|
SAM Riboswitch
The SAM riboswitch (also known as the S-box leader and the SAM-I riboswitch) is found upstream of a number of genes which code for proteins involved in methionine or cysteine biosynthesis in Gram-positive bacteria. Two SAM riboswitches in ''Bacillus subtilis'' that were experimentally studied act at the level of transcription termination control. The predicted secondary structure consists of a complex stem-loop region followed by a single stem-loop terminator region. An alternative and mutually exclusive form involves bases in the 3' segment of helix 1 with those in the 5' region of helix 5 to form a structure termed the anti-terminator form. When SAM is unbound, the anti-terminator sequence sequesters the terminator sequence so the terminator is unable to form, allowing the polymerase to read-through the downstream gene.Winkler, W., Nahvi, A., Sudarsan, N., Barrick, J., and Breaker, R. (2003) An mRNA structure that controls gene expression by binding S-adenosylmethionine. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
S-Adenosyl Methionine
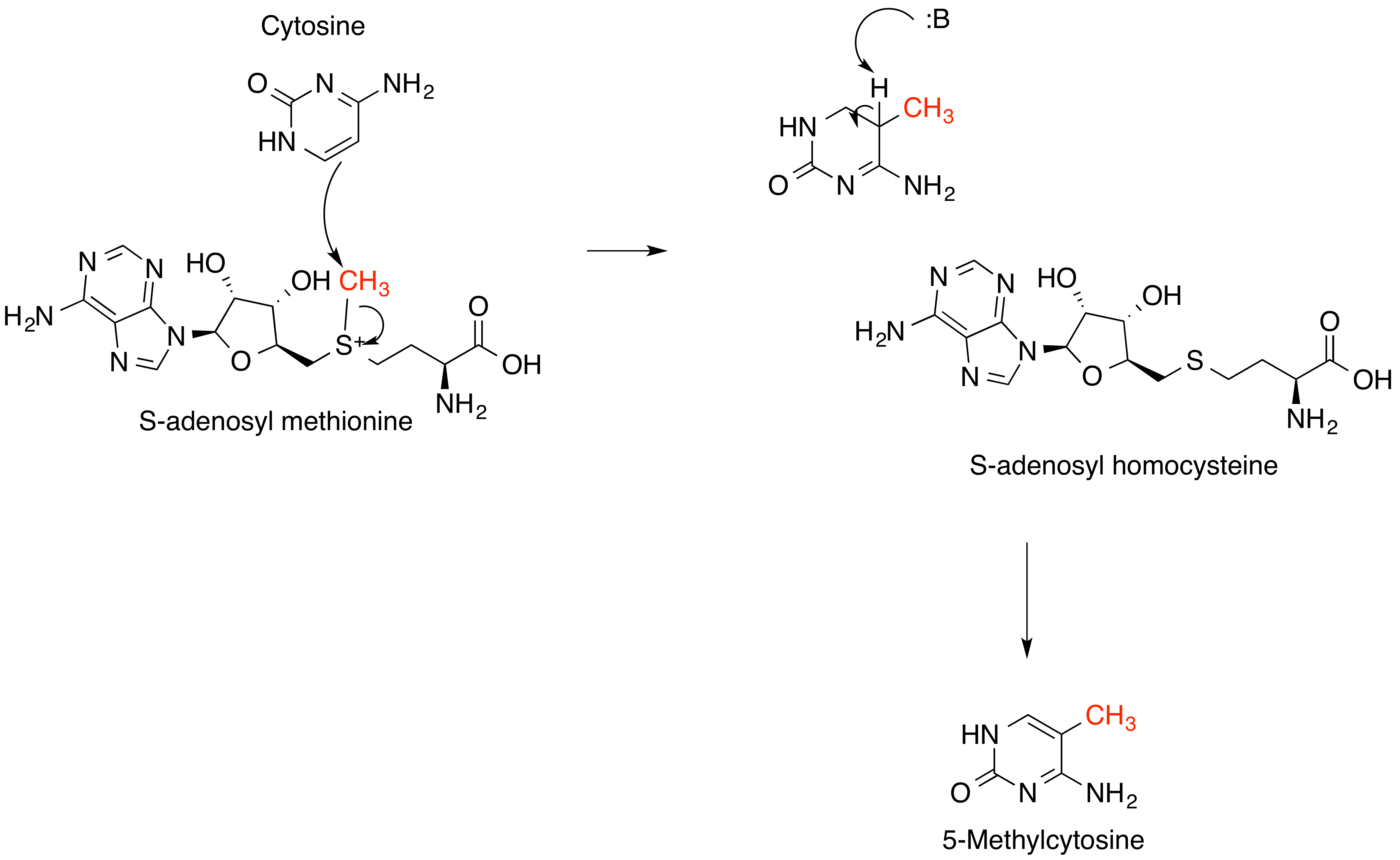
''S''-Adenosyl methionine (SAM), also known under the commercial names of SAMe, SAM-e, or AdoMet, is a common cosubstrate involved in methyl group transfers, transsulfuration, and aminopropylation. Although these anabolic reactions occur throughout the body, most SAM is produced and consumed in the liver. More than 40 methyl transfers from SAM are known, to various substrates such as nucleic acids, proteins, lipids and secondary metabolites. It is made from adenosine triphosphate (ATP) and methionine by methionine adenosyltransferase. SAM was first discovered by Giulio Cantoni in 1952. In bacteria, SAM is bound by the SAM riboswitch, which regulates genes involved in methionine or cysteine biosynthesis. In eukaryotic cells, SAM serves as a regulator of a variety of processes including DNA, tRNA, and rRNA methylation; immune response; amino acid metabolism; transsulfuration; and more. In plants, SAM is crucial to the biosynthesis of ethylene, an important plant hormone and sig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM-VI Riboswitch
SAM-VI is a member of the riboswitch family. It is predominantly found in ''Bifidobacterium'' and exhibits some similarities to the SAM-III ( Smk box) riboswitch class, but lacks most of the highly conserved nucleotides of SAM-III class. SAM-VI aptamers bind the cofactor S-adenosylmethinine SAM (a key metabolite in sulphur metabolism) and discriminate strongly against S-adenosylhomocysteine SAH. The class was discovered by further analysis of Bifido-''meK'' motif RNAs. See also * SAM-I riboswitch * SAM-II riboswitch * SAM-III riboswitch * SAM-IV riboswitch SAM-IV riboswitches are a kind of riboswitch that specifically binds S-adenosylmethionine (SAM), a cofactor used in many methylation reactions. Originally identified by bioinformatics, SAM-IV riboswitches are largely confined to the Actinomyce ... * SAM-V riboswitch References {{reflist Cis-regulatory RNA elements Riboswitch ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM-V Riboswitch
SAM-V riboswitch is the fifth known riboswitch to bind S-adenosyl methionine (SAM). It was first discovered in the marine bacterium '' Candidatus Pelagibacter ubique'' and can also be found in marine metagenomes. SAM-V features a similar consensus sequence and secondary structure as the binding site of SAM-II riboswitch, but bioinformatics scans cluster the two aptamers independently. These similar binding pockets suggest that the two riboswitches have undergone convergent evolution. SAM-binding was confirmed using equilibrium dialysis. The riboswitch has been characterised as a 'tandem riboswitch' - it is able to regulate both translation and transcription. When SAM is present in high concentration, SAM-II will bind its ligand and form a terminator stem to halt transcription. If SAM exists in lower concentrations, SAM-V will be transcribed and, if SAM concentration should then increase, it can bind SAM and occlude the Shine-Dalgarno sequence of the downstream open reading fr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ligand
In coordination chemistry, a ligand is an ion or molecule (functional group) that binds to a central metal atom to form a coordination complex. The bonding with the metal generally involves formal donation of one or more of the ligand's electron pairs, often through Lewis bases. The nature of metal–ligand bonding can range from covalent to ionic. Furthermore, the metal–ligand bond order can range from one to three. Ligands are viewed as Lewis bases, although rare cases are known to involve Lewis acidic "ligands". Metals and metalloids are bound to ligands in almost all circumstances, although gaseous "naked" metal ions can be generated in a high vacuum. Ligands in a complex dictate the reactivity of the central atom, including ligand substitution rates, the reactivity of the ligands themselves, and redox. Ligand selection requires critical consideration in many practical areas, including bioinorganic and medicinal chemistry, homogeneous catalysis, and environmental chemi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM-IV Riboswitch
SAM-IV riboswitches are a kind of riboswitch that specifically binds S-adenosylmethionine (SAM), a cofactor used in many methylation reactions. Originally identified by bioinformatics, SAM-IV riboswitches are largely confined to the Actinomycetales, an order of Bacteria. Conserved features of SAM-IV riboswitch and experiments imply that they probably share a similar SAM-binding site to another class of SAM-binding riboswitches called SAM-I riboswitches. However, the scaffolds of these two types of riboswitch appear to be quite distinct. The structural relationship between these riboswitch types has been studied. See also * SAM-I riboswitch * SAM-II riboswitch The SAM-II riboswitch is a RNA element found predominantly in Alphaproteobacteria that binds S-adenosyl methionine (SAM). Its structure and sequence appear to be unrelated to the SAM riboswitch found in Gram-positive bacteria. This SAM riboswit ... * SAM-III riboswitch * SAM-V riboswitch * SAM-VI riboswitch R ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM–SAH Riboswitch
The SAM–SAH riboswitch is a conserved RNA structure in certain bacteria that binds ''S''-adenosylmethionine (SAM) and ''S''-adenosylhomocysteine (SAH) and is therefore presumed to be a riboswitch. SAM–SAH riboswitches do not share any apparent structural resemblance to known riboswitches that bind SAM or SAH. The binding affinities for both compounds are similar, but binding for SAH is at least somewhat stronger. SAM–SAH riboswitches are exclusively found in Rhodobacterales, an order of alphaproteobacteria. They are always found in the apparent 5' untranslated regions of ''metK'' genes, which encode the enzyme ( Methionine adenosyltransferase) that synthesizes SAM. Given this gene association, it was proposed that SAM–SAH riboswitches more likely function as SAM-sensing RNAs. SAM–SAH riboswitches are relatively small among known riboswitches, which might relate to their inability to discriminate against SAH. However, the ability to reject SAH as a ligand might not ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SMK Box Riboswitch
The SMKbox riboswitch (also known as SAM-III) is an RNA element that regulates gene expression in bacteria. The SMK box riboswitch is found in the 5' UTR of the MetK gene in lactic acid bacteria. The structure of this element changes upon binding to S-adenosyl methionine (SAM) to a conformation that blocks the shine-dalgarno sequence and blocks translation of the gene. There are other known SAM-binding riboswitches such as SAM-I and SAM-II, but these appear to share no similarity in sequence or structure to SAM-III. Structure The crystal structure of the riboswitch from ''E. faecalis'' was solved by X-ray crystallography. The structure showed that the most conserved nucleotides involved in SAM binding were organised around a junction between three helices. In some species there are large insertions of up to 210 nucleotides within this structure. See also * SAH riboswitch * SAM-I riboswitch * SAM-II riboswitch * SAM-IV riboswitch SAM-IV riboswitches are a kind of ri ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SAM-II Riboswitch
The SAM-II riboswitch is a RNA element found predominantly in Alphaproteobacteria that binds S-adenosyl methionine (SAM). Its structure and sequence appear to be unrelated to the SAM riboswitch found in Gram-positive bacteria. This SAM riboswitch is located upstream of the metA and metC genes in Agrobacterium tumefaciens, and other methionine and SAM biosynthesis genes in other alpha-proteobacteria. Like the other SAM riboswitch, it probably functions to turn off expression of these genes in response to elevated SAM levels. A significant variant of SAM-II riboswitches was found in ''Pelagibacter ubique'' and related marine bacteria and called SAM-V. Also, like many structured RNAs, SAM-II riboswitches can tolerate long loops between their stems. Structure The SAM-II riboswitch is short with less than 70 nucleotides and is structurally relatively simple being composed of a single hairpin A hairpin or hair pin is a long device used to hold a person's hair in place. It ma ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Kink Turn
Kink or KINK may refer to: Common uses * Kink (sexuality), a colloquial term for non-normative sexual behavior * Kink, a curvature, bend, or twist Geography * Kink, Iran, a village in Iran * The Kink, a man-made geographic feature in remote eastern Alaska Arts, entertainment, and media * ''Kink'' (film), a documentary about the internet pornography company Kink.com * ''Kink'', an autobiography written by Dave Davies, guitarist for the Kinks *Kink.com, a BDSM-focused Internet pornography company * The Kinks, a British rock band * ''The Kink'' (novel), a 1927 detective novel by Lynn Brock Radio and television * ''KinK'', a Canadian documentary television series profiling some of the more unusual edges of human sexuality * KINK and kink.fm, a radio station in Portland, Oregon, United States * Kink FM, a radio station in the Netherlands People named Kink * Dick Kink (1921–1971), American politician * KiNK (Strahil Velchev), a music producer and DJ in Sofia, Bulgaria * George Ki ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pseudoknot
__NOTOC__ A pseudoknot is a nucleic acid secondary structure containing at least two stem-loop structures in which half of one stem is intercalated between the two halves of another stem. The pseudoknot was first recognized in the turnip yellow mosaic virus in 1982. Pseudoknots fold into knot-shaped three-dimensional conformations but are not true topological knots. Prediction and identification The structural configuration of pseudoknots does not lend itself well to bio-computational detection due to its context-sensitivity or "overlapping" nature. The base pairing in pseudoknots is not well nested; that is, base pairs occur that "overlap" one another in sequence position. This makes the presence of pseudoknots in RNA sequences more difficult to predict by the standard method of dynamic programming, which use a recursive scoring system to identify paired stems and consequently, most cannot detect non-nested base pairs. The newer method of stochastic context-free grammars su ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
X-ray Crystallography
X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. By measuring the angles and intensities of these diffracted beams, a crystallographer can produce a three-dimensional picture of the density of electrons within the crystal. From this electron density, the mean positions of the atoms in the crystal can be determined, as well as their chemical bonds, their crystallographic disorder, and various other information. Since many materials can form crystals—such as salts, metals, minerals, semiconductors, as well as various inorganic, organic, and biological molecules—X-ray crystallography has been fundamental in the development of many scientific fields. In its first decades of use, this method determined the size of atoms, the lengths and types of chemical bonds, and the atomic-scale differences among various mat ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |

4-3D-balls.png)

