|
Tac-Promoter
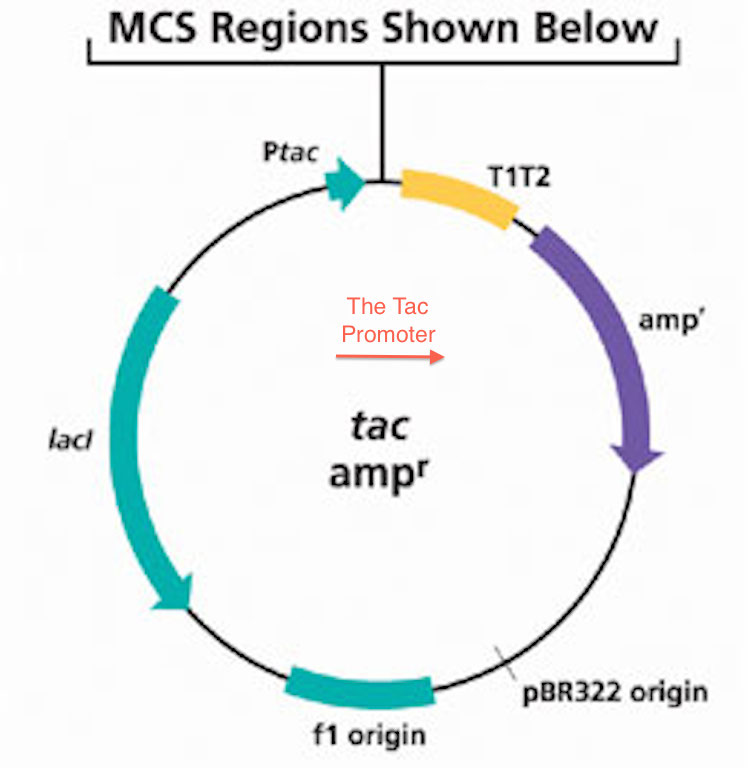
The Tac-Promoter (abbreviated as Ptac), or tac vector is a synthetically produced DNA promoter, produced from the combination of promoters from the trp and lac operons. It is commonly used for protein production in ''Escherichia coli''. Two hybrid promoters functional in ''Escherichia coli'' were constructed. These hybrid promoters, tacI and tacII, were derived from sequences of the trp and the lac UV5 promoters. In the first hybrid promoter (tacI), the DNA upstream of position –20 with respect to the transcriptional start site was derived from the trp promoter. The DNA downstream of position –20 was derived from the lac UV5 promoter. In the second hybrid promoter (tacII), the DNA upstream of position –11 at the Hpa I site within the Pribnow box was derived from the trp promoter. The DNA downstream of position –11 is a 46-base-pair synthetic DNA fragment that specifies part of the hybrid Pribnow box and the entire lac operator. It also specifies a Shine–Dalgarno sequence ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Promoter (genetics)
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism. Promoters control gene expression in bacteria and eukaryotes. RNA polymerase must attach to DNA near a gene for transcription to occur. Promoter DNA sequences provide an enzyme binding site. The -10 sequence is TATAAT. -35 sequences are conserved on average, but not in most promoters. Artificial promoters with conserved -10 and -35 elements transcribe more slowly. All D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tac Promoter
TAC, or tac, may refer to: People * Pablo Tac, US scholar * Pham Cong Tac, a leader of the Cao Dai religion Places * Tác, a village in Fejér County, Hungary Organisations * TAC (building automation), a Swedish building automation company * Tactical Air Command, a former command of the US Air Force * Technical Advisory Council, a committee advising the US FCC * Technical Assistance Center, a company's internal support group for external customers * The Aleut Corporation, Alaska, United States, a Native Regional Corporation * The Analysis Corporation, a private intelligence firm * The Ant Commandos, a company which produces video game console peripherals * The Asatru Community, an inclusive Norse Pagan/Heathen sect; see Kindred (Heathenism) * The Athletics Congress, US sports governing body, became USATF in 1992 * The Architects Collaborative, Cambridge, Massachusetts, US * Thomas Aquinas College, a college with multiple campuses in the US * Treatment Action Campaign, a South A ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Yeast Artificial Chromosome
Yeast artificial chromosomes (YACs) are genetically engineered chromosomes derived from the DNA of the yeast, ''Saccharomyces cerevisiae'', which is then ligated into a bacterial plasmid. By inserting large fragments of DNA, from 100–1000 kb, the inserted sequences can be cloned and physically mapped using a process called chromosome walking. This is the process that was initially used for the Human Genome Project, however due to stability issues, YACs were abandoned for the use of Bacterial artificial chromosomes (BAC). Beginning with the initial research of the Rankin et al., Strul et al., and Hsaio et al., the inherently fragile chromosome was stabilized by discovering the necessary autonomously replicating sequence (ARS); a refined YAC utilizing this data was described in 1983 by Murray et al. The primary components of a YAC are the ARS, centromere, and telomeres from ''S. cerevisiae''. Additionally, selectable marker genes, such as antibiotic resistance and a visible ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Human Artificial Chromosome
A human artificial chromosome (HAC) is a microchromosome that can act as a new chromosome in a population of human cells. That is, instead of 46 chromosomes, the cell could have 47 with the 47th being very small, roughly 6–10megabases (Mb) in size instead of 50–250Mb for natural chromosomes, and able to carry new genes introduced by human researchers. Ideally, researchers could integrate different genes that perform a variety of functions, includindisease defense Alternative methods of creating transgenes, such as utilizing yeast artificial chromosomes and bacterial artificial chromosomes, lead to unpredictable problems. The genetic material introduced by these vectors not only leads to different expression levels, but the inserts also disrupt the original genome. HACs differ in this regard, as they are entirely separate chromosomes. This separation from existing genetic material assumes that no insertional mutants would arise. This stability and accuracy makes HACs preferable t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Isopropyl β-D-1-thiogalactopyranoside
Isopropyl β--1-thiogalactopyranoside (IPTG) is a molecular biology reagent. This compound is a molecular mimic of allolactose, a lactose metabolite that triggers Transcription (genetics), transcription of the lac operon, ''lac'' operon, and it is therefore used to induce protein expression where the gene is under the control of the lac operator. Mechanism of action Like allolactose, IPTG binds to the lac repressor and releases the tetrameric repressor from the lac operator in an allosteric manner, thereby allowing the transcription of genes in the lac operon, such as the gene coding for beta-galactosidase, a hydrolase enzyme that catalyzes the hydrolysis of β-galactosides into monosaccharides. But unlike allolactose, the sulfur (sulfur, S) atom creates a chemical bond which is non-hydrolyzable by the cell, preventing the cell from metabolizing or degrading the inducer. Therefore, its concentration remains constant during an experiment. IPTG uptake by ''E. coli'' can be inde ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Regulation Of Gene Expression
Regulation of gene expression, or gene regulation, includes a wide range of mechanisms that are used by cells to increase or decrease the production of specific gene products (protein or RNA). Sophisticated programs of gene expression are widely observed in biology, for example to trigger developmental pathways, respond to environmental stimuli, or adapt to new food sources. Virtually any step of gene expression can be modulated, from Transcriptional regulation, transcriptional initiation, to RNA processing, and to the post-translational modification of a protein. Often, one gene regulator controls another, and so on, in a gene regulatory network. Gene regulation is essential for viruses, prokaryotes and eukaryotes as it increases the versatility and adaptability of an organism by allowing the cell to express protein when needed. Although as early as 1951, Barbara McClintock showed interaction between two genetic loci, Activator (''Ac'') and Dissociator (''Ds''), in the color f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcription (genetics)
Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules called non-coding RNAs (ncRNAs). mRNA comprises only 1–3% of total RNA samples. Less than 2% of the human genome can be transcribed into mRNA ( Human genome#Coding vs. noncoding DNA), while at least 80% of mammalian genomic DNA can be actively transcribed (in one or more types of cells), with the majority of this 80% considered to be ncRNA. Both DNA and RNA are nucleic acids, which use base pairs of nucleotides as a complementary language. During transcription, a DNA sequence is read by an RNA polymerase, which produces a complementary, antiparallel RNA strand called a primary transcript. Transcription proceeds in the following general steps: # RNA polymerase, together with one or more general transcription factors, binds to promoter DNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that synthesizes RNA from a DNA template. Using the enzyme helicase, RNAP locally opens the double-stranded DNA so that one strand of the exposed nucleotides can be used as a template for the synthesis of RNA, a process called transcription. A transcription factor and its associated transcription mediator complex must be attached to a DNA binding site called a promoter region before RNAP can initiate the DNA unwinding at that position. RNAP not only initiates RNA transcription, it also guides the nucleotides into position, facilitates attachment and elongation, has intrinsic proofreading and replacement capabilities, and termination recognition capability. In eukaryotes, RNAP can build chains as long as 2.4 million nucleotides. RNAP produces RNA that, functionally, is either for protein coding, i.e. messenger RNA (mRNA); or n ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sigma Factor
A sigma factor (σ factor or specificity factor) is a protein needed for initiation of transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to gene promoters. It is homologous to archaeal transcription factor B and to eukaryotic factor TFIIB. The specific sigma factor used to initiate transcription of a given gene will vary, depending on the gene and on the environmental signals needed to initiate transcription of that gene. Selection of promoters by RNA polymerase is dependent on the sigma factor that associates with it. They are also found in plant chloroplasts as a part of the bacteria-like plastid-encoded polymerase (PEP). The sigma factor, together with RNA polymerase, is known as the RNA polymerase holoenzyme. Every molecule of RNA polymerase holoenzyme contains exactly one sigma factor subunit, which in the model bacterium ''Escherichia coli'' is one of those listed below. The number of sigma f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Recombinant Proteins
Recombinant DNA (rDNA) molecules are DNA molecules formed by laboratory methods of genetic recombination (such as molecular cloning) that bring together genetic material from multiple sources, creating sequences that would not otherwise be found in the genome. Recombinant DNA is the general name for a piece of DNA that has been created by combining at least two fragments from two different sources. Recombinant DNA is possible because DNA molecules from all organisms share the same chemical structure, and differ only in the nucleotide sequence within that identical overall structure. Recombinant DNA molecules are sometimes called chimeric DNA, because they can be made of material from two different species, like the mythical chimera. R-DNA technology uses palindromic sequences and leads to the production of sticky and blunt ends. The DNA sequences used in the construction of recombinant DNA molecules can originate from any species. For example, plant DNA may be joined to bacter ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the central dogma of molecular biology first formulated by Francis Crick in 1958, further developed in his 1970 article, and expanded by the subsequent discoveries of reverse transcription and RNA replication. The process of gene expression is used by all known life—eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenotype, '' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |