|
Tudor Domain
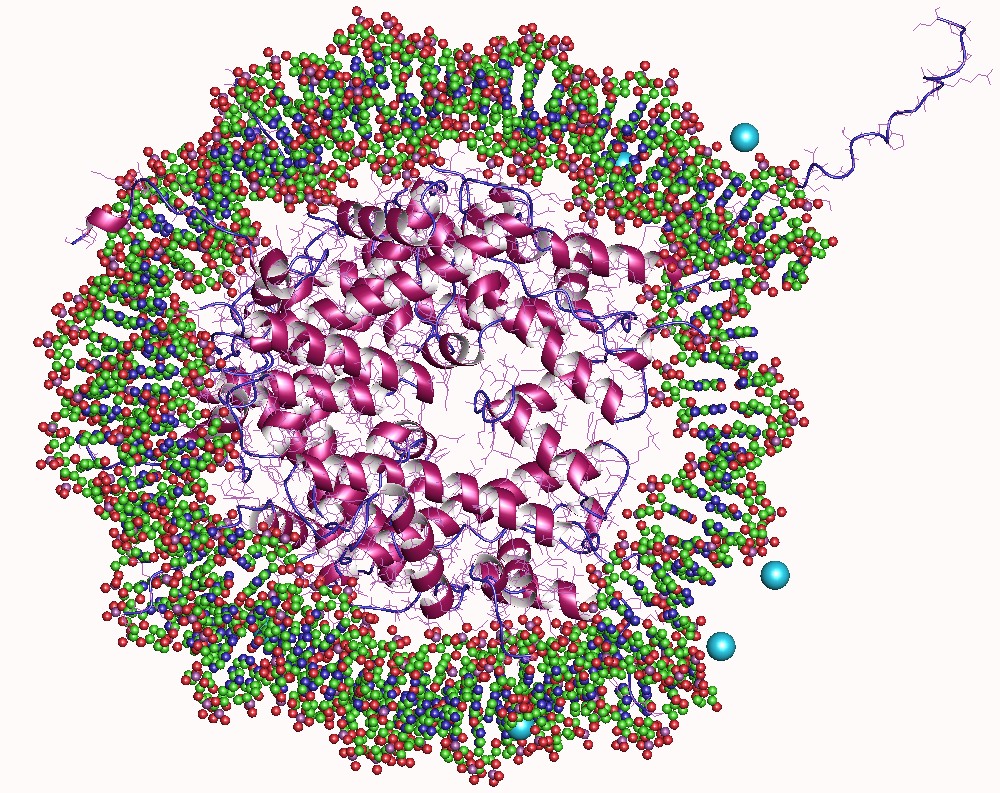
In molecular biology, a Tudor domain is a conserved protein structural domain originally identified in the Tudor protein encoded in Drosophila ''Drosophila'' () is a genus of flies, belonging to the family Drosophilidae, whose members are often called "small fruit flies" or (less frequently) pomace flies, vinegar flies, or wine flies, a reference to the characteristic of many s .... The Tudor gene was found in a Drosophila screen for maternal factors that regulate embryonic development or fertility. Mutations here are lethal for offspring, inspiring the name Tudor, as a reference to the Tudor King Henry VIII and the several miscarriages experienced by his wives. Structure A Tudor domain is a protein region approximately 60 amino acids in length, which folds into an SH3-like structure with a five-stranded antiparallel beta-barrel form. Tudor domains can further be organized into functional units consisting of either a single Tudor domain, tandem Tudor domains, or hybr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid resid ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TP53BP1
Tumor suppressor p53-binding protein 1 also known as p53-binding protein 1 or 53BP1 is a protein that in humans is encoded by the ''TP53BP1'' gene. Clinical significance 53BP1 is underexpressed in most cases of triple-negative breast cancer. DNA repair DNA double-strand breaks (DSBs) are cytotoxic damages that can be repaired either by the homologous recombinational repair (HR) pathway or by the non-homologous end-joining (NHEJ) pathway. NHEJ, although faster than HR, is less accurate. The early divergent step between the two pathways is end resection, and this step is regulated by numerous factors. In particular, BRCA1 and 53BP1 play a role in determining the balance between the two pathways. 53BP1 restricts resection and promotes NHEJ. Age-associated deficient repair Ordinarily during the G1 phase of the cell cycle, when a sister chromatid is unavailable for HR, NHEJ is the predominant pathway for repairing DNA double-strand breaks (DSBs). However, as individuals ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
AKAP1
A kinase anchor protein 1, mitochondrial is an enzyme that in humans is encoded by the ''AKAP1'' gene. Function The A-kinase anchor proteins (AKAPs) are a group of structurally diverse proteins that have the common function of binding to the regulatory subunit of protein kinase A (PKA) and confining the holoenzyme to discrete locations within the cell. This gene encodes a member of the AKAP family. The encoded protein binds to type I and type II regulatory subunits of PKA and anchors them to the mitochondrion. This protein is speculated to be involved in the cAMP-dependent signal transduction pathway and in directing RNA to a specific cellular compartment. Interactions AKAP1 has been shown to interact Advocates for Informed Choice, dba interACT or interACT Advocates for Intersex Youth, is a 501(c)(3) nonprofit organization using innovative strategies to advocate for the legal and human rights of children with intersex traits. The organizat ... with: * C3orf15, * ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Arginine
Arginine is the amino acid with the formula (H2N)(HN)CN(H)(CH2)3CH(NH2)CO2H. The molecule features a guanidino group appended to a standard amino acid framework. At physiological pH, the carboxylic acid is deprotonated (−CO2−) and both the amino and guanidino groups are protonated, resulting in a cation. Only the -arginine (symbol Arg or R) enantiomer is found naturally. Arg residues are common components of proteins. It is encoded by the codons CGU, CGC, CGA, CGG, AGA, and AGG. The guanidine group in arginine is the precursor for the biosynthesis of nitric oxide. Like all amino acids, it is a white, water-soluble solid. History Arginine was first isolated in 1886 from yellow lupin seedlings by the German chemist Ernst Schulze and his assistant Ernst Steiger. He named it from the Greek ''árgyros'' (ἄργυρος) meaning "silver" due to the silver-white appearance of arginine nitrate crystals. In 1897, Schulze and Ernst Winterstein (1865–1949) determined the struct ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Methylation
In the chemical sciences, methylation denotes the addition of a methyl group on a substrate, or the substitution of an atom (or group) by a methyl group. Methylation is a form of alkylation, with a methyl group replacing a hydrogen atom. These terms are commonly used in chemistry, biochemistry, soil science, and the biological sciences. In biological systems, methylation is catalyzed by enzymes; such methylation can be involved in modification of heavy metals, regulation of gene expression, regulation of protein function, and RNA processing. In vitro methylation of tissue samples is also one method for reducing certain histological staining artifacts. The reverse of methylation is demethylation. In biology In biological systems, methylation is accomplished by enzymes. Methylation can modify heavy metals, regulate gene expression, RNA processing and protein function. It has been recognized as a key process underlying epigenetics. Methanogenesis Methanogenesis, the proce ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta Sheet
The beta sheet, (β-sheet) (also β-pleated sheet) is a common motif of the regular protein secondary structure. Beta sheets consist of beta strands (β-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. A β-strand is a stretch of polypeptide chain typically 3 to 10 amino acids long with backbone in an extended conformation. The supramolecular association of β-sheets has been implicated in the formation of the fibrils and protein aggregates observed in amyloidosis, notably Alzheimer's disease. History The first β-sheet structure was proposed by William Astbury in the 1930s. He proposed the idea of hydrogen bonding between the peptide bonds of parallel or antiparallel extended β-strands. However, Astbury did not have the necessary data on the bond geometry of the amino acids in order to build accurate models, especially since he did not then know that the peptide bond was planar. A refined version was ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Survival Of Motor Neuron
Survival of motor neuron or survival motor neuron (SMN) is a protein that in humans is encoded by the ''SMN1'' and '' SMN2'' genes. SMN is found in the cytoplasm of all animal cells and also in the nuclear gems. It functions in transcriptional regulation, telomerase regeneration and cellular trafficking. SMN deficiency, primarily due to mutations in ''SMN1'', results in widespread splicing defects, especially in spinal motor neurons, and is one cause of spinal muscular atrophy. Research also showed a possible role of SMN in neuronal migration and/or differentiation. Function The SMN protein contains GEMIN2-binding, Tudor and YG-Box domains. It localizes to both the cytoplasm and the nucleus. Within the nucleus, the protein localizes to subnuclear bodies called gems which are found near coiled bodies containing high concentrations of small ribonucleoproteins (snRNPs). This protein forms heteromeric complexes with proteins such as GEMIN2 and GEMIN4, and also interacts with se ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wrapped into 30-nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 90 micrometers (0.09 mm) of 30 nm diameter chromatin fibers. There are five families of histones which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B dimers and a H3-H4 tetramer. The tight wrapping of DNA around his ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homology (biology)
In biology, homology is similarity due to shared ancestry between a pair of structures or genes in different taxa. A common example of homologous structures is the forelimbs of vertebrates, where the wings of bats and birds, the arms of primates, the front flippers of whales and the forelegs of four-legged vertebrates like dogs and crocodiles are all derived from the same ancestral tetrapod structure. Evolutionary biology explains homologous structures adapted to different purposes as the result of descent with modification from a common ancestor. The term was first applied to biology in a non-evolutionary context by the anatomist Richard Owen in 1843. Homology was later explained by Charles Darwin's theory of evolution in 1859, but had been observed before this, from Aristotle onwards, and it was explicitly analysed by Pierre Belon in 1555. In developmental biology, organs that developed in the embryo in the same manner and from similar origins, such as from matc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Fission Yeast
''Schizosaccharomyces pombe'', also called "fission yeast", is a species of yeast used in traditional brewing and as a model organism in molecular and cell biology. It is a unicellular eukaryote, whose cells are rod-shaped. Cells typically measure 3 to 4 micrometres in diameter and 7 to 14 micrometres in length. Its genome, which is approximately 14.1 million base pairs, is estimated to contain 4,970 protein-coding genes and at least 450 non-coding RNAs. These cells maintain their shape by growing exclusively through the cell tips and divide by medial fission to produce two daughter cells of equal size, which makes them a powerful tool in cell cycle research. Fission yeast was isolated in 1893 by Paul Lindner from East African millet beer. The species name ''pombe'' is the Swahili word for beer. It was first developed as an experimental model in the 1950s: by Urs Leupold for studying genetics, and by Murdoch Mitchison for studying the cell cycle. Paul Nurse, a fission yeast ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Structure Of JMJD2A Hybrid Tudor Domains
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residues in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |