|
TAR DNA-binding Protein 43
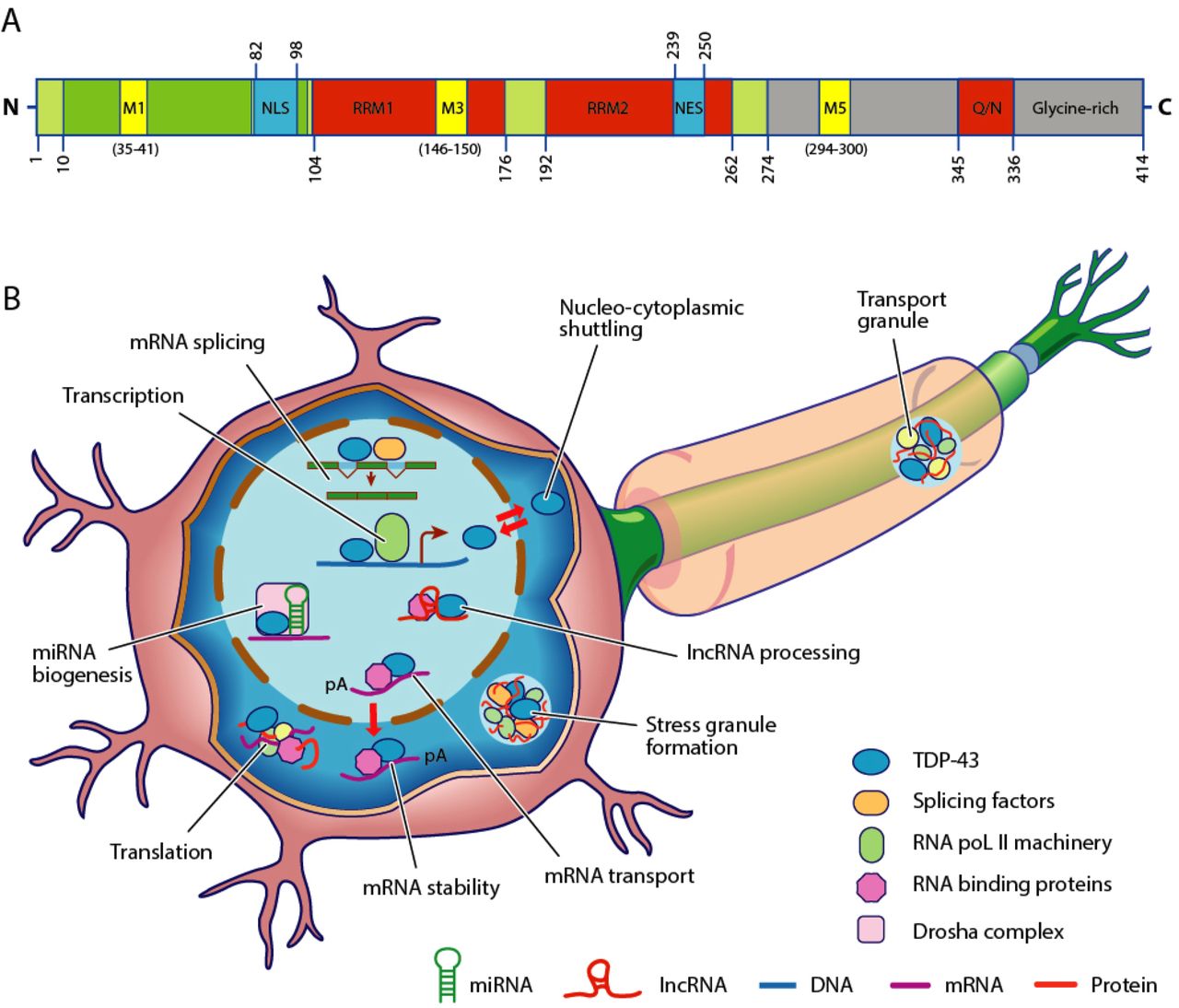
TAR DNA-binding protein 43 (TDP-43, transactive response DNA binding protein 43 kDa) is a protein that in humans is encoded by the ''TARDBP'' gene. Structure TDP-43 is 414 amino acid residues long. It consists of 4 domains: an N-terminal domain spanning residues 1-76 (NTD) with a well-defined fold that has been shown to form a dimer or oligomer; 2 highly conserved folded RNA recognition motifs spanning residues 106-176 (RRM1) and 191-259 (RRM2), respectively, required to bind target RNA and DNA; an unstructured C-terminal domain encompassing residues 274-414 (CTD), which contains a glycine-rich region, is involved in protein-protein interactions, and harbors most of the mutations associated with familial amyotrophic lateral sclerosis. The entire protein devoid of large solubilising tags has been purified. The full-length protein is a dimer. The dimer is formed due to a self-interaction between two NTD domains, where the dimerisation can be propagated to form higher-order ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Atomic Mass Unit
The dalton or unified atomic mass unit (symbols: Da or u) is a non-SI unit of mass widely used in physics and chemistry. It is defined as of the mass of an unbound neutral atom of carbon-12 in its nuclear and electronic ground state and at rest. The atomic mass constant, denoted ''m''u, is defined identically, giving . This unit is commonly used in physics and chemistry to express the mass of atomic-scale objects, such as atoms, molecules, and elementary particles, both for discrete instances and multiple types of ensemble averages. For example, an atom of helium-4 has a mass of . This is an intrinsic property of the isotope and all helium-4 atoms have the same mass. Acetylsalicylic acid (aspirin), , has an average mass of approximately . However, there are no acetylsalicylic acid molecules with this mass. The two most common masses of individual acetylsalicylic acid molecules are , having the most common isotopes, and , in which one carbon is carbon-13. The molecular mass ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TMEM106B
Transmembrane protein 106B is a protein that is encoded by the ''TMEM106B'' gene. It is found primarily within Neuron, neurons and Oligodendrocyte, oligodendrocytes in the central nervous system with its subcellular location being in Lysosome, lysosomal membranes. TMEM106B helps facilitate important functions for maintaining a healthy lysosome, and therefore certain mutations and Polymorphism (biology), polymorphisms can lead to issues with proper lysosomal function. Lysosomes are in charge of clearing out mis-folded proteins and other debris, and thus, play an important role in Neurodegenerative disease, neurodegenerative diseases that are driven by the accumulation of various mis-folded proteins and aggregates. Due to its impact on lysosomal function, TMEM106B has been investigated and found to be associated to multiple neurodegenerative diseases. Structure Gene In humans,''TMEM106B'' is found on chromosome 7 at positions 12211270 - 12243367, totaling 32097 base pairs. Th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Solenoid Protein Domain
Solenoid protein domains are a highly modular type of protein domain. They consist of a chain of nearly identical folds, often simply called tandem repeats. They are extremely common among all types of proteins, though exact figures are unknown. "Repeats" in molecular biology In proteins, a "repeat" is any sequence block that returns more than one time in the sequence, either in an identical or a highly similar form. Repetitiveness does not in itself indicate anything about the structure of the protein. As a "rule of thumb", short repetitive sequences (e.g. those below the length of 10 amino acids) may be intrinsically disordered, and not part of any folded protein domains. Repeats that are at least 30 to 40 amino acids long, are far more likely to be folded as part of a domain. Such long repeats are frequently indicative of the presence of a solenoid domain in the protein. Examples of disordered repetitive sequences include the 7-mer peptide repeats found in the RPB1 subun ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sulfate
The sulfate or sulphate ion is a polyatomic anion with the empirical formula . Salts, acid derivatives, and peroxides of sulfate are widely used in industry. Sulfates occur widely in everyday life. Sulfates are salts of sulfuric acid and many are prepared from that acid. Spelling "Sulfate" is the spelling recommended by IUPAC, but "sulphate" was traditionally used in British English. Structure The sulfate anion consists of a central sulfur atom surrounded by four equivalent oxygen atoms in a tetrahedral arrangement. The symmetry is the same as that of methane. The sulfur atom is in the +6 oxidation state while the four oxygen atoms are each in the −2 state. The sulfate ion carries an overall charge of −2 and it is the conjugate base of the bisulfate (or hydrogensulfate) ion, , which is in turn the conjugate base of , sulfuric acid. Organic sulfate esters, such as dimethyl sulfate, are covalent compounds and esters of sulfuric acid. The tetrahedral molecular geometry of th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helix
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence. The alpha helix is also called a classic Pauling–Corey–Branson α-helix. The name 3.613-helix is also used for this type of helix, denoting the average number of residues per helical turn, with 13 atoms being involved in the ring formed by the hydrogen bond. Among types of local structure in proteins, the α-helix is the most extreme and the most predictable from sequence, as well as the most prevalent. Discovery In the early 1930s, William Astbury showed that there were drastic changes in the X-ray fiber diffraction of moist wool or hair fibers upon significant stretching. The data suggested that the unstretched fibers had a coiled molecular structure with a characteristic repeat of ≈. Astb ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta Sheet
The beta sheet, (β-sheet) (also β-pleated sheet) is a common motif of the regular protein secondary structure. Beta sheets consist of beta strands (β-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. A β-strand is a stretch of polypeptide chain typically 3 to 10 amino acids long with backbone in an extended conformation. The supramolecular association of β-sheets has been implicated in the formation of the fibrils and protein aggregates observed in amyloidosis, notably Alzheimer's disease. History The first β-sheet structure was proposed by William Astbury in the 1930s. He proposed the idea of hydrogen bonding between the peptide bonds of parallel or antiparallel extended β-strands. However, Astbury did not have the necessary data on the bond geometry of the amino acids in order to build accurate models, especially since he did not then know that the peptide bond was planar. A refined versi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin Ligase
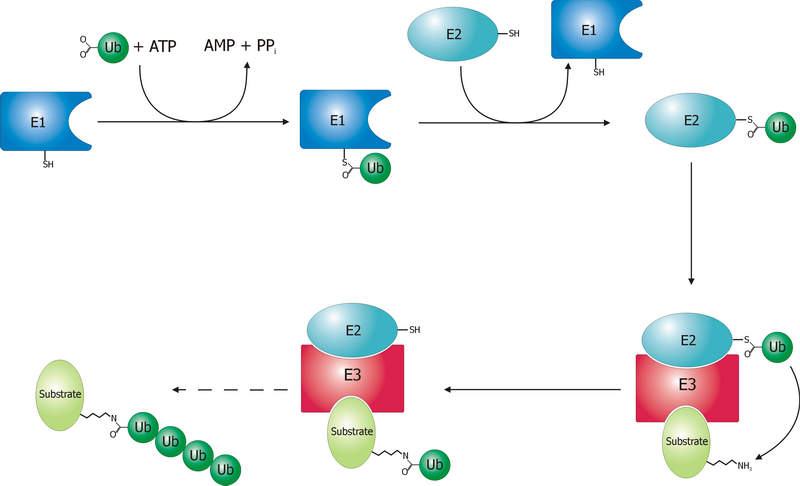
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases reg ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin-like Protein
Ubiquitin-like proteins (UBLs) are a family of small proteins involved in post-translational modification of other proteins in a cell, usually with a regulatory function. The UBL protein family derives its name from the first member of the class to be discovered, ubiquitin (Ub), best known for its role in regulating protein degradation through covalent modification of other proteins. Following the discovery of ubiquitin, many additional evolutionarily related members of the group were described, involving parallel regulatory processes and similar chemistry. UBLs are involved in a widely varying array of cellular functions including autophagy, protein trafficking, inflammation and immune responses, transcription, DNA repair, RNA splicing, and cellular differentiation. Discovery Ubiquitin itself was first discovered in the 1970s and originally named "ubiquitous immunopoietic polypeptide". Subsequently, other proteins with sequence similarity to ubiquitin were occasionally reported i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell (biology)
The cell is the basic structural and functional unit of life forms. Every cell consists of a cytoplasm enclosed within a membrane, and contains many biomolecules such as proteins, DNA and RNA, as well as many small molecules of nutrients and metabolites.Cell Movements and the Shaping of the Vertebrate Body in Chapter 21 of Molecular Biology of the Cell '' fourth edition, edited by Bruce Alberts (2002) published by Garland Science. The Alberts text discusses how the "cellular building blocks" move to shape developing embryos. It is also common to describe small molecules such as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Oligomer
In chemistry and biochemistry, an oligomer () is a molecule that consists of a few repeating units which could be derived, actually or conceptually, from smaller molecules, monomers.Quote: ''Oligomer molecule: A molecule of intermediate relative molecular mass, the structure of which essentially comprises a small plurality of units derived, actually or conceptually, from molecules of lower relative molecular mass.'' The name is composed of Greek elements '' oligo-'', "a few" and '' -mer'', "parts". An adjective form is ''oligomeric''. The oligomer concept is contrasted to that of a polymer, which is usually understood to have a large number of units, possibly thousands or millions. However, there is no sharp distinction between these two concepts. One proposed criterion is whether the molecule's properties vary significantly with the removal of one or a few of the units. An oligomer with a specific number of units is referred to by the Greek prefix denoting that number, wi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Splicing
RNA splicing is a process in molecular biology where a newly-made precursor messenger RNA (pre-mRNA) transcript is transformed into a mature messenger RNA (mRNA). It works by removing all the introns (non-coding regions of RNA) and ''splicing'' back together exons (coding regions). For nuclear-encoded genes, splicing occurs in the nucleus either during or immediately after transcription. For those eukaryotic genes that contain introns, splicing is usually needed to create an mRNA molecule that can be translated into protein. For many eukaryotic introns, splicing occurs in a series of reactions which are catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs). There exist self-splicing introns, that is, ribozymes that can catalyze their own excision from their parent RNA molecule. The process of transcription, splicing and translation is called gene expression, the central dogma of molecular biology. Splicing pathways Several methods of RNA splici ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |