protection from environmental stresses. These relationships can be essential for growth of a particular organism or group of organisms (syntrophy).
Bacterial growth follows four phases. When a population of bacteria first enter a high-nutrient environment that allows growth, the cells need to adapt to their new environment. The first phase of growth is the Stationary phase (biology), lag phase, a period of slow growth when the cells are adapting to the high-nutrient environment and preparing for fast growth. The lag phase has high biosynthesis rates, as proteins necessary for rapid growth are produced. The second phase of growth is the Stationary phase (biology), logarithmic phase, also known as the exponential phase. The log phase is marked by rapid exponential growth. The rate at which cells grow during this phase is known as the ''growth rate'' (''k''), and the time it takes the cells to double is known as the ''generation time'' (''g''). During log phase, nutrients are metabolised at maximum speed until one of the nutrients is depleted and starts limiting growth. The third phase of growth is the ''Stationary phase (biology), stationary phase'' and is caused by depleted nutrients. The cells reduce their metabolic activity and consume non-essential cellular proteins. The stationary phase is a transition from rapid growth to a stress response state and there is increased gene expression, expression of genes involved in DNA repair, antioxidant, antioxidant metabolism and active transport, nutrient transport. The final phase is the Stationary phase (biology), death phase where the bacteria run out of nutrients and die.
Genetics
 Most bacteria have a single circular
Most bacteria have a single circular chromosome
A chromosome is a long DNA molecule with part or all of the genetic material of an organism. In most chromosomes the very long thin DNA fibers are coated with packaging proteins; in eukaryotic cells the most important of these proteins are ...
that can range in size from only 160,000 base pairs in the endosymbiotic
An ''endosymbiont'' or ''endobiont'' is any organism that lives within the body or cells of another organism most often, though not always, in a mutualistic relationship.
(The term endosymbiosis is from the Greek: ἔνδον ''endon'' "within ...
bacteria ''Candidatus Carsonella ruddii, Carsonella ruddii'', to 12,200,000 base pairs (12.2 Mbp) in the soil-dwelling bacteria ''Sorangium cellulosum''. There are many exceptions to this, for example some '' Streptomyces'' and ''Borrelia'' species contain a single linear chromosome, while some ''Vibrio'' species contain more than one chromosome. Bacteria can also contain plasmids, small extra-chromosomal molecules of DNA that may contain genes for various useful functions such as antibiotic resistance, metabolic capabilities, or various virulence, virulence factors.
Bacteria genomes usually encode a few hundred to a few thousand genes. The genes in bacterial genomes are usually a single continuous stretch of DNA and although several different types of introns do exist in bacteria, these are much rarer than in eukaryotes.
Bacteria, as asexual organisms, inherit an identical copy of the parent's genomes and are Clonal colony, clonal. However, all bacteria can evolve by selection on changes to their genetic material DNA caused by genetic recombination or mutations. Mutations come from errors made during the replication of DNA or from exposure to mutagens. Mutation rates vary widely among different species of bacteria and even among different clones of a single species of bacteria. Genetic changes in bacterial genomes come from either random mutation during replication or "stress-directed mutation", where genes involved in a particular growth-limiting process have an increased mutation rate.
Some bacteria also transfer genetic material between cells. This can occur in three main ways. First, bacteria can take up exogenous DNA from their environment, in a process called transformation (genetics), transformation. Many bacteria can natural competence, naturally take up DNA from the environment, while others must be chemically altered in order to induce them to take up DNA. The development of competence in nature is usually associated with stressful environmental conditions, and seems to be an adaptation for facilitating repair of DNA damage in recipient cells. The second way bacteria transfer genetic material is by transduction (genetics), transduction, when the integration of a bacteriophage introduces foreign DNA into the chromosome. Many types of bacteriophage exist, some infect and lytic cycle, lyse their host (biology), host bacteria, while others insert into the bacterial chromosome. Bacteria resist phage infection through restriction modification systems that degrade foreign DNA, and a system that uses CRISPR sequences to retain fragments of the genomes of phage that the bacteria have come into contact with in the past, which allows them to block virus replication through a form of RNA interference. The third method of gene transfer is conjugation
Conjugation or conjugate may refer to:
Linguistics
* Grammatical conjugation, the modification of a verb from its basic form
* Emotive conjugation or Russell's conjugation, the use of loaded language
Mathematics
* Complex conjugation, the chang ...
, whereby DNA is transferred through direct cell contact. In ordinary circumstances, transduction, conjugation, and transformation involve transfer of DNA between individual bacteria of the same species, but occasionally transfer may occur between individuals of different bacterial species and this may have significant consequences, such as the transfer of antibiotic resistance. In such cases, gene acquisition from other bacteria or the environment is called horizontal gene transfer and may be common under natural conditions.
Behaviour
Movement
electrochemical gradient
An electrochemical gradient is a gradient of electrochemical potential, usually for an ion that can move across a membrane. The gradient consists of two parts, the chemical gradient, or difference in solute concentration across a membrane, and ...
across the membrane for power.
 Bacteria can use flagella in different ways to generate different kinds of movement. Many bacteria (such as ''Escherichia coli, E. coli'') have two distinct modes of movement: forward movement (swimming) and tumbling. The tumbling allows them to reorient and makes their movement a three-dimensional random walk. Bacterial species differ in the number and arrangement of flagella on their surface; some have a single flagellum (''Flagellum#Flagellar arrangement schemes, monotrichous''), a flagellum at each end (''Flagellum#Flagellar arrangement schemes, amphitrichous''), clusters of flagella at the poles of the cell (''Flagellum#Flagellar arrangement schemes, lophotrichous''), while others have flagella distributed over the entire surface of the cell (''Flagellum#Flagellar arrangement schemes, peritrichous''). The flagella of a unique group of bacteria, the spirochaetes, are found between two membranes in the periplasmic space. They have a distinctive helix, helical body that twists about as it moves.
Two other types of bacterial motion are called twitching motility that relies on a structure called the pilus#Type IV pili, type IV pilus, and Bacterial gliding, gliding motility, that uses other mechanisms. In twitching motility, the rod-like pilus extends out from the cell, binds some substrate, and then retracts, pulling the cell forward.
Motile bacteria are attracted or repelled by certain stimulus (physiology), stimuli in behaviours called ''taxis, taxes'': these include chemotaxis, phototaxis, taxis, energy taxis, and magnetotaxis. In one peculiar group, the myxobacteria, individual bacteria move together to form waves of cells that then differentiate to form fruiting bodies containing spores. The myxobacteria move only when on solid surfaces, unlike ''E. coli'', which is motile in liquid or solid media.
Several ''Listeria'' and ''Shigella'' species move inside host cells by usurping the
Bacteria can use flagella in different ways to generate different kinds of movement. Many bacteria (such as ''Escherichia coli, E. coli'') have two distinct modes of movement: forward movement (swimming) and tumbling. The tumbling allows them to reorient and makes their movement a three-dimensional random walk. Bacterial species differ in the number and arrangement of flagella on their surface; some have a single flagellum (''Flagellum#Flagellar arrangement schemes, monotrichous''), a flagellum at each end (''Flagellum#Flagellar arrangement schemes, amphitrichous''), clusters of flagella at the poles of the cell (''Flagellum#Flagellar arrangement schemes, lophotrichous''), while others have flagella distributed over the entire surface of the cell (''Flagellum#Flagellar arrangement schemes, peritrichous''). The flagella of a unique group of bacteria, the spirochaetes, are found between two membranes in the periplasmic space. They have a distinctive helix, helical body that twists about as it moves.
Two other types of bacterial motion are called twitching motility that relies on a structure called the pilus#Type IV pili, type IV pilus, and Bacterial gliding, gliding motility, that uses other mechanisms. In twitching motility, the rod-like pilus extends out from the cell, binds some substrate, and then retracts, pulling the cell forward.
Motile bacteria are attracted or repelled by certain stimulus (physiology), stimuli in behaviours called ''taxis, taxes'': these include chemotaxis, phototaxis, taxis, energy taxis, and magnetotaxis. In one peculiar group, the myxobacteria, individual bacteria move together to form waves of cells that then differentiate to form fruiting bodies containing spores. The myxobacteria move only when on solid surfaces, unlike ''E. coli'', which is motile in liquid or solid media.
Several ''Listeria'' and ''Shigella'' species move inside host cells by usurping the cytoskeleton
The cytoskeleton is a complex, dynamic network of interlinking protein filaments present in the cytoplasm of all cells, including those of bacteria and archaea. In eukaryotes, it extends from the cell nucleus to the cell membrane and is com ...
, which is normally used to move organelles inside the cell. By promoting actin biopolymer, polymerisation at one pole of their cells, they can form a kind of tail that pushes them through the host cell's cytoplasm.
Communication
A few bacteria have chemical systems that generate light. This bioluminescence often occurs in bacteria that live in association with fish, and the light probably serves to attract fish or other large animals. Bacteria often function as multicellular aggregates known as biofilms, exchanging a variety of molecular signals for Cell signaling, inter-cell communication, and engaging in coordinated multicellular behaviour. The communal benefits of multicellular cooperation include a cellular division of labour, accessing resources that cannot effectively be used by single cells, collectively defending against antagonists, and optimising population survival by differentiating into distinct cell types. For example, bacteria in biofilms can have more than 500 times increased resistance to Bactericide, antibacterial agents than individual "planktonic" bacteria of the same species. One type of inter-cellular communication by a molecular signal is calledquorum sensing
In biology, quorum sensing or quorum signalling (QS) is the ability to detect and respond to cell population density by gene regulation. As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at ...
, which serves the purpose of determining whether there is a local population density that is sufficiently high that it is productive to invest in processes that are only successful if large numbers of similar organisms behave similarly, as in excreting digestive enzymes or emitting light.
Quorum sensing allows bacteria to coordinate gene expression, and enables them to produce, release and detect autoinducers or pheromones which accumulate with the growth in cell population.
Classification and identification

 Scientific classification, Classification seeks to describe the diversity of bacterial species by naming and grouping organisms based on similarities. Bacteria can be classified on the basis of cell structure, Cell metabolism, cellular metabolism or on differences in cell components, such as DNA, fatty acids, pigments,
Scientific classification, Classification seeks to describe the diversity of bacterial species by naming and grouping organisms based on similarities. Bacteria can be classified on the basis of cell structure, Cell metabolism, cellular metabolism or on differences in cell components, such as DNA, fatty acids, pigments, antigen
In immunology, an antigen (Ag) is a molecule or molecular structure or any foreign particulate matter or a pollen grain that can bind to a specific antibody or T-cell receptor. The presence of antigens in the body may trigger an immune respons ...
s and quinones. While these schemes allowed the identification and classification of bacterial strains, it was unclear whether these differences represented variation between distinct species or between strains of the same species. This uncertainty was due to the lack of distinctive structures in most bacteria, as well as lateral gene transfer between unrelated species. Due to lateral gene transfer, some closely related bacteria can have very different morphologies and metabolisms. To overcome this uncertainty, modern bacterial classification emphasises molecular systematics, using genetic techniques such as guanine cytosine GC-content, ratio determination, genome-genome hybridisation, as well as DNA sequencing, sequencing genes that have not undergone extensive lateral gene transfer, such as the ribosomal DNA, rRNA gene. Classification of bacteria is determined by publication in the International Journal of Systematic Bacteriology, and Bergey's Manual of Systematic Bacteriology. The International Committee on Systematic Bacteriology (ICSB) maintains international rules for the naming of bacteria and taxonomic categories and for the ranking of them in the International Code of Nomenclature of Bacteria.
Historically, bacteria were considered a part of the plants, Plantae, the Plant kingdom, and were called "Schizomycetes" (fission-fungi). For this reason, collective bacteria and other microorganisms in a host are often called "flora".
The term "bacteria" was traditionally applied to all microscopic, single-cell prokaryotes. However, molecular systematics showed prokaryotic life to consist of two separate domain (biology), domains, originally called Eubacteria and Archaebacteria, but now called Bacteria and Archaea that evolved independently from an ancient common ancestor. The archaea and eukaryotes are more closely related to each other than either is to the bacteria. These two domains, along with Eukarya, are the basis of the three-domain system, which is currently the most widely used classification system in microbiology. However, due to the relatively recent introduction of molecular systematics and a rapid increase in the number of genome sequences that are available, bacterial classification remains a changing and expanding field. For example, Thomas Cavalier-Smith, Cavalier-Smith argued that the Archaea and Eukaryotes evolved from Gram-positive bacteria.
The identification of bacteria in the laboratory is particularly relevant in medicine, where the correct treatment is determined by the bacterial species causing an infection. Consequently, the need to identify human pathogens was a major impetus for the development of techniques to identify bacteria.
The ''Gram stain
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish b ...
'', developed in 1884 by Hans Christian Gram, characterises bacteria based on the structural characteristics of their cell walls. The thick layers of peptidoglycan in the "Gram-positive" cell wall stain purple, while the thin "Gram-negative" cell wall appears pink. By combining morphology and Gram-staining, most bacteria can be classified as belonging to one of four groups (Gram-positive cocci, Gram-positive bacilli, Gram-negative cocci and Gram-negative bacilli). Some organisms are best identified by stains other than the Gram stain, particularly mycobacteria or ''Nocardia'', which show acid-fastness, acid fastness on Ziehl-Neelsen stain, Ziehl–Neelsen or similar stains. Other organisms may need to be identified by their growth in special media, or by other techniques, such as serology.
Microbiological culture, Culture techniques are designed to promote the growth and identify particular bacteria, while restricting the growth of the other bacteria in the sample. Often these techniques are designed for specific specimens; for example, a sputum sample will be treated to identify organisms that cause pneumonia, while feces, stool specimens are cultured on selective media to identify organisms that cause diarrhea, while preventing growth of non-pathogenic bacteria. Specimens that are normally sterile, such as blood, urine or cerebrospinal fluid, spinal fluid, are cultured under conditions designed to grow all possible organisms. Once a pathogenic organism has been isolated, it can be further characterised by its morphology, growth patterns (such as aerobic organism, aerobic or anaerobic organism, anaerobic growth), hemolysis (microbiology), patterns of hemolysis, and staining.
As with bacterial classification, identification of bacteria is increasingly using molecular methods, and Matrix-assisted laser desorption/ionization, mass spectroscopy. Most bacteria have not been characterised and there are may species that cannot be grown in the laboratory. Diagnostics using DNA-based tools, such as polymerase chain reaction, are increasingly popular due to their specificity and speed, compared to culture-based methods. These methods also allow the detection and identification of "viable but nonculturable" cells that are metabolically active but non-dividing. However, even using these improved methods, the total number of bacterial species is not known and cannot even be estimated with any certainty. Following present classification, there are a little less than 9,300 known species of prokaryotes, which includes bacteria and archaea; but attempts to estimate the true number of bacterial diversity have ranged from 107 to 109 total species—and even these diverse estimates may be off by many orders of magnitude.
Phyla
Valid phyla
The following phyla have been validly published according to the Bacteriological Code: * Acidobacteriota *Actinomycetota
The ''Actinomycetota'' (or ''Actinobacteria'') are a phylum of all gram-positive bacteria. They can be terrestrial or aquatic. They are of great economic importance to humans because agriculture and forests depend on their contributions to s ...
* Aquificota
* Armatimonadota
* Atribacterota
* Bacillota
The Bacillota (synonym Firmicutes) are a phylum of bacteria, most of which have gram-positive cell wall structure. The renaming of phyla such as Firmicutes in 2021 remains controversial among microbiologists, many of whom continue to use the earl ...
* Bacteroidota
* Balneolota
* Bdellovibrionota
* Caldisericota
* Calditrichota
* Campylobacterota
* Chlamydiota
* Chlorobiota
* Chloroflexota
* Chrysiogenota
* Coprothermobacterota
* Deferribacterota
* Deinococcota
* Dictyoglomota
* Elusimicrobiota
* Fibrobacterota
* Fusobacteriota
* Gemmatimonadota
* Ignavibacteriota
* Lentisphaerota
* Mycoplasmatota
* Myxococcota
* Nitrospinota
* Nitrospirota
* Planctomycetota
* Pseudomonadota
* Rhodothermota
* Spirochaetota
* Synergistota
* Thermodesulfobacteriota
* Thermomicrobiota
* Thermotogota
* Verrucomicrobiota
Provisional phyla
The following phyla have been proposed, but have not been validly published according to the Bacteriological Code (including those that have ''candidatus'' status): * "''Candidatus'' Abawacabacteria" * "Abditibacteriota" * "''Candidatus'' Absconditabacteria" * "''Candidatus'' Acetothermia" * "''Candidatus'' Adlerbacteria" * "''Candidatus'' Aerophobetes" * "''Candidatus'' Amesbacteria" * "''Candidatus'' Aminicenantes" * "''Candidatus'' Andersenbacteria" * "''Candidatus'' Azambacteria" * "''Candidatus'' Beckwithbacteria" * "''Candidatus'' Berkelbacteria" * "''Candidatus'' Binatota" * "''Candidatus'' Bipolaricaulota" * "''Candidatus'' Blackallbacteria" * "''Candidatus'' Blackburnbacteria" * "''Candidatus'' Brennerbacteria" * "''Candidatus'' Brownbacteria" * "''Candidatus'' Buchananbacteria" * "''Candidatus'' Caldatribacteriota" * "''Candidatus'' Calescamantes" * "''Candidatus'' Campbellbacteria" * "''Candidatus'' Chisholmbacteria" * "''Candidatus'' Cloacimonetes" * "''Candidatus'' Coatesbacteria" * "''Candidatus'' Collierbacteria" * "''Candidatus'' Colwellbacteria" * "''Candidatus'' Cryosericota" * "''Candidatus'' Curtissbacteria" * "Cyanobacteria" * "''Candidatus'' Dadabacteria" * "''Candidatus'' Daviesbacteria" * "''Candidatus'' Delongbacteria" * "''Candidatus'' Delphibacteria" * "''Candidatus'' Dependentiae" * "''Candidatus'' Desantisbacteria" * "''Candidatus'' Dojkabacteria" * "''Candidatus'' Dormibacteraeota" * "''Candidatus'' Doudnabacteria" * "''Candidatus'' Edwardsbacteria" * "''Candidatus'' Eisenbacteria" * "''Candidatus'' Elulimicrobiota" * "''Candidatus'' Eremiobacterota" * "''Candidatus'' Falkowbacteria" * "''Candidatus'' Fermentibacteria" * "''Candidatus'' Fertabacteria" * "''Candidatus'' Fervidibacteria" * "''Candidatus'' Firestonebacteria" * "''Candidatus'' Fischerbacteria" * "''Candidatus'' Fraserbacteria" * "''Candidatus'' Genascibacteria" * "''Candidatus'' Giovannonibacteria" * "''Candidatus'' Glassbacteria" * "''Candidatus'' Goldbacteria" * "''Candidatus'' Gottesmanbacteria" * "''Candidatus'' Gracilibacteria" * "''Candidatus'' Gribaldobacteria" * "''Candidatus'' Handelsmanbacteria" * "''Candidatus'' Harrisonbacteria" * "''Candidatus'' Howlettbacteria" * "''Candidatus'' Hugbacteria" * "''Candidatus'' Hydrogenedentes" * "''Candidatus'' Hydrothermae" * "''Candidatus'' Hydrothermota" * "''Candidatus'' Jacksonbacteria" * "''Candidatus'' Jorgensenbacteria" * "''Candidatus'' Kaiserbacteria" * "''Candidatus'' Kapabacteria" * "''Candidatus'' Katanobacteria" * "''Candidatus'' Kerfeldbacteria" * "''Candidatus'' Komeilibacteria" * "''Candidatus'' Krumholzibacteriota" * "''Candidatus'' Kryptonia" * "''Candidatus'' Kuenenbacteria" * "''Candidatus'' Lambdaproteobacteria" * "''Candidatus'' Latescibacteria" * "''Candidatus'' Levybacteria" * "''Candidatus'' Lindowbacteria" * "''Candidatus'' Liptonbacteria" * "''Candidatus'' Lloydbacteria" * "''Candidatus'' Magasanikbacteria" * "''Candidatus'' Margulisbacteria" * "''Candidatus'' Marinimicrobia" * "''Candidatus'' Mcinerneyibacteriota" * "''Candidatus'' Melainabacteria" * "''Candidatus'' Microgenomates" * "''Candidatus'' Modulibacteria" * "''Candidatus'' Moisslbacteria" * "''Candidatus'' Montesolbacteria" * "''Candidatus'' Moranbacteria" * "''Candidatus'' Muirbacteria" * "''Candidatus'' Muproteobacteria" * "''Candidatus'' Nealsonbacteria" * "''Candidatus'' Niyogibacteria" * "''Candidatus'' Nomurabacteria" * "''Candidatus'' Omnitrophica" * "''Candidatus'' Pacebacteria" * "''Candidatus'' Parcubacteria" * "''Candidatus'' Parcunitrobacteria" * "''Candidatus'' Peregrinibacteria" * "''Candidatus'' Poribacteria" * "''Candidatus'' Portnoybacteria" * "''Candidatus'' Pyropristinus" * "''Candidatus'' Ratteibacteria" * "''Candidatus'' Raymondbacteria" * "''Candidatus'' Riflebacteria" * "''Candidatus'' Roizmanbacteria" * "''Candidatus'' Rokubacteria" * "''Candidatus'' Ryanbacteria" * "''Candidatus'' Saccharibacteria" * "''Candidatus'' Saganbacteria" * "''Candidatus'' Schekmanbacteria" * "''Candidatus'' Shapirobacteria" * "''Candidatus'' Spechtbacteria" * "''Candidatus'' Stahlbacteria" * "''Candidatus'' Staskawiczbacteria" * "''Candidatus'' Sumerlaeota" * "''Candidatus'' Sungbacteria" * "''Candidatus'' Tagabacteria" * "''Candidatus'' Taylorbacteria" * "''Candidatus'' Tectomicrobia" * "''Candidatus'' Terrybacteria" * "''Candidatus'' Teskebacteria" * "''Candidatus'' Tianyabacteria" * "''Candidatus'' Torokbacteria" * "''Candidatus'' Uhrbacteria" * "''Candidatus'' Veblenbacteria" * "''Candidatus'' Vogelbacteria" * "''Candidatus'' Wallbacteria" * "''Candidatus'' Wildermuthbacteria" * "''Candidatus'' Wirthbacteria" * "''Candidatus'' Woesebacteria" * "''Candidatus'' Wolfebacteria" * "''Candidatus'' Woykebacteria" * "''Candidatus'' Yanofskybacteria" * "''Candidatus'' Yonathbacteria" * "''Candidatus'' Zambryskibacteria" * "''Candidatus'' Zixibacteria"Genera ''incertae sedis''
The following bacteria genera have not been assigned to a phylum, class, or order: * "Fermentobadaceae" Haiying 1995 ** "Guhaiyingella" Haiying 1995 * Not assigned to a family: ** "''Candidatus'' Aegiribacteria" Hamilton et al. 2016 ** Archaeoscillatoriopsis Schopf 1993 ** "Eoleptonema" Awramik et al. 1983 ** "''Candidatus'' Epulonipiscium" corrig. Montgomery and Pollak 1988 ** "''Candidatus'' Ovibacter" corrig. Fenchel and Thar 2004 ** "''Primaevifilum''" Schopf 1983 ** "''Rappaport (bacterium), Rappaport''" Waldman Ben-Asher et al. 2017Interactions with other organisms
 Despite their apparent simplicity, bacteria can form complex associations with other organisms. These symbiosis, symbiotic associations can be divided into parasitism, Mutualism (biology), mutualism and commensalism.
Despite their apparent simplicity, bacteria can form complex associations with other organisms. These symbiosis, symbiotic associations can be divided into parasitism, Mutualism (biology), mutualism and commensalism.
Commensals
The word "commensalism" is derived from the word "commensal", meaning "eating at the same table" and all plants and animals are colonised by commensal bacteria. In humans and other animals millions of them live on the skin, the airways, the gut and other orifices. Referred to as "normal flora", or "commensals", these bacteria usually cause no harm but may occasionally invade other sites of the body and cause infection. ''Escherichia coli'' is a commensal in the human gut but can cause urinary tract infections. Similarly, streptoccoci, which are part of the normal flora of the human mouth, can cause subacute bacterial endocarditis, heart disease.Predators
Some species of bacteria kill and then consume other microorganisms, these species are called ''predatory bacteria''. These include organisms such as ''Myxococcus xanthus'', which forms swarms of cells that kill and digest any bacteria they encounter. Other bacterial predators either attach to their prey in order to digest them and absorb nutrients or invade another cell and multiply inside the cytosol. These predatory bacteria are thought to have evolved from Detritivore, saprophages that consumed dead microorganisms, through adaptations that allowed them to entrap and kill other organisms.Mutualists
Certain bacteria form close spatial associations that are essential for their survival. One such mutualistic association, called interspecies hydrogen transfer, occurs between clusters of anaerobic bacteria that consume organic acids, such as butyric acid or propionic acid, and produce hydrogen, and methanogenic archaea that consume hydrogen. The bacteria in this association are unable to consume the organic acids as this reaction produces hydrogen that accumulates in their surroundings. Only the intimate association with the hydrogen-consuming archaea keeps the hydrogen concentration low enough to allow the bacteria to grow. In soil, microorganisms that reside in the Rhizosphere (ecology), rhizosphere (a zone that includes the root surface and the soil that adheres to the root after gentle shaking) carry out nitrogen fixation, converting nitrogen gas to nitrogenous compounds. This serves to provide an easily absorbable form of nitrogen for many plants, which cannot fix nitrogen themselves. Many other bacteria are found as symbionts Bacteria in the human body, in humans and other organisms. For example, the presence of over 1,000 bacterial species in the normal human gut flora of the intestines can contribute to gut immunity, synthesise vitamins, such as folic acid, vitamin K and biotin, convert Milk protein, sugars to lactic acid (see ''Lactobacillus''), as well as fermenting complex undigestible carbohydrates. The presence of this gut flora also inhibits the growth of potentially pathogenic bacteria (usually through competitive exclusion) and these beneficial bacteria are consequently sold as probiotic dietary supplements. Nearly all Life, animal life is dependent on bacteria for survival as only bacteria and some archaea possess the genes and enzymes necessary to synthesize Vitamin B12, vitamin B12, also known as cobalamin, and provide it through the food chain. Vitamin B12 is a water-soluble vitamin that is involved in themetabolism
Metabolism (, from el, μεταβολή ''metabolē'', "change") is the set of life-sustaining chemical reactions in organisms. The three main functions of metabolism are: the conversion of the energy in food to energy available to run c ...
of every cell of the human body. It is a cofactor (biochemistry), cofactor in DNA replication, DNA synthesis, and in both fatty acid metabolism, fatty acid and amino acid metabolism. It is particularly important in the normal functioning of the nervous system via its role in the myelinogenesis, synthesis of myelin.
Pathogens
 The body is continually exposed to many species of bacteria, including beneficial commensals, which grow on the skin and mucous membranes, and saprophytes, which grow mainly in the soil and in decomposition, decaying matter. The blood and tissue fluids contain nutrients sufficient to sustain the growth of many bacteria. The body has defence mechanisms that enable it to resist microbial invasion of its tissues and give it a natural immune system, immunity or innate immunity, innate resistance against many microorganisms. Unlike some
The body is continually exposed to many species of bacteria, including beneficial commensals, which grow on the skin and mucous membranes, and saprophytes, which grow mainly in the soil and in decomposition, decaying matter. The blood and tissue fluids contain nutrients sufficient to sustain the growth of many bacteria. The body has defence mechanisms that enable it to resist microbial invasion of its tissues and give it a natural immune system, immunity or innate immunity, innate resistance against many microorganisms. Unlike some virus
A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea.
Since Dmitri Ivanovsk ...
es, bacteria evolve relatively slowly so many bacterial diseases also occur in other animals.
If bacteria form a parasitic association with other organisms, they are classed as pathogens. Pathogenic bacteria are a major cause of human death and disease and cause infections such as tetanus
Tetanus, also known as lockjaw, is a bacterial infection caused by ''Clostridium tetani'', and is characterized by muscle spasms. In the most common type, the spasms begin in the jaw and then progress to the rest of the body. Each spasm usually ...
(caused by ''Clostridium tetani
''Clostridium tetani'' is a common soil bacterium and the causative agent of tetanus. Vegetative cells of ''Clostridium tetani'' are usually rod-shaped and up to 2.5 μm long, but they become enlarged and tennis racket- or drumstick-shaped wh ...
''), typhoid fever, diphtheria, syphilis, cholera, foodborne illness, leprosy
Leprosy, also known as Hansen's disease (HD), is a long-term infection by the bacteria ''Mycobacterium leprae'' or ''Mycobacterium lepromatosis''. Infection can lead to damage of the nerves, respiratory tract, skin, and eyes. This nerve damag ...
(caused by ''Mycobacterium leprae'') and tuberculosis
Tuberculosis (TB) is an infectious disease usually caused by '' Mycobacterium tuberculosis'' (MTB) bacteria. Tuberculosis generally affects the lungs, but it can also affect other parts of the body. Most infections show no symptoms, i ...
(caused by ''Mycobacterium tuberculosis''). A pathogenic cause for a known medical disease may only be discovered many years later, as was the case with ''Helicobacter pylori'' and Timeline of peptic ulcer disease and Helicobacter pylori, peptic ulcer disease. Bacterial diseases are also important in agriculture, with bacteria causing leaf spot, fire blight and Wilting, wilts in plants, as well as Johne's disease, Mastitis in dairy cattle, mastitis, salmonellosis, salmonella and anthrax in farm animals.
 Each species of pathogen has a characteristic spectrum of interactions with its human host (biology), hosts. Some organisms, such as ''Staphylococcus'' or ''Streptococcus'', can cause skin infections, pneumonia, meningitis and sepsis, a systemic Inflammation, inflammatory response producing shock (circulatory), shock, massive vasodilator, vasodilation and death. Yet these organisms are also part of the normal human flora and usually exist on the skin or in the nose without causing any disease at all. Other organisms invariably cause disease in humans, such as ''Rickettsia'', which are obligate intracellular parasites able to grow and reproduce only within the cells of other organisms. One species of ''Rickettsia'' causes typhus, while another causes Rocky Mountain spotted fever. ''Chlamydia (bacterium), Chlamydia'', another phylum of obligate intracellular parasites, contains species that can cause pneumonia or urinary tract infection and may be involved in coronary heart disease. Some species, such as ''
Each species of pathogen has a characteristic spectrum of interactions with its human host (biology), hosts. Some organisms, such as ''Staphylococcus'' or ''Streptococcus'', can cause skin infections, pneumonia, meningitis and sepsis, a systemic Inflammation, inflammatory response producing shock (circulatory), shock, massive vasodilator, vasodilation and death. Yet these organisms are also part of the normal human flora and usually exist on the skin or in the nose without causing any disease at all. Other organisms invariably cause disease in humans, such as ''Rickettsia'', which are obligate intracellular parasites able to grow and reproduce only within the cells of other organisms. One species of ''Rickettsia'' causes typhus, while another causes Rocky Mountain spotted fever. ''Chlamydia (bacterium), Chlamydia'', another phylum of obligate intracellular parasites, contains species that can cause pneumonia or urinary tract infection and may be involved in coronary heart disease. Some species, such as ''Pseudomonas aeruginosa
''Pseudomonas aeruginosa'' is a common encapsulated, gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, ''P. aerug ...
'', ''Burkholderia cenocepacia'', and ''Mycobacterium avium complex, Mycobacterium avium'', are opportunistic infection, opportunistic pathogens and cause disease mainly in people who are immunosuppression, immunosuppressed or have cystic fibrosis. Some bacteria produce Microbial toxin, toxins, which cause diseases. These are endotoxins, which come from broken bacterial cells, and exotoxins, which are produced by bacteria and released into the environment. The bacterium ''Clostridium botulinum'' for example, produces a powerful exotoxin that cause respiratory paralysis, and ''Salmonellae'' produce an endotoxin that causes gastroenteritis. Some exotoxins can be converted to toxoids, which are used as vaccines to prevent the disease.
Bacterial infections may be treated with antibiotics, which are classified as Bactericide, bacteriocidal if they kill bacteria or bacteriostatic if they just prevent bacterial growth. There are many types of antibiotics, and each class enzyme inhibitor, inhibits a process that is different in the pathogen from that found in the host. An example of how antibiotics produce selective toxicity are chloramphenicol and puromycin, which inhibit the bacterial ribosome, but not the structurally different eukaryotic ribosome. Antibiotics are used both in treating human disease and in intensive farming to promote animal growth, where they may be contributing to the rapid development of antibiotic resistance in bacterial populations. Infections can be prevented by antiseptic measures such as sterilising the skin prior to piercing it with the needle of a syringe, and by proper care of indwelling catheters. Surgical and dental instruments are also sterilization (microbiology), sterilised to prevent contamination by bacteria. Disinfectants such as bleach are used to kill bacteria or other pathogens on surfaces to prevent contamination and further reduce the risk of infection.
Significance in technology and industry
Bacteria, often lactic acid bacteria, such as ''Lactobacillus'' species and ''Lactococcus'' species, in combination with yeasts and Mold (fungus), moulds, have been used for thousands of years in the preparation of fermentation (food), fermented foods, such as cheese, Pickling, pickles, soy sauce, sauerkraut, vinegar, wine andyogurt
Yogurt (; , from tr, yoğurt, also spelled yoghurt, yogourt or yoghourt) is a food produced by bacterial fermentation of milk. The bacteria used to make yogurt are known as ''yogurt cultures''. Fermentation of sugars in the milk by these bac ...
.
The ability of bacteria to degrade a variety of organic compounds is remarkable and has been used in waste processing and bioremediation. Bacteria capable of digesting the hydrocarbons in petroleum are often used to clean up oil spills. Fertiliser was added to some of the beaches in Prince William Sound in an attempt to promote the growth of these naturally occurring bacteria after the 1989 Exxon Valdez oil spill, ''Exxon Valdez'' oil spill. These efforts were effective on beaches that were not too thickly covered in oil. Bacteria are also used for the bioremediation of industrial toxic wastes. In the chemical industry, bacteria are most important in the production of enantiomerically pure chemicals for use as pharmaceutical company, pharmaceuticals or agrichemicals.
Bacteria can also be used in the place of pesticides in the biological pest control. This commonly involves ''Bacillus thuringiensis'' (also called BT), a Gram-positive, soil dwelling bacterium. Subspecies of this bacteria are used as a Lepidopteran-specific insecticides under trade names such as Dipel and Thuricide. Because of their specificity, these pesticides are regarded as environmentally friendly, with little or no effect on humans, wildlife, pollinators and most other beneficial insects.
Because of their ability to quickly grow and the relative ease with which they can be manipulated, bacteria are the workhorses for the fields of molecular biology, genetics and biochemistry. By making mutations in bacterial DNA and examining the resulting phenotypes, scientists can determine the function of genes, enzyme
Enzymes () are proteins that act as biological catalysts by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products ...
s and metabolic pathways in bacteria, then apply this knowledge to more complex organisms. This aim of understanding the biochemistry of a cell reaches its most complex expression in the synthesis of huge amounts of enzyme kinetics, enzyme kinetic and gene expression data into mathematical models of entire organisms. This is achievable in some well-studied bacteria, with models of ''Escherichia coli'' metabolism now being produced and tested. This understanding of bacterial metabolism and genetics allows the use of biotechnology to bioengineering, bioengineer bacteria for the production of therapeutic proteins, such as insulin, growth factors, or antibody, antibodies.
Because of their importance for research in general, samples of bacterial strains are isolated and preserved in Biological Resource Centers. This ensures the availability of the strain to scientists worldwide.
History of bacteriology
Bacteria were first observed by the Dutch microscopist Antonie van Leeuwenhoek in 1676, using a single-lens microscope of his own design. He then published his observations in a series of letters to the Royal Society of London. Bacteria were Leeuwenhoek's most remarkable microscopic discovery. They were just at the limit of what his simple lenses could make out and, in one of the most striking hiatuses in the history of science, no one else would see them again for over a century. His observations had also included protozoans which he called animalcules, and his findings were looked at again in the light of the more recent findings of cell theory. Christian Gottfried Ehrenberg introduced the word "bacterium" in 1828. In fact, his ''Bacterium (genus), Bacterium'' was a genus that contained non-spore-forming rod-shaped bacteria, as opposed to ''Bacillus'', a genus of spore-forming rod-shaped bacteria defined by Ehrenberg in 1835. Louis Pasteur demonstrated in 1859 that the growth of microorganisms causes the fermentation (food), fermentation process, and that this growth is not due to spontaneous generation (yeasts and Mold (fungus), molds, commonly associated with fermentation, are not bacteria, but ratherfungi
A fungus ( : fungi or funguses) is any member of the group of eukaryotic organisms that includes microorganisms such as yeasts and molds, as well as the more familiar mushrooms. These organisms are classified as a kingdom, separately from ...
). Along with his contemporary Robert Koch, Pasteur was an early advocate of the germ theory of disease. Before them, Ignaz Semmelweis and Joseph Lister had realised the importance of sanitized hands in medical work. Semmelweis ideas was rejected and his book on the topic condemned by the medical community, but after Lister doctors started sanitizing their hands in the 1870s. While Semmelweis who started with rules about handwashing in his hospital in the 1840s predated the spread of the ideas about germs themselves and attributed diseases to "decomposing animal organic matter", Lister was active later.
Robert Koch, a pioneer in medical microbiology, worked on cholera, anthrax and tuberculosis
Tuberculosis (TB) is an infectious disease usually caused by '' Mycobacterium tuberculosis'' (MTB) bacteria. Tuberculosis generally affects the lungs, but it can also affect other parts of the body. Most infections show no symptoms, i ...
. In his research into tuberculosis Koch finally proved the germ theory, for which he received a Nobel Prize in Physiology or Medicine, Nobel Prize in 1905. In Koch's postulates, he set out criteria to test if an organism is the cause of a disease, and these postulates are still used today.
Ferdinand Cohn is said to be a founder of bacteriology, studying bacteria from 1870. Cohn was the first to classify bacteria based on their morphology.
Though it was known in the nineteenth century that bacteria are the cause of many diseases, no effective antiseptic, antibacterial treatments were available. In 1910, Paul Ehrlich developed the first antibiotic, by changing dyes that selectively stained ''Treponema pallidum''—the spirochaete that causes syphilis—into compounds that selectively killed the pathogen. Ehrlich had been awarded a 1908 Nobel Prize for his work on immunology, and pioneered the use of stains to detect and identify bacteria, with his work being the basis of the Gram stain
In microbiology and bacteriology, Gram stain (Gram staining or Gram's method), is a method of staining used to classify bacterial species into two large groups: gram-positive bacteria and gram-negative bacteria. The name comes from the Danish b ...
and the Ziehl–Neelsen stain.
A major step forward in the study of bacteria came in 1977 when Carl Woese recognised that archaea have a separate line of evolutionary descent from bacteria. This new phylogenetic taxonomy
Taxonomy is the practice and science of categorization or classification.
A taxonomy (or taxonomical classification) is a scheme of classification, especially a hierarchical classification, in which things are organized into groups or types. ...
depended on the sequencing of 16S ribosomal RNA, and divided prokaryotes into two evolutionary domains, as part of the three-domain system.
See also
* Genetically modified bacteria * Marine prokaryotesReferences
Bibliography
* * * * * *External links
On-line text book on bacteriology (2015)
{{Authority control Bacteria, Bacteriology Domains (biology)